-
Notifications
You must be signed in to change notification settings - Fork 8
Open
Labels
bugSomething isn't workingSomething isn't working
Description
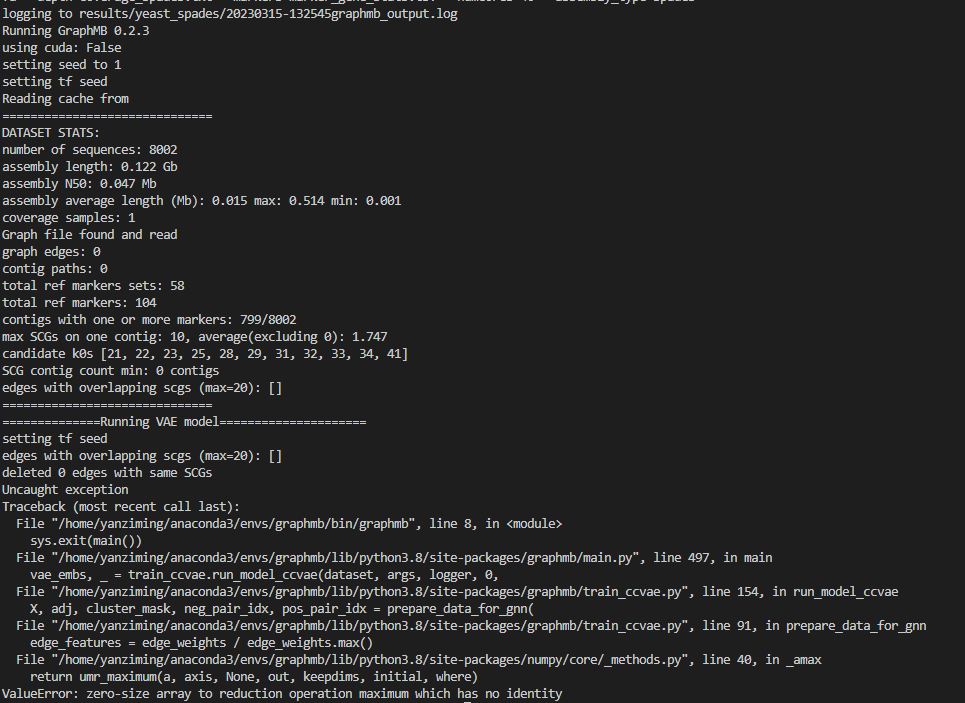
I used spades.py tool to get the following error, I do not know how to solve?
The command is as follows:
graphmb --assembly data/yeast_spades/ --outdir results/yeast_spades/ --assembly_name contigs_over_1000.fasta --graph_file strain_graph.gfa --depth coverage_spades.txt --markers marker_gene_stats.tsv --numcores 40 --assembly_type spades
I would appreciate it if you could help me.
Thank you.
Reactions are currently unavailable
Metadata
Metadata
Assignees
Labels
bugSomething isn't workingSomething isn't working