diff --git a/CHANGELOG.md b/CHANGELOG.md
index b654c49..24d9439 100644
--- a/CHANGELOG.md
+++ b/CHANGELOG.md
@@ -2,6 +2,76 @@
All notable changes to this project will be documented in this file.
+## [0.6.0] - 2026-02-26
+
+### Breaking changes
+- **Removed `set_backend()`, `get_backend_info()`, `reset_backend()`** — only one backend (C++ native) exists since v0.5.0, so the multi-backend API was dead code. Use `from mcpower.backends import get_backend` if you need the backend instance directly
+- **Removed `set_heterogeneity()` and `set_heteroskedasticity()`** — heterogeneity and heteroskedasticity are now controlled exclusively through scenario configurations (`set_scenario_configs()`). The optimistic scenario uses zero perturbation; realistic/doomer scenarios apply these automatically
+- **Removed dead scipy fallback code** from `distributions.py` — scipy was never a runtime dependency since v0.5.0, so the fallback paths were unreachable dead code. The module now cleanly fails with an `ImportError` if the C++ native backend is missing
+- **`_create_power_plot()` returns `fig`** — the function now accepts a `show=True` parameter and always returns the matplotlib figure object. Set `show=False` to suppress `plt.show()` for programmatic use
+- **`apply()` made private (`_apply()`)** — the method is now `_apply()` and called automatically by `find_power()` / `find_sample_size()`. Direct calls should use `model._apply()` instead
+- **`[all]` extra no longer includes `statsmodels`** — use `pip install mcpower[lme]` to get statsmodels for mixed-effects models
+
+### Added
+- **`test_formula` parameter** on `find_power()` and `find_sample_size()` — test a reduced model against data generated from the full model to evaluate power under model misspecification. For example, generate data with `y = x1 + x2 + x3` but test with `test_formula="y ~ x1 + x2"` to see power when `x3` is omitted. Supports interactions, factors, and mixed models.
+- **C++ non-normal residual generation** — scenario perturbations now generate heavy-tailed (Student-t) and skewed (chi-squared) residuals directly in C++ via `residual_dist`/`residual_df` parameters in `generate_y()`, replacing the Python-side post-hoc perturbation approach. Applies to all model types (OLS and LME)
+- **`optimistic` scenario** is now a first-class entry in `DEFAULT_SCENARIO_CONFIG` with all-zero perturbation values, eliminating the special `scenario_config=None` code path. Custom scenarios inherit from the optimistic baseline, ensuring all required keys exist
+
+### Fixed
+- **`set_variable_type()` docstring listed wrong distribution types** — documented non-existent `"skewed"` type; now lists all supported types: `right_skewed`, `left_skewed`, `high_kurtosis`, `uniform`
+- **`set_scenario_configs()` docstring referenced non-existent keys** — `"effect_size_jitter"` and `"distribution_jitter"` replaced with actual keys (`correlation_noise_sd`, `distribution_change_prob`, etc.)
+- **String factor levels crash in LME variance computation** — `proportions[level - 1]` crashed when factor levels were strings (e.g. `"Japan"`). Now looks up level position in the label list
+- **Division by zero on constant-variance columns** — `upload_data()` normalization produced `inf`/`NaN` when a column had zero variance. Now raises `ValueError` with the column name
+- **Pending state not cleared after `_apply()`** — calling `_apply()` twice could re-apply the same effects. Pending fields are now reset after each `_apply()` call
+- **Parser crash on unbalanced parentheses** — unmatched `)` caused `paren_count` to go negative, producing silent misparses. Now raises `ValueError`
+- **Update checker wrote cache inside installed package** — moved cache file to `~/.cache/mcpower/update_cache.json`
+- **Update checker unbounded response read** — `response.read()` now limited to 1 MB
+- **`scenario_config` dict access on `None`** — added `None` guards for optional scenario configuration lookups
+- **NaN values in uploaded data** — `upload_data()` now rejects data containing NaN values with a clear error message listing affected columns
+- **Formula minus-sign silently dropped terms** — `y = x1 - x2` silently ignored `x2`. Now raises `ValueError` explaining that term removal with `-` is not supported
+- **`_create_table` crash on empty rows** — formatter now handles empty row lists by computing column widths from headers only
+- **`_create_power_plot` crash when `first_achieved` not in sample sizes** — added bounds check before `.index()` call
+- **Redundant `_validate_cluster_sample_size` call** — removed duplicate validation in `find_power()` (already called per-sample-size in `find_sample_size()`)
+
+### Changed
+- **`upload_data()` returns `self`** for method chaining consistency
+- **Assert statements replaced with `RuntimeError`** — internal assertions now raise proper exceptions instead of using `assert`
+- **Removed "(not yet implemented)" from mixed-model docstrings** — mixed model testing has been implemented since v0.4.2
+- **Thread-safe RNG in data generation** — replaced global `np.random.seed()` with local `np.random.RandomState()` for thread safety
+- **Update checker runs in a background thread** — no longer blocks `import mcpower` on slow networks
+- **Module-level deduplication for update checker** — prevents redundant version checks within the same Python session
+- **Removed unused `cluster_column_indices` parameter** from `_lme_analysis_wrapper()` and `_lme_analysis_statsmodels()` — was explicitly marked unused and kept only for API compatibility
+- **Scenario formatters iterate dynamically** — no longer hardcode scenario names, enabling custom scenario display
+
+### Packaging
+- **`tqdm` added as core dependency** (`>=4.60.0`) — used for progress bars
+- **Removed stale pytest warning filter** for `"Mixed-effects models are experimental"` (warning was removed in v0.5.4)
+- **NumPy minimum version relaxed** to `>=1.26.0` (was `>=2.0.0`) in both build-requires and runtime dependencies
+- **`scikit-build-core` bumped** to `>=0.10` (was `>=0.5`)
+- **`statsmodels` added to `[dev]` extras** for test/development convenience
+- **Documentation URL** now points to the GitHub wiki
+- **Changelog URL** added to project URLs
+- **Removed unused pytest markers** (`unit`, `integration`) — only `lme` marker remains
+- **Per-module mypy overrides** replace blanket `ignore_missing_imports`
+
+### Documentation
+- Updated README requirements section: added `tqdm`, specified `NumPy (>=1.26.0)`
+- Changed `pip install mcpower[all]` → `pip install mcpower[lme]` for statsmodels installation
+- Wiki documentation review and cleanup: fixed broken links, corrected API signatures (`set_scenario_configs` parameter name), removed stale `apply()` and `set_heterogeneity()` wiki pages, fixed formula redundancy in Model Specification, corrected Tukey return value docs, added mixed-model caveats
+
+### Technical
+- Removed ~150 lines of dead scipy fallback shims from `distributions.py`
+- Removed `_BACKEND` sentinel variable (only one backend exists)
+- C++ `generate_y()` now accepts `residual_dist` and `residual_df` parameters for non-normal error generation
+- `suppress_output` test fixture now actually suppresses stdout (was a no-op)
+- Removed unused `correlation_matrix_3x3` test fixture
+- Removed empty `tests/mcpower/` artifact directory
+- Added unit tests for `ResultsProcessor` (`test_results.py`)
+- Added unit tests for `normalize_upload_input` (`test_upload_data_utils.py`)
+- Added integration tests for `test_formula` feature (`test_test_formula.py`)
+- Added unit tests for `test_formula_utils` (`test_test_formula_utils.py`)
+- Rewrote optimizer tests to test native backend directly (removed dead scipy fallback tests)
+
## [0.5.4] - 2026-02-22
### Changed
diff --git a/README.md b/README.md
index a230021..2bd003a 100644
--- a/README.md
+++ b/README.md
@@ -21,6 +21,10 @@
It's a Python package, but prefer a graphical interface? **[MCPower GUI](https://github.com/pawlenartowicz/mcpower-gui)** is a standalone desktop app — no Python installation required. Download ready-to-run executables for Windows, Linux, and macOS from the [releases page](https://github.com/pawlenartowicz/mcpower-gui/releases/latest).
+| Model setup | Results |
+|:---:|:---:|
+| 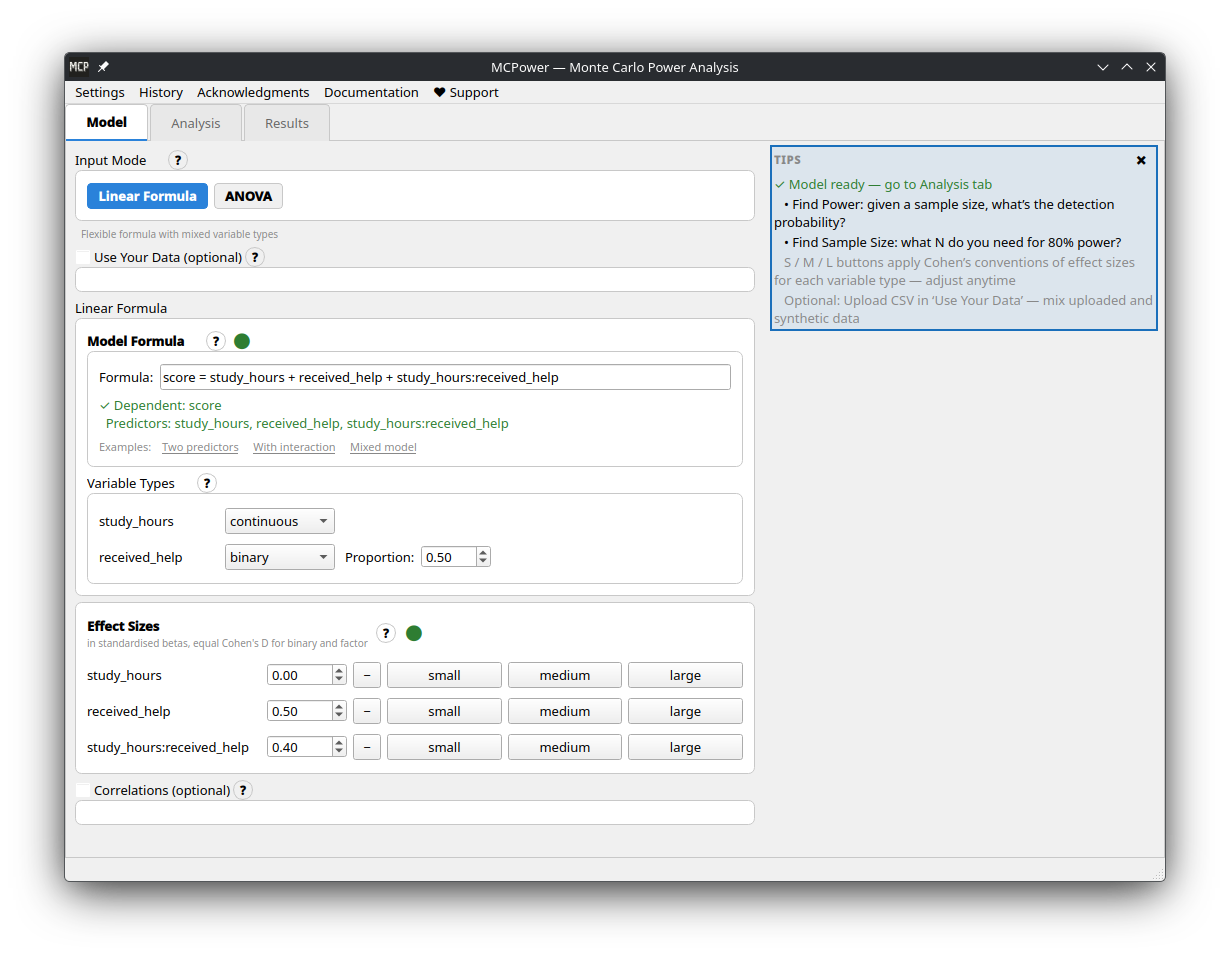 |
| 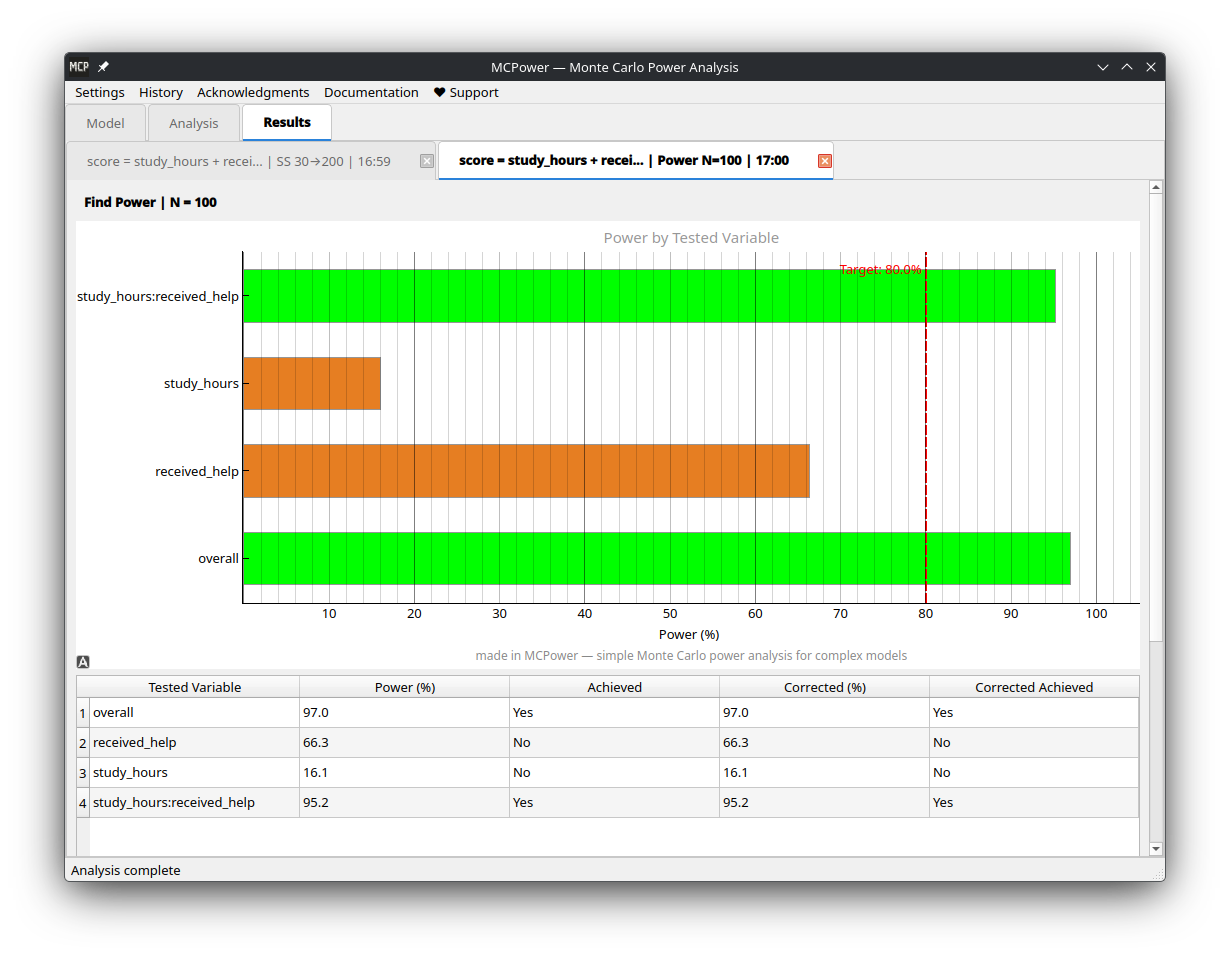 |
+
## Why MCPower?
Traditional power formulas break down with interactions, correlated predictors, categorical variables, or non-normal data. MCPower simulates instead — generates thousands of datasets like yours, fits your model, and counts how often the effects are detected.
@@ -297,19 +301,20 @@ model.set_effects("group[2]=0.4, group[3]=0.6, covariate=0.3")
# Use "vs" syntax for pairwise comparisons + correction="tukey"
model.find_power(
sample_size=150,
- target_test="group[0] vs group[1], group[0] vs group[2]",
+ target_test="group[1] vs group[2], group[1] vs group[3]",
correction="tukey"
)
```
### Test Individual Assumption Violations
```python
-# Manually add specific violations (without full scenario analysis)
-model.set_heterogeneity(0.2) # Effect sizes vary between people
-model.set_heteroskedasticity(0.15) # Violation of equal variance assumption
+# Add specific violations via custom scenario configs
+model.set_scenario_configs({
+ "my_test": {"heterogeneity": 0.2, "heteroskedasticity": 0.15}
+})
-# Run with your manual settings (no automatic scenario variations)
-model.find_sample_size(target_test="treatment")
+# Run with scenario variations
+model.find_sample_size(target_test="treatment", scenarios=True)
```
### Mixed-Effects Models
@@ -392,7 +397,7 @@ model.find_power(sample_size=200, progress_callback=False)
| **Factor effects** | **`model.set_effects("var[2]=0.5, var[3]=0.7")`** |
| Correlated predictors | `model.set_correlations("corr(var1, var2)=0.4")` |
| Multiple testing correction | Add `correction="FDR"`, `"Holm"`, `"Bonferroni"`, or `"Tukey"`|
-| Post-hoc pairwise comparison | `target_test="group[0] vs group[1]"` with `correction="tukey"` |
+| Post-hoc pairwise comparison | `target_test="group[1] vs group[2]"` with `correction="tukey"` |
| Mixed model (random intercept) | `MCPower("y ~ x + (1\|group)")` + `model.set_cluster(...)` |
| Random slopes | `MCPower("y ~ x + (1+x\|group)")` + `set_cluster(..., random_slopes=["x"], slope_variance=0.1)` |
| Nested random effects | `MCPower("y ~ x + (1\|A/B)")` + two `set_cluster()` calls |
@@ -424,7 +429,7 @@ model.find_power(sample_size=200, progress_callback=False)
- For simple models where all assumptions are clearly met.
- For large analyses with tens of thousands of observations, tiny effects, or very low alpha levels.
-## What Makes Scenarios Different? (Be careful, unvalidated, preliminary scenarios)
+## What Makes Scenarios Different? (Rule-of-thumb scenarios)
**Traditional power analysis assumes perfect conditions.** MCPower's scenarios add realistic "messiness":
@@ -478,8 +483,8 @@ model.set_variable_type("treatment=(factor,3), education=(factor,4)")
# Set effects for specific levels
model.set_effects("treatment[2]=0.5, treatment[3]=0.7, education[2]=0.3")
-# Or set same effect for all levels of a factor
-model.set_effects("treatment=0.5") # Applies to treatment[2] and treatment[3]
+# Each non-reference level needs its own effect
+model.set_effects("treatment[2]=0.5, treatment[3]=0.7")
# Important: Factors cannot be used in correlations
# This will error: model.set_correlations("corr(treatment, education)=0.3")
@@ -508,12 +513,31 @@ model.set_alpha(0.01) # Stricter significance (p < 0.01)
model.set_simulations(10000) # High precision (slower)
```
+### Model Misspecification Testing
+
+Use `test_formula` to generate data with one model but test with a simpler one -- useful for evaluating the power impact of omitting variables:
+
+```python
+# Generate with 3 predictors, test with 2 (omitting x3)
+model = MCPower("y = x1 + x2 + x3")
+model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+model.find_power(100, test_formula="y = x1 + x2")
+
+# Generate with clusters, test without (ignoring clustering)
+model = MCPower("y ~ treatment + (1|school)")
+model.set_cluster("school", ICC=0.2, n_clusters=20)
+model.set_effects("treatment=0.5")
+model.find_power(1000, test_formula="y ~ treatment")
+```
+
+See the [Test Formula Tutorial](https://github.com/pawlenartowicz/MCPower/wiki/Tutorial-Test-Formula) for details.
+
### Formula Syntax
```python
# These are equivalent:
-"y = x1 + x2 + x1*x2" # Assignment style
-"y ~ x1 + x2 + x1*x2" # R-style formula
-"x1 + x2 + x1*x2" # Predictors only
+"y = x1 + x2 + x1:x2" # Assignment style
+"y ~ x1 + x2 + x1:x2" # R-style formula
+"x1 + x2 + x1:x2" # Predictors only
# Interactions:
"x1*x2" # Main effects + interaction (x1 + x2 + x1:x2)
@@ -538,9 +562,8 @@ model.set_correlations("(x1, x2)=0.3, (x1, x3)=-0.2")
## Requirements
- Python ≥ 3.10
-- NumPy, matplotlib, joblib
+- NumPy (≥1.26.0), matplotlib, joblib, tqdm
- pandas (optional, for DataFrame input — install with `pip install mcpower[pandas]`)
-- statsmodels (optional, for mixed-effects models — install with `pip install mcpower[all]`)
## Documentation
@@ -549,11 +572,11 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
- [Quick Start](https://github.com/pawlenartowicz/MCPower/wiki/Quick-Start)
- [Model Specification](https://github.com/pawlenartowicz/MCPower/wiki/Model-Specification)
-- [Variable Types](https://github.com/pawlenartowicz/MCPower/wiki/Variable-Types)
-- [Effect Sizes](https://github.com/pawlenartowicz/MCPower/wiki/Effect-Sizes)
-- [Mixed-Effects Models](https://github.com/pawlenartowicz/MCPower/wiki/Mixed-Effects-Models) (random intercepts, slopes, nested effects)
-- [ANOVA & Post-Hoc Tests](https://github.com/pawlenartowicz/MCPower/wiki/ANOVA-and-Post-Hoc-Tests)
-- [Scenario Analysis](https://github.com/pawlenartowicz/MCPower/wiki/Scenario-Analysis)
+- [Variable Types](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Variable-Types)
+- [Effect Sizes](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Effect-Sizes)
+- [Mixed-Effects Models](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Mixed-Effects) (random intercepts, slopes, nested effects)
+- [ANOVA & Post-Hoc Tests](https://github.com/pawlenartowicz/MCPower/wiki/Tutorial-ANOVA-PostHoc)
+- [Scenario Analysis](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Scenario-Analysis)
- [API Reference](https://github.com/pawlenartowicz/MCPower/wiki/API-Reference)
## Need Help?
@@ -568,8 +591,8 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
- ✅ C++ native backend (pybind11 + Eigen, 3x speedup)
- ✅ Mixed Effects Models (random intercepts, random slopes, nested effects) — [validated against lme4](https://github.com/pawlenartowicz/MCPower/wiki/Concept-LME-Validation)
- 🚧 Logistic Regression (coming soon)
-- 🚧 ANOVA (coming soon)
-- 🚧 Guide about methods, corrections (coming soon)
+- ✅ ANOVA (factor variables as ANOVA, post-hoc pairwise comparisons)
+- ✅ Guide about methods, corrections
- 📋 2 groups comparison with alternative tests
- 📋 Robust regression methods
@@ -578,16 +601,18 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
GPL v3. If you use MCPower in research, please cite:
-Lenartowicz, P. (2025). MCPower: Monte Carlo Power Analysis for Statistical Models. Zenodo. DOI: 10.5281/zenodo.16502734
+Lenartowicz, P. (2025). MCPower: Monte Carlo Power Analysis for Complex Statistical Models (Version ) [Computer software]. Zenodo. https://doi.org/10.5281/zenodo.16502734
+
+*Replace `` with the version you used — check with `import mcpower; print(mcpower.__version__)`.*
```bibtex
@software{mcpower2025,
- author = {Pawel Lenartowicz},
- title = {MCPower: Monte Carlo Power Analysis for Statistical Models},
- year = {2025},
+ author = {Lenartowicz, Pawe{\l}},
+ title = {{MCPower}: Monte Carlo Power Analysis for Complex Statistical Models},
+ year = {2025},
publisher = {Zenodo},
- doi = {10.5281/zenodo.16502734},
- url = {https://doi.org/10.5281/zenodo.16502734}
+ doi = {10.5281/zenodo.16502734},
+ url = {https://doi.org/10.5281/zenodo.16502734}
}
```
diff --git a/cpp/src/bindings.cpp b/cpp/src/bindings.cpp
index 26fee22..8c02998 100644
--- a/cpp/src/bindings.cpp
+++ b/cpp/src/bindings.cpp
@@ -110,7 +110,9 @@ py::array_t generate_y_wrapper(
py::array_t effects,
double heterogeneity,
double heteroskedasticity,
- int seed
+ int seed,
+ int residual_dist,
+ double residual_df
) {
auto X_buf = X.request();
auto effects_buf = effects.request();
@@ -129,7 +131,8 @@ py::array_t generate_y_wrapper(
);
Eigen::VectorXd y = generate_y(
- X_map, effects_map, heterogeneity, heteroskedasticity, seed
+ X_map, effects_map, heterogeneity, heteroskedasticity, seed,
+ residual_dist, residual_df
);
py::array_t result(n);
@@ -447,7 +450,9 @@ PYBIND11_MODULE(mcpower_native, m) {
py::arg("heterogeneity") = 0.0,
py::arg("heteroskedasticity") = 0.0,
py::arg("seed") = -1,
- "Generate dependent variable with heterogeneity and heteroskedasticity"
+ py::arg("residual_dist") = 0,
+ py::arg("residual_df") = 10.0,
+ "Generate dependent variable with heterogeneity, heteroskedasticity, and non-normal residuals"
);
// LME analysis (q=1 random intercept)
diff --git a/cpp/src/ols.cpp b/cpp/src/ols.cpp
index 7d04ec2..11bd62f 100644
--- a/cpp/src/ols.cpp
+++ b/cpp/src/ols.cpp
@@ -151,19 +151,14 @@ Eigen::VectorXd generate_y(
const Eigen::Ref& effects,
double heterogeneity,
double heteroskedasticity,
- int seed
+ int seed,
+ int residual_dist,
+ double residual_df
) {
const int n = static_cast(X.rows());
const int p = static_cast(X.cols());
- // Set up random generator
std::mt19937 gen;
- if (seed >= 0) {
- gen.seed(static_cast(seed));
- } else {
- std::random_device rd;
- gen.seed(rd());
- }
std::normal_distribution normal(0.0, 1.0);
// Linear predictor with heterogeneity
@@ -176,9 +171,12 @@ Eigen::VectorXd generate_y(
// Heterogeneity: vary effect sizes per observation
linear_pred.setZero();
- // Change seed for heterogeneity noise
+ // Seed at offset +1 for heterogeneity noise
if (seed >= 0) {
gen.seed(static_cast(seed + 1));
+ } else {
+ std::random_device rd;
+ gen.seed(rd());
}
for (int j = 0; j < p; ++j) {
@@ -192,14 +190,43 @@ Eigen::VectorXd generate_y(
}
}
- // Generate errors
+ // Generate errors — seed at offset +2
if (seed >= 0) {
gen.seed(static_cast(seed + 2));
+ } else {
+ std::random_device rd;
+ gen.seed(rd());
}
Eigen::VectorXd error(n);
- for (int i = 0; i < n; ++i) {
- error(i) = normal(gen);
+
+ if (residual_dist == 1) {
+ // Heavy-tailed: Student's t distribution
+ double df = std::max(residual_df, 3.0);
+ std::student_t_distribution t_dist(df);
+ double theoretical_scale = 1.0 / std::sqrt(df / (df - 2.0));
+ for (int i = 0; i < n; ++i) {
+ error(i) = t_dist(gen) * theoretical_scale;
+ }
+ } else if (residual_dist == 2) {
+ // Skewed: chi-squared, centered and scaled
+ double df = std::max(residual_df, 3.0);
+ std::chi_squared_distribution chi2_dist(df);
+ double scale = 1.0 / std::sqrt(2.0 * df);
+ for (int i = 0; i < n; ++i) {
+ error(i) = (chi2_dist(gen) - df) * scale;
+ }
+ } else {
+ // Normal (default)
+ for (int i = 0; i < n; ++i) {
+ error(i) = normal(gen);
+ }
+ }
+
+ // Empirical re-standardization to SD = 1
+ double empirical_sd = std::sqrt(error.array().square().mean());
+ if (empirical_sd > FLOAT_NEAR_ZERO) {
+ error /= empirical_sd;
}
// Apply heteroskedasticity
diff --git a/cpp/src/ols.hpp b/cpp/src/ols.hpp
index ad1f9b6..9e046eb 100644
--- a/cpp/src/ols.hpp
+++ b/cpp/src/ols.hpp
@@ -65,6 +65,8 @@ class OLSAnalyzer {
* @param heterogeneity SD of effect size variation
* @param heteroskedasticity Correlation between predictor and error variance
* @param seed Random seed (-1 for random)
+ * @param residual_dist Error distribution: 0=normal, 1=heavy_tailed (t), 2=skewed (chi2)
+ * @param residual_df Degrees of freedom for non-normal residuals (min clamped to 3)
* @return Response vector (n_samples,)
*/
Eigen::VectorXd generate_y(
@@ -72,7 +74,9 @@ Eigen::VectorXd generate_y(
const Eigen::Ref& effects,
double heterogeneity,
double heteroskedasticity,
- int seed
+ int seed,
+ int residual_dist = 0,
+ double residual_df = 10.0
);
} // namespace mcpower
diff --git a/docs/screenshots/gui-model-setup.png b/docs/screenshots/gui-model-setup.png
new file mode 100644
index 0000000..7f87a53
Binary files /dev/null and b/docs/screenshots/gui-model-setup.png differ
diff --git a/docs/screenshots/gui-results.png b/docs/screenshots/gui-results.png
new file mode 100644
index 0000000..f84152d
Binary files /dev/null and b/docs/screenshots/gui-results.png differ
diff --git a/mcpower/__init__.py b/mcpower/__init__.py
index a52560c..675a4f3 100644
--- a/mcpower/__init__.py
+++ b/mcpower/__init__.py
@@ -16,7 +16,6 @@
from importlib.metadata import version as _get_version
-from .backends import get_backend_info, set_backend
from .model import MCPower
from .progress import PrintReporter, ProgressReporter, SimulationCancelled, TqdmReporter
@@ -27,14 +26,14 @@
__all__ = [
"MCPower",
"SimulationCancelled",
- "set_backend",
- "get_backend_info",
"ProgressReporter",
"PrintReporter",
"TqdmReporter",
]
+import threading as _threading
+
from .utils.updates import _check_for_updates
-_check_for_updates(__version__)
+_threading.Thread(target=_check_for_updates, args=(__version__,), daemon=True).start()
diff --git a/mcpower/backends/__init__.py b/mcpower/backends/__init__.py
index 7bb03f8..8b24a73 100644
--- a/mcpower/backends/__init__.py
+++ b/mcpower/backends/__init__.py
@@ -3,11 +3,9 @@
This module provides a unified interface for compute backends.
The only supported backend is native C++ (compiled via pybind11).
-
-Users can override via set_backend('c++' | 'default') or pass a ComputeBackend instance.
"""
-from typing import Optional, Protocol, Union, runtime_checkable
+from typing import Optional, Protocol, runtime_checkable
import numpy as np
@@ -24,6 +22,7 @@ def ols_analysis(
f_crit: float,
t_crit: float,
correction_t_crits: np.ndarray,
+ # correction_method encoding: 0=none, 1=Bonferroni, 2=FDR (BH), 3=Holm
correction_method: int,
) -> np.ndarray:
"""Run OLS regression and return significance flags.
@@ -40,9 +39,15 @@ def generate_y(
heterogeneity: float,
heteroskedasticity: float,
seed: int,
+ residual_dist: int = 0,
+ residual_df: float = 10.0,
) -> np.ndarray:
"""Generate the dependent variable ``y = X @ effects + error``.
+ Args:
+ residual_dist: Error distribution (0=normal, 1=heavy_tailed, 2=skewed).
+ residual_df: Degrees of freedom for non-normal residuals.
+
Returns:
1-D array of length ``n_samples``.
"""
@@ -88,12 +93,8 @@ def lme_analysis(
...
-# Valid backend names for set_backend()
-_BACKEND_NAMES = {"default", "c++"}
-
# Global backend instance
_backend_instance: Optional[ComputeBackend] = None
-_backend_forced = False
def get_backend() -> ComputeBackend:
@@ -101,7 +102,7 @@ def get_backend() -> ComputeBackend:
Get the active compute backend.
On first call, instantiates the C++ native backend.
- Subsequent calls return the cached instance unless reset_backend() is called.
+ Subsequent calls return the cached instance.
Raises:
ImportError: If the C++ extension is not compiled/installed.
@@ -117,64 +118,7 @@ def get_backend() -> ComputeBackend:
return _backend_instance
-def set_backend(backend: Union[str, ComputeBackend]) -> None:
- """
- Set the compute backend.
-
- Args:
- backend: One of:
- - 'default' -- use native C++ backend
- - 'c++' -- force native C++ backend
- - A ComputeBackend instance
-
- Raises:
- ImportError: If the C++ backend is not available.
- ValueError: If the string is not recognized.
- """
- global _backend_instance, _backend_forced
-
- if isinstance(backend, str):
- name = backend.lower().strip()
- if name not in _BACKEND_NAMES:
- raise ValueError(f"Unknown backend {backend!r}. Choose from: {', '.join(sorted(_BACKEND_NAMES))}")
-
- from .native import NativeBackend
-
- _backend_instance = NativeBackend()
- _backend_forced = name != "default"
- else:
- _backend_instance = backend
- _backend_forced = True
-
-
-def reset_backend() -> None:
- """Reset backend to automatic selection."""
- global _backend_instance, _backend_forced
- _backend_instance = None
- _backend_forced = False
-
-
-def get_backend_info() -> dict:
- """
- Get information about the current backend.
-
- Returns:
- Dictionary with backend name, type, and whether it was forced.
- """
- backend = get_backend()
- name = type(backend).__name__
- return {
- "name": name,
- "is_native": name == "NativeBackend",
- "module": type(backend).__module__,
- "forced": _backend_forced,
- }
-
-
__all__ = [
"ComputeBackend",
"get_backend",
- "set_backend",
- "reset_backend",
- "get_backend_info",
]
diff --git a/mcpower/backends/native.py b/mcpower/backends/native.py
index acd7633..2338cfe 100644
--- a/mcpower/backends/native.py
+++ b/mcpower/backends/native.py
@@ -17,6 +17,11 @@
mcpower_native = None
+def _prep(arr: np.ndarray, dtype=np.float64) -> np.ndarray:
+ """Ensure array is contiguous with the expected dtype for C++ interop."""
+ return np.ascontiguousarray(arr, dtype=dtype)
+
+
class NativeBackend:
"""
C++ compute backend using pybind11 bindings.
@@ -46,8 +51,8 @@ def _initialize_tables(self) -> None:
t3_ppf = manager.load_t3_ppf_table()
# Ensure correct dtypes
- norm_cdf = np.ascontiguousarray(norm_cdf.astype(np.float64))
- t3_ppf = np.ascontiguousarray(t3_ppf.astype(np.float64))
+ norm_cdf = _prep(norm_cdf)
+ t3_ppf = _prep(t3_ppf)
# Initialize C++ tables (generation tables only)
mcpower_native.init_tables(norm_cdf, t3_ppf)
@@ -77,10 +82,10 @@ def ols_analysis(
Returns:
Array: [f_sig, uncorrected..., corrected...]
"""
- X = np.ascontiguousarray(X, dtype=np.float64)
- y = np.ascontiguousarray(y, dtype=np.float64)
- target_indices = np.ascontiguousarray(target_indices, dtype=np.int32)
- correction_t_crits = np.ascontiguousarray(correction_t_crits, dtype=np.float64)
+ X = _prep(X)
+ y = _prep(y)
+ target_indices = _prep(target_indices, np.int32)
+ correction_t_crits = _prep(correction_t_crits)
return mcpower_native.ols_analysis(X, y, target_indices, f_crit, t_crit, correction_t_crits, correction_method) # type: ignore[no-any-return]
@@ -91,6 +96,8 @@ def generate_y(
heterogeneity: float,
heteroskedasticity: float,
seed: int,
+ residual_dist: int = 0,
+ residual_df: float = 10.0,
) -> np.ndarray:
"""
Generate dependent variable.
@@ -101,14 +108,16 @@ def generate_y(
heterogeneity: Effect size variation SD
heteroskedasticity: Error-predictor correlation
seed: Random seed (-1 for random)
+ residual_dist: Error distribution (0=normal, 1=heavy_tailed, 2=skewed)
+ residual_df: Degrees of freedom for non-normal residuals
Returns:
Response vector (n_samples,)
"""
- X = np.ascontiguousarray(X, dtype=np.float64)
- effects = np.ascontiguousarray(effects, dtype=np.float64)
+ X = _prep(X)
+ effects = _prep(effects)
- return mcpower_native.generate_y(X, effects, heterogeneity, heteroskedasticity, seed) # type: ignore[no-any-return]
+ return mcpower_native.generate_y(X, effects, heterogeneity, heteroskedasticity, seed, residual_dist, residual_df) # type: ignore[no-any-return]
def generate_X(
self,
@@ -137,11 +146,11 @@ def generate_X(
Returns:
Design matrix (n_samples, n_vars)
"""
- correlation_matrix = np.ascontiguousarray(correlation_matrix, dtype=np.float64)
- var_types = np.ascontiguousarray(var_types, dtype=np.int32)
- var_params = np.ascontiguousarray(var_params, dtype=np.float64)
- upload_normal = np.ascontiguousarray(upload_normal, dtype=np.float64)
- upload_data = np.ascontiguousarray(upload_data, dtype=np.float64)
+ correlation_matrix = _prep(correlation_matrix)
+ var_types = _prep(var_types, np.int32)
+ var_params = _prep(var_params)
+ upload_normal = _prep(upload_normal)
+ upload_data = _prep(upload_data)
return mcpower_native.generate_X( # type: ignore[no-any-return]
n_samples,
@@ -185,12 +194,15 @@ def lme_analysis(
Returns:
Array: [f_sig, uncorrected..., corrected..., wald_flag]
or empty array on failure
+
+ wald_flag: 1.0 if the Wald test was used as fallback for the overall
+ significance test (instead of the likelihood ratio test), 0.0 otherwise.
"""
- X = np.ascontiguousarray(X, dtype=np.float64)
- y = np.ascontiguousarray(y, dtype=np.float64)
- cluster_ids = np.ascontiguousarray(cluster_ids, dtype=np.int32)
- target_indices = np.ascontiguousarray(target_indices, dtype=np.int32)
- correction_z_crits = np.ascontiguousarray(correction_z_crits, dtype=np.float64)
+ X = _prep(X)
+ y = _prep(y)
+ cluster_ids = _prep(cluster_ids, np.int32)
+ target_indices = _prep(target_indices, np.int32)
+ correction_z_crits = _prep(correction_z_crits)
return mcpower_native.lme_analysis( # type: ignore[no-any-return]
X,
@@ -240,14 +252,17 @@ def lme_analysis_general(
Returns:
Array: [f_sig, uncorrected..., corrected..., wald_flag]
or empty array on failure
+
+ wald_flag: 1.0 if the Wald test was used as fallback for the overall
+ significance test (instead of the likelihood ratio test), 0.0 otherwise.
"""
- X = np.ascontiguousarray(X, dtype=np.float64)
- y = np.ascontiguousarray(y, dtype=np.float64)
- Z = np.ascontiguousarray(Z, dtype=np.float64)
- cluster_ids = np.ascontiguousarray(cluster_ids, dtype=np.int32)
- target_indices = np.ascontiguousarray(target_indices, dtype=np.int32)
- correction_z_crits = np.ascontiguousarray(correction_z_crits, dtype=np.float64)
- warm_theta = np.ascontiguousarray(warm_theta, dtype=np.float64)
+ X = _prep(X)
+ y = _prep(y)
+ Z = _prep(Z)
+ cluster_ids = _prep(cluster_ids, np.int32)
+ target_indices = _prep(target_indices, np.int32)
+ correction_z_crits = _prep(correction_z_crits)
+ warm_theta = _prep(warm_theta)
return mcpower_native.lme_analysis_general( # type: ignore[no-any-return]
X,
@@ -301,15 +316,18 @@ def lme_analysis_nested(
Returns:
Array: [f_sig, uncorrected..., corrected..., wald_flag]
or empty array on failure
+
+ wald_flag: 1.0 if the Wald test was used as fallback for the overall
+ significance test (instead of the likelihood ratio test), 0.0 otherwise.
"""
- X = np.ascontiguousarray(X, dtype=np.float64)
- y = np.ascontiguousarray(y, dtype=np.float64)
- parent_ids = np.ascontiguousarray(parent_ids, dtype=np.int32)
- child_ids = np.ascontiguousarray(child_ids, dtype=np.int32)
- child_to_parent = np.ascontiguousarray(child_to_parent, dtype=np.int32)
- target_indices = np.ascontiguousarray(target_indices, dtype=np.int32)
- correction_z_crits = np.ascontiguousarray(correction_z_crits, dtype=np.float64)
- warm_theta = np.ascontiguousarray(warm_theta, dtype=np.float64)
+ X = _prep(X)
+ y = _prep(y)
+ parent_ids = _prep(parent_ids, np.int32)
+ child_ids = _prep(child_ids, np.int32)
+ child_to_parent = _prep(child_to_parent, np.int32)
+ target_indices = _prep(target_indices, np.int32)
+ correction_z_crits = _prep(correction_z_crits)
+ warm_theta = _prep(warm_theta)
return mcpower_native.lme_analysis_nested( # type: ignore[no-any-return]
X,
diff --git a/mcpower/core/results.py b/mcpower/core/results.py
index 45b178a..dfe6b77 100644
--- a/mcpower/core/results.py
+++ b/mcpower/core/results.py
@@ -54,6 +54,7 @@ def calculate_powers(
# Individual powers
individual_powers = {}
individual_powers_corrected = {}
+ non_overall_tests = [t for t in target_tests if t != "overall"]
for test in target_tests:
if test == "overall":
@@ -62,7 +63,6 @@ def calculate_powers(
individual_powers_corrected[test] = np.mean(results_corrected_array[:, 0]) * 100
else:
# Find position among non-'overall' tests and add 1 for F-test offset
- non_overall_tests = [t for t in target_tests if t != "overall"]
pos = non_overall_tests.index(test)
col_idx = pos + 1 # +1 because column 0 is F-test
individual_powers[test] = np.mean(results_array[:, col_idx]) * 100
diff --git a/mcpower/core/scenarios.py b/mcpower/core/scenarios.py
index 454f8e3..2d2dd01 100644
--- a/mcpower/core/scenarios.py
+++ b/mcpower/core/scenarios.py
@@ -13,35 +13,51 @@
from ..utils.visualization import _create_power_plot
# Default scenario configurations.
+# "optimistic" is the zero-perturbation baseline — also used as the default
+# scenario_config when scenarios=False and as a template for custom scenarios
+# (ensures all required keys exist).
# "realistic" introduces moderate assumption violations; "doomer" introduces
# severe violations. Each simulation iteration draws random perturbations
# from these parameters (correlation noise, distribution swaps, etc.).
DEFAULT_SCENARIO_CONFIG = {
+ "optimistic": {
+ "heterogeneity": 0.0,
+ "heteroskedasticity": 0.0,
+ "correlation_noise_sd": 0.0,
+ "distribution_change_prob": 0.0,
+ "new_distributions": ["right_skewed", "left_skewed", "uniform"],
+ # Mixed model perturbations (only consumed when cluster_specs present)
+ "random_effect_dist": "normal",
+ "random_effect_df": 5,
+ "icc_noise_sd": 0.0,
+ # Residual distribution perturbations (all model types)
+ "residual_dists": ["heavy_tailed", "skewed"],
+ "residual_change_prob": 0.0,
+ "residual_df": 10,
+ },
"realistic": {
"heterogeneity": 0.2,
- "heteroskedasticity": 0.1,
- "correlation_noise_sd": 0.2,
- "distribution_change_prob": 0.3,
+ "heteroskedasticity": 0.15,
+ "correlation_noise_sd": 0.15,
+ "distribution_change_prob": 0.5,
"new_distributions": ["right_skewed", "left_skewed", "uniform"],
- # LME-specific keys (only consumed when cluster_specs present)
"random_effect_dist": "heavy_tailed",
- "random_effect_df": 5,
+ "random_effect_df": 10,
"icc_noise_sd": 0.15,
- "residual_dist": "heavy_tailed",
- "residual_change_prob": 0.3,
- "residual_df": 10,
+ "residual_dists": ["heavy_tailed", "skewed"],
+ "residual_change_prob": 0.5,
+ "residual_df": 8,
},
"doomer": {
"heterogeneity": 0.4,
- "heteroskedasticity": 0.2,
- "correlation_noise_sd": 0.4,
- "distribution_change_prob": 0.6,
+ "heteroskedasticity": 0.35,
+ "correlation_noise_sd": 0.30,
+ "distribution_change_prob": 0.8,
"new_distributions": ["right_skewed", "left_skewed", "uniform"],
- # LME-specific keys (only consumed when cluster_specs present)
"random_effect_dist": "heavy_tailed",
- "random_effect_df": 3,
+ "random_effect_df": 5,
"icc_noise_sd": 0.30,
- "residual_dist": "heavy_tailed",
+ "residual_dists": ["heavy_tailed", "skewed"],
"residual_change_prob": 0.8,
"residual_df": 5,
},
@@ -111,15 +127,7 @@ def run_power_analysis(
if progress is not None:
progress.start()
- # Optimistic (user's original settings)
- results["optimistic"] = run_find_power_func(
- sample_size=sample_size,
- target_tests=target_tests,
- correction=correction,
- scenario_config=None,
- )
-
- # Realistic & Doomer scenarios
+ # Run all scenarios (optimistic is always present as zero-perturbation baseline)
for scenario_name, config in self.configs.items():
results[scenario_name] = run_find_power_func(
sample_size=sample_size,
@@ -175,15 +183,7 @@ def run_sample_size_analysis(
if progress is not None:
progress.start()
- # Optimistic
- results["optimistic"] = run_sample_size_func(
- sample_sizes=sample_sizes,
- target_tests=target_tests,
- correction=correction,
- scenario_config=None,
- )
-
- # Other scenarios
+ # Run all scenarios (optimistic is always present as zero-perturbation baseline)
for scenario_name, config in self.configs.items():
results[scenario_name] = run_sample_size_func(
sample_sizes=sample_sizes,
@@ -209,8 +209,9 @@ def run_sample_size_analysis(
def _create_scenario_plots(self, results: Dict) -> None:
"""Create visualizations for scenario analysis."""
scenarios = results["scenarios"]
- scenario_names = ["optimistic", "realistic", "doomer"]
- scenario_labels = ["Optimistic", "Realistic", "Doomer"]
+ # Derive scenario order from results: optimistic first, then config keys
+ scenario_names = ["optimistic"] + [k for k in scenarios if k != "optimistic"]
+ scenario_labels = [name.title() for name in scenario_names]
first_scenario = scenarios.get("optimistic", {})
if "results" not in first_scenario or "sample_sizes_tested" not in first_scenario["results"]:
@@ -286,7 +287,7 @@ def apply_lme_perturbations(
if icc_noise_sd == 0.0 and re_dist == "normal":
return None
- rng = np.random.RandomState(sim_seed + 5000 if sim_seed is not None else None)
+ rng = np.random.RandomState(sim_seed + 6 if sim_seed is not None else None)
# ICC jitter: multiplicative noise on tau_squared per grouping variable
tau_squared_multipliers: Dict[str, float] = {}
@@ -304,70 +305,6 @@ def apply_lme_perturbations(
}
-def apply_lme_residual_perturbations(
- y: np.ndarray,
- scenario_config: Dict,
- sim_seed: Optional[int],
-) -> np.ndarray:
- """Replace normal residuals with non-normal if coin flip succeeds.
-
- For each simulation, independently flips a coin (probability
- ``residual_change_prob``) to decide whether residuals are replaced.
- If activated, reproduces the original N(0,1) errors via the known
- seed, generates replacements from t(df) or shifted χ², and applies
- the correction ``y += (new_error - original_error)``.
-
- Args:
- y: Dependent variable array (modified in-place).
- scenario_config: Scenario parameters with residual keys.
- sim_seed: Random seed for reproducibility.
-
- Returns:
- The (possibly modified) dependent variable array.
- """
- residual_dist = scenario_config.get("residual_dist", "normal")
- residual_change_prob = scenario_config.get("residual_change_prob", 0.0)
- residual_df = scenario_config.get("residual_df", 10)
-
- if residual_dist == "normal" or residual_change_prob <= 0.0:
- return y
-
- rng = np.random.RandomState(sim_seed + 6000 if sim_seed is not None else None)
-
- # Coin flip: should this simulation have non-normal residuals?
- if rng.random() > residual_change_prob:
- return y
-
- n = len(y)
-
- # Reproduce the original N(0,1) errors using the same seed as generate_y
- # generate_y uses sim_seed + 2 for error generation
- original_rng = np.random.RandomState(sim_seed + 2 if sim_seed is not None else None)
- original_errors = original_rng.standard_normal(n)
-
- # Generate replacement errors
- replacement_rng = np.random.RandomState(sim_seed + 6001 if sim_seed is not None else None)
-
- if residual_dist == "heavy_tailed":
- # t(df) scaled to have variance 1
- df = max(residual_df, 3)
- raw = replacement_rng.standard_t(df, size=n)
- # t(df) has variance df/(df-2), scale to unit variance
- scale = 1.0 / np.sqrt(df / (df - 2))
- new_errors = raw * scale
- elif residual_dist == "skewed":
- # Shifted chi-squared: mean=0, variance=1
- df = max(residual_df, 3)
- raw = replacement_rng.chisquare(df, size=n)
- new_errors = (raw - df) / np.sqrt(2 * df)
- else:
- return y
-
- # Apply correction: swap out original errors for new ones
- y = y + (new_errors - original_errors)
- return y
-
-
def apply_per_simulation_perturbations(
correlation_matrix: np.ndarray,
var_types: np.ndarray,
@@ -393,19 +330,22 @@ def apply_per_simulation_perturbations(
if scenario_config is None:
return correlation_matrix, var_types
- rng = np.random.RandomState(sim_seed)
+ rng = np.random.RandomState(sim_seed + 5 if sim_seed is not None else None)
# Perturb correlation matrix
perturbed_corr = correlation_matrix
- if correlation_matrix is not None and scenario_config["correlation_noise_sd"] > 0:
+ if correlation_matrix is not None and scenario_config.get("correlation_noise_sd", 0) > 0:
perturbed_corr = correlation_matrix.copy()
noise = rng.normal(0, scenario_config["correlation_noise_sd"], correlation_matrix.shape)
noise = (noise + noise.T) / 2 # Keep symmetric
perturbed_corr += noise
+ # Clip off-diagonal correlations to [-0.8, 0.8] to prevent near-singular
+ # matrices that cause Cholesky decomposition failures in data generation.
perturbed_corr = np.clip(perturbed_corr, -0.8, 0.8)
np.fill_diagonal(perturbed_corr, 1.0)
- # Ensure positive semi-definiteness via eigenvalue clipping
+ # Nearest correlation matrix repair via spectral clipping: set negative
+ # eigenvalues to zero and reconstruct, then re-normalize to unit diagonal.
eigvals, eigvecs = np.linalg.eigh(perturbed_corr)
if np.any(eigvals < 0):
eigvals = np.maximum(eigvals, 0.0)
@@ -417,7 +357,7 @@ def apply_per_simulation_perturbations(
# Perturb variable types
perturbed_var_types = var_types.copy()
- if scenario_config["distribution_change_prob"] > 0:
+ if scenario_config.get("distribution_change_prob", 0) > 0:
type_mapping = {"right_skewed": 2, "left_skewed": 3, "uniform": 5}
new_type_codes = [type_mapping[distribution] for distribution in scenario_config["new_distributions"]]
diff --git a/mcpower/core/simulation.py b/mcpower/core/simulation.py
index 266a39e..2223324 100644
--- a/mcpower/core/simulation.py
+++ b/mcpower/core/simulation.py
@@ -61,7 +61,7 @@ def __init__(
Args:
n_simulations: Number of Monte Carlo iterations.
seed: Base random seed. Each iteration uses
- ``seed + 4 * sim_id``.
+ ``seed + 12 * sim_id``.
alpha: Significance level for hypothesis tests.
parallel: Parallel processing mode (unused inside the
runner itself; parallelism is handled at the
@@ -143,12 +143,19 @@ def run_power_simulations(
if metadata.cluster_specs:
from ..stats.lme_solver import compute_lme_critical_values
- n_fixed = len(metadata.target_indices)
- # n_fixed_effects = number of columns in X_expanded (excluding intercept)
- # This equals the total effect count minus cluster effects
- n_fixed_total = len(metadata.effect_sizes)
- if metadata.cluster_effect_indices:
- n_fixed_total -= len(metadata.cluster_effect_indices)
+ # Use test formula dimensions when subsetting with random effects
+ if metadata.test_column_indices is not None and metadata.test_has_random_effects:
+ if metadata.test_target_indices is None:
+ raise RuntimeError("test_target_indices must be set when test_column_indices is present")
+ n_fixed = len(metadata.test_target_indices)
+ n_fixed_total = metadata.test_effect_count
+ else:
+ n_fixed = len(metadata.target_indices)
+ # n_fixed_effects = number of columns in X_expanded (excluding intercept)
+ # This equals the total effect count minus cluster effects
+ n_fixed_total = len(metadata.effect_sizes)
+ if metadata.cluster_effect_indices:
+ n_fixed_total -= len(metadata.cluster_effect_indices)
chi2_crit, z_crit, correction_z_crits = compute_lme_critical_values(
self.alpha, n_fixed_total, n_fixed, metadata.correction_method
)
@@ -162,19 +169,18 @@ def run_power_simulations(
raise SimulationCancelled("Simulation cancelled by user")
- sim_seed = self.seed + 4 * sim_id if self.seed is not None else None
-
- # Apply perturbations if in scenario mode
- if scenario_config is not None and apply_perturbations_func is not None:
- perturbed_corr, perturbed_types = apply_perturbations_func(
- metadata.correlation_matrix,
- metadata.var_types,
- scenario_config,
- sim_seed,
- )
- else:
- perturbed_corr = metadata.correlation_matrix
- perturbed_types = metadata.var_types
+ sim_seed = self.seed + 12 * sim_id if self.seed is not None else None
+
+ # Apply per-simulation perturbations (correlation noise, distribution swaps)
+ # Zero-valued params in optimistic scenario are no-ops
+ if apply_perturbations_func is None:
+ raise RuntimeError("apply_perturbations_func must be provided")
+ perturbed_corr, perturbed_types = apply_perturbations_func(
+ metadata.correlation_matrix,
+ metadata.var_types,
+ scenario_config,
+ sim_seed,
+ )
result = self._single_simulation(
sim_id=sim_id,
@@ -326,7 +332,9 @@ def _single_simulation(
first_spec = next(iter(metadata.cluster_specs.values()))
sample_size = first_spec["n_clusters"] * first_spec["cluster_size"]
- # Check if strict mode with uploaded data
+ # Strict-mode bootstrap: resample whole rows from uploaded data to
+ # preserve exact inter-variable relationships, then generate y from
+ # the bootstrapped X. This bypasses the normal X-generation pipeline.
if metadata.preserve_correlation == "strict" and metadata.uploaded_raw_data is not None:
# Strict mode: bootstrap uploaded data + generate created variables separately
from ..stats.data_generation import bootstrap_uploaded_data
@@ -336,7 +344,7 @@ def _single_simulation(
sample_size,
metadata.uploaded_raw_data,
metadata.uploaded_var_metadata,
- sim_seed,
+ sim_seed + 3 if sim_seed is not None else None,
)
# Merge uploaded and created non-factor variables
@@ -367,7 +375,7 @@ def _single_simulation(
X_factors = X_uploaded_factors
else:
# Mixed: generate all factors, replace uploaded factor columns
- X_factors = _generate_factors(sample_size, metadata.factor_specs, sim_seed)
+ X_factors = _generate_factors(sample_size, metadata.factor_specs, sim_seed + 3 if sim_seed is not None else None)
# Overwrite uploaded factor dummy columns with bootstrapped data
if X_uploaded_factors.shape[1] > 0:
col_offset = 0
@@ -400,14 +408,14 @@ def _single_simulation(
X_non_factors = np.empty((sample_size, 0), dtype=float)
# Generate factor variables (as dummy variables)
- X_factors = _generate_factors(sample_size, metadata.factor_specs, sim_seed)
+ X_factors = _generate_factors(sample_size, metadata.factor_specs, sim_seed + 3 if sim_seed is not None else None)
# Compute LME perturbations (ICC jitter, non-normal RE dist)
lme_perturbations = None
- if metadata.cluster_specs and scenario_config is not None:
+ if metadata.cluster_specs:
from ..core.scenarios import apply_lme_perturbations
- lme_perturbations = apply_lme_perturbations(metadata.cluster_specs, scenario_config, sim_seed)
+ lme_perturbations = apply_lme_perturbations(metadata.cluster_specs, scenario_config or {}, sim_seed)
# Generate cluster random effects (independent of upload mode)
re_result = None # Phase 2: random effects result for slopes/nesting
@@ -448,6 +456,16 @@ def _single_simulation(
# Create extended design matrix with interactions (excludes cluster effects)
X_expanded = create_X_extended_func(X)
+ # Test formula column subsetting: use reduced design matrix for analysis
+ if metadata.test_column_indices is not None:
+ X_test = X_expanded[:, metadata.test_column_indices]
+ if metadata.test_target_indices is None:
+ raise RuntimeError("test_target_indices must be set when test_column_indices is present")
+ test_target_indices = metadata.test_target_indices
+ else:
+ X_test = X_expanded
+ test_target_indices = metadata.target_indices
+
# Split effect sizes: fixed effects vs cluster effects
# Use precomputed values (Phase 2 optimization)
if metadata.cluster_effect_indices:
@@ -457,6 +475,21 @@ def _single_simulation(
fixed_effect_sizes = metadata.fixed_effect_sizes_cached
cluster_effect_sizes = None
+ # Residual coin flip: decide whether this simulation uses non-normal errors
+ residual_dist = 0 # normal
+ residual_df = 10.0
+ residual_change_prob = scenario_config.get("residual_change_prob", 0.0) if scenario_config else 0.0
+ if residual_change_prob > 0:
+ if scenario_config is None:
+ raise RuntimeError("scenario_config must be provided when residual_change_prob > 0")
+ coin_rng = np.random.RandomState(sim_seed + 7 if sim_seed is not None else None)
+ if coin_rng.random() < residual_change_prob:
+ residual_dists = scenario_config.get("residual_dists", ["heavy_tailed", "skewed"])
+ picked = coin_rng.choice(residual_dists)
+ dist_map = {"heavy_tailed": 1, "skewed": 2}
+ residual_dist = dist_map.get(picked, 0)
+ residual_df = float(scenario_config.get("residual_df", 10))
+
# Generate dependent variable with fixed effects only
y = generate_y_func(
X_expanded=X_expanded,
@@ -464,6 +497,8 @@ def _single_simulation(
heterogeneity=metadata.heterogeneity,
heteroskedasticity=metadata.heteroskedasticity,
sim_seed=sim_seed,
+ residual_dist=residual_dist,
+ residual_df=residual_df,
)
# Add cluster random effects contribution
@@ -478,12 +513,6 @@ def _single_simulation(
if re_result is not None and not np.allclose(re_result.slope_contribution, 0):
y = y + re_result.slope_contribution
- # Apply LME residual perturbations (non-normal residuals)
- if metadata.cluster_specs and scenario_config is not None:
- from ..core.scenarios import apply_lme_residual_perturbations
-
- y = apply_lme_residual_perturbations(y, scenario_config, sim_seed)
-
# Determine cluster IDs for the solver
cluster_ids: Optional[np.ndarray]
if re_result is not None:
@@ -496,23 +525,25 @@ def _single_simulation(
cluster_ids = metadata.cluster_ids_template
# Route to correct analysis method
- if cluster_ids is not None:
+ # When test_formula specifies no random effects, use OLS even if generation has clusters
+ use_lme = cluster_ids is not None and not (metadata.test_column_indices is not None and not metadata.test_has_random_effects)
+ if use_lme:
# Mixed model path (LME)
from ..stats.mixed_models import _lme_analysis_wrapper
+ assert cluster_ids is not None # narrowed by use_lme guard above
lme_result = _lme_analysis_wrapper(
- X_expanded,
+ X_test,
y,
- metadata.target_indices,
+ test_target_indices,
cluster_ids,
- metadata.cluster_column_indices,
metadata.correction_method,
self.alpha,
backend="custom",
verbose=metadata.verbose,
- chi2_crit=getattr(metadata, "lme_chi2_crit", None),
- z_crit=getattr(metadata, "lme_z_crit", None),
- correction_z_crits=getattr(metadata, "lme_correction_z_crits", None),
+ chi2_crit=metadata.lme_chi2_crit,
+ z_crit=metadata.lme_z_crit,
+ correction_z_crits=metadata.lme_correction_z_crits,
re_result=re_result,
)
@@ -539,16 +570,20 @@ def _single_simulation(
else:
# Standard OLS path
results = analyze_func(
- X_expanded,
+ X_test,
y,
- metadata.target_indices,
+ test_target_indices,
self.alpha,
metadata.correction_method,
)
diagnostics = None
- # Extract results: [f_sig, uncorr..., corr..., (wald_flag)]
- n_targets = len(metadata.target_indices)
+ # Result array layout: [F_sig, uncorrected[n_targets], corrected[n_targets], wald_flag?]
+ # - F_sig (index 0): overall model F-test significance (1.0 or 0.0)
+ # - uncorrected[1..n]: per-target t-test significance without correction
+ # - corrected[n+1..2n]: per-target significance with multiple-comparison correction
+ # - wald_flag (optional, LME only): 1.0 if Wald test was used instead of LRT
+ n_targets = len(test_target_indices)
f_significant = bool(results[0])
uncorrected = results[1 : 1 + n_targets].astype(bool)
corrected = results[1 + n_targets : 1 + 2 * n_targets].astype(bool)
@@ -560,19 +595,19 @@ def _single_simulation(
wald_flag = bool(results[expected_len])
# Post-hoc pairwise contrasts (OLS path only)
- if metadata.posthoc_specs and cluster_ids is None:
+ if metadata.posthoc_specs and not use_lme:
from ..stats.ols import compute_posthoc_contrasts
ph_uncorr, ph_corr, regular_override = compute_posthoc_contrasts(
- X_expanded,
+ X_test,
y,
metadata.posthoc_specs,
metadata.posthoc_method,
metadata.posthoc_t_crit,
metadata.posthoc_tukey_crits,
- target_indices=metadata.target_indices,
+ target_indices=test_target_indices,
correction_method=metadata.correction_method,
- correction_t_crits_combined=getattr(metadata, "posthoc_correction_t_crits_combined", None),
+ correction_t_crits_combined=metadata.posthoc_correction_t_crits_combined,
)
# If FDR/Holm combined correction was applied, override regular corrected
@@ -645,7 +680,7 @@ class SimulationMetadata:
correction_method: Encoded multiple-comparison correction
(0=none, 1=Bonferroni, 2=BH, 3=Holm).
heterogeneity: SD of random effect-size multiplier.
- heteroskedasticity: Correlation between first predictor and error SD.
+ heteroskedasticity: Correlation between predicted values and error SD.
preserve_correlation: Upload correlation mode
(``"no"``/``"partial"``/``"strict"``).
uploaded_raw_data: Normalised raw data for strict-mode bootstrap.
@@ -728,6 +763,13 @@ def __init__(
self.posthoc_method: str = "t-test"
self.posthoc_tukey_crits: Dict[str, float] = {}
self.posthoc_t_crit: float = 0.0
+ self.posthoc_correction_t_crits_combined: Optional[np.ndarray] = None
+
+ # Test formula fields (for model misspecification testing)
+ self.test_column_indices: Optional[np.ndarray] = None
+ self.test_target_indices: Optional[np.ndarray] = None
+ self.test_effect_count: Optional[int] = None # p for critical value computation
+ self.test_has_random_effects: bool = False # Whether test formula has (1|group) etc.
def _compute_fixed_effect_variance(registry) -> float:
@@ -779,8 +821,13 @@ def _compute_fixed_effect_variance(registry) -> float:
factor_info = registry._factors[factor_name]
proportions = factor_info.get("proportions")
if proportions is not None:
- # level is 1-indexed; proportions list is 0-indexed
- p_k = proportions[level - 1]
+ level_labels = factor_info.get("level_labels")
+ if level_labels is not None:
+ # String level labels — look up position by label
+ p_k = proportions[level_labels.index(str(level))]
+ else:
+ # Integer levels are 1-indexed; proportions list is 0-indexed
+ p_k = proportions[level - 1]
else:
# Equal proportions (default)
n_levels = factor_info["n_levels"]
@@ -814,6 +861,7 @@ def prepare_metadata(
model,
target_tests: List[str],
correction: Optional[str] = None,
+ test_formula_effects: Optional[List[str]] = None,
) -> SimulationMetadata:
"""
Prepare simulation metadata from model state.
@@ -825,6 +873,9 @@ def prepare_metadata(
model: MCPowerModel instance
target_tests: List of effects to test
correction: Multiple comparison correction method
+ test_formula_effects: Optional list of effect names from a test
+ formula. When provided, the metadata will include column
+ indices for subsetting X_expanded to the test model.
Returns:
SimulationMetadata instance
@@ -960,8 +1011,6 @@ def prepare_metadata(
upload_data_values=model.upload_data_values if model.upload_data_values is not None else np.zeros((2, 2), dtype=np.float64),
effect_sizes=effect_sizes,
correction_method=correction_method,
- heterogeneity=model.heterogeneity,
- heteroskedasticity=model.heteroskedasticity,
preserve_correlation=model._preserve_correlation,
uploaded_raw_data=model._uploaded_raw_data,
uploaded_var_metadata=model._uploaded_var_metadata,
@@ -982,4 +1031,20 @@ def prepare_metadata(
metadata.posthoc_specs = model._posthoc_specs
metadata.posthoc_method = "tukey" if is_tukey_correction else "t-test"
+ # Test formula column subsetting
+ if test_formula_effects is not None:
+ from ..utils.test_formula_utils import _compute_test_column_indices, _remap_target_indices
+
+ # Get all non-cluster effect names in registry order
+ all_effect_names = [name for name in registry._effects if name not in registry.cluster_effect_names]
+
+ test_col_indices = _compute_test_column_indices(all_effect_names, test_formula_effects)
+ metadata.test_column_indices = test_col_indices
+ metadata.test_effect_count = len(test_col_indices)
+
+ # Remap target indices to X_test space
+ # Only remap targets that exist in the test formula
+ valid_targets = np.array([idx for idx in target_indices if idx in test_col_indices], dtype=np.int64)
+ metadata.test_target_indices = _remap_target_indices(valid_targets, test_col_indices)
+
return metadata
diff --git a/mcpower/core/variables.py b/mcpower/core/variables.py
index e311606..338d40b 100644
--- a/mcpower/core/variables.py
+++ b/mcpower/core/variables.py
@@ -367,76 +367,42 @@ def expand_factors(self) -> None:
level_labels = factor_info.get("level_labels")
reference_level = factor_info.get("reference_level", 1)
+ # Compute non-reference levels once
if level_labels is not None:
- # Named levels: skip the reference, create dummies for the rest
- non_ref_labels = [lb for lb in level_labels if lb != str(reference_level)]
- for label in non_ref_labels:
- dummy_name = f"{factor_name}[{label}]"
-
- # Create dummy predictor
- dummy_pred = PredictorVar(
- name=dummy_name,
- var_type="factor_dummy",
- is_dummy=True,
- factor_source=factor_name,
- factor_level=label,
- column_index=col_idx,
- level_labels=level_labels,
- )
- new_predictors[dummy_name] = dummy_pred
-
- # Create main effect for dummy
- dummy_eff = Effect(
- name=dummy_name,
- effect_type="main",
- var_names=[dummy_name],
- column_index=col_idx,
- factor_source=factor_name,
- factor_level=label,
- )
- new_effects[dummy_name] = dummy_eff
-
- # Store dummy mapping
- self._factor_dummies[dummy_name] = {
- "factor_name": factor_name,
- "level": label,
- }
-
- col_idx += 1
+ non_ref = [lb for lb in level_labels if lb != str(reference_level)]
else:
- # Original integer-indexed behavior
- for level in range(2, n_levels + 1):
- dummy_name = f"{factor_name}[{level}]"
-
- # Create dummy predictor
- dummy_pred = PredictorVar(
- name=dummy_name,
- var_type="factor_dummy",
- is_dummy=True,
- factor_source=factor_name,
- factor_level=level,
- column_index=col_idx,
- )
- new_predictors[dummy_name] = dummy_pred
-
- # Create main effect for dummy
- dummy_eff = Effect(
- name=dummy_name,
- effect_type="main",
- var_names=[dummy_name],
- column_index=col_idx,
- factor_source=factor_name,
- factor_level=level,
- )
- new_effects[dummy_name] = dummy_eff
+ non_ref = list(range(2, n_levels + 1))
+
+ for level in non_ref:
+ dummy_name = f"{factor_name}[{level}]"
+
+ dummy_pred = PredictorVar(
+ name=dummy_name,
+ var_type="factor_dummy",
+ is_dummy=True,
+ factor_source=factor_name,
+ factor_level=level,
+ column_index=col_idx,
+ level_labels=level_labels if level_labels is not None else None,
+ )
+ new_predictors[dummy_name] = dummy_pred
+
+ dummy_eff = Effect(
+ name=dummy_name,
+ effect_type="main",
+ var_names=[dummy_name],
+ column_index=col_idx,
+ factor_source=factor_name,
+ factor_level=level,
+ )
+ new_effects[dummy_name] = dummy_eff
- # Store dummy mapping
- self._factor_dummies[dummy_name] = {
- "factor_name": factor_name,
- "level": level,
- }
+ self._factor_dummies[dummy_name] = {
+ "factor_name": factor_name,
+ "level": level,
+ }
- col_idx += 1
+ col_idx += 1
# Handle interactions involving factors — Cartesian product of
# non-reference dummy levels across all factor components.
@@ -503,6 +469,13 @@ def get_effect_sizes(self) -> np.ndarray:
def get_var_types(self) -> np.ndarray:
"""Get variable types as numpy array (for data generation)."""
+ # Type codes: 0-5 are parametric distributions generated from scratch.
+ # 97/98/99 are sentinel codes for uploaded-data variables whose values
+ # come from bootstrapped/quantile-matched empirical data rather than
+ # parametric generation:
+ # 97 = uploaded_factor (factor from uploaded data)
+ # 98 = uploaded_binary (binary from uploaded data)
+ # 99 = uploaded_data (continuous from uploaded data)
type_mapping = {
"normal": 0,
"binary": 1,
@@ -717,28 +690,20 @@ def register_cluster(
def _reindex_predictors(self) -> None:
"""Reindex all predictors to maintain order: non_factor | cluster_effect | dummies."""
- col_idx = 0
+ non_factor = []
+ cluster = []
+ dummies = []
- # Non-factor predictors first
- for name in sorted(self._predictors.keys(), key=lambda x: self._predictors[x].column_index or 0):
- pred = self._predictors[name]
- if not pred.is_factor and not pred.is_dummy and pred.var_type != "cluster_effect":
- pred.column_index = col_idx
- col_idx += 1
-
- # Cluster effect predictors second
- for name in sorted(self._predictors.keys(), key=lambda x: self._predictors[x].column_index or 0):
- pred = self._predictors[name]
- if pred.var_type == "cluster_effect":
- pred.column_index = col_idx
- col_idx += 1
-
- # Factor dummies last
for name in sorted(self._predictors.keys(), key=lambda x: self._predictors[x].column_index or 0):
pred = self._predictors[name]
if pred.is_dummy:
- pred.column_index = col_idx
- col_idx += 1
+ dummies.append(pred)
+ elif pred.var_type == "cluster_effect":
+ cluster.append(pred)
+ elif not pred.is_factor:
+ non_factor.append(pred)
+
+ for col_idx, pred in enumerate(non_factor + cluster + dummies):
+ pred.column_index = col_idx
- # Update effect indices
self._update_effect_indices()
diff --git a/mcpower/model.py b/mcpower/model.py
index 4fa2c4b..9a5f813 100644
--- a/mcpower/model.py
+++ b/mcpower/model.py
@@ -123,10 +123,9 @@ def __init__(self, data_generation_formula: str):
self._pending_factor_levels: Optional[str] = None
self._pending_effects: Optional[str] = None
self._pending_correlations: Optional[Union[str, np.ndarray]] = None
- self._pending_heterogeneity: Optional[float] = None
- self._pending_heteroskedasticity: Optional[float] = None
self._pending_data: Optional[Dict[str, Any]] = None
self._pending_clusters: Dict[str, Dict] = {} # {grouping_var: {n_clusters, cluster_size, icc}}
+ self._effects_set: bool = False # True after set_effects() has been called
# Detect mixed model formula
if self._registry._random_effects_parsed:
@@ -134,8 +133,6 @@ def __init__(self, data_generation_formula: str):
# Applied state
self._applied = False
- self.heterogeneity = 0.0
- self.heteroskedasticity = 0.0
# Data storage
self.upload_normal_values: Optional[np.ndarray] = None
@@ -385,6 +382,7 @@ def set_effects(self, effects_string: str):
raise ValueError("effects_string cannot be empty")
self._pending_effects = effects_string
+ self._effects_set = True
self._applied = False
return self
@@ -432,13 +430,16 @@ def set_variable_type(self, variable_types_string: str):
- ``"normal"`` — standard normal (default).
- ``"binary"`` or ``"binary(p)"`` — Bernoulli with proportion *p*
(default 0.5).
- - ``"skewed"`` — heavy-tailed (t-distribution, df=3).
+ - ``"right_skewed"`` — positively skewed distribution.
+ - ``"left_skewed"`` — negatively skewed distribution.
+ - ``"high_kurtosis"`` — heavy-tailed (t-distribution, df=3).
+ - ``"uniform"`` — uniform distribution.
- ``"factor(k)"`` — categorical with *k* levels (creates *k-1*
dummy variables).
- ``"factor(k, p1, p2, ...)"`` — factor with custom level
proportions.
- Example: ``"x1=binary, x2=skewed, x3=factor(3)"``.
+ Example: ``"x1=binary, x2=right_skewed, x3=factor(3)"``.
Returns:
self: For method chaining.
@@ -479,62 +480,6 @@ def set_factor_levels(self, spec: str):
self._applied = False
return self
- def set_heterogeneity(self, heterogeneity: float):
- """Set heterogeneity (random variation) in effect sizes.
-
- When non-zero, each simulation draws a per-simulation effect-size
- multiplier from a normal distribution with mean 1 and the given
- standard deviation. This models uncertainty about the true effect
- size — for example, ``heterogeneity=0.1`` means effect sizes vary
- by roughly +/- 10% across simulations.
-
- This setting is deferred until ``apply()`` is called.
-
- Args:
- heterogeneity: Standard deviation of the random effect-size
- multiplier. Must be non-negative. Default is 0 (no variation).
-
- Returns:
- self: For method chaining.
-
- Raises:
- TypeError: If *heterogeneity* is not numeric.
- """
- if not isinstance(heterogeneity, (int, float)):
- raise TypeError("heterogeneity must be a number")
-
- self._pending_heterogeneity = float(heterogeneity)
- self._applied = False
- return self
-
- def set_heteroskedasticity(self, heteroskedasticity_correlation: float):
- """Set heteroskedasticity (non-constant error variance).
-
- Introduces a correlation between the first predictor's values and
- the error standard deviation, producing variance that increases (or
- decreases) with the predictor. This violates the homoskedasticity
- assumption and typically reduces power.
-
- This setting is deferred until ``apply()`` is called.
-
- Args:
- heteroskedasticity_correlation: Correlation between the first
- predictor and the error standard deviation, in the range
- [-1, 1]. Default is 0 (homoskedastic errors).
-
- Returns:
- self: For method chaining.
-
- Raises:
- TypeError: If the value is not numeric.
- """
- if not isinstance(heteroskedasticity_correlation, (int, float)):
- raise TypeError("heteroskedasticity_correlation must be a number")
-
- self._pending_heteroskedasticity = float(heteroskedasticity_correlation)
- self._applied = False
- return self
-
def set_cluster(
self,
grouping_var: str,
@@ -769,6 +714,7 @@ def upload_data(
"preserve_factor_level_names": preserve_factor_level_names,
}
self._applied = False
+ return self
def set_scenario_configs(self, configs_dict: Dict):
"""Set custom scenario configurations for robustness analysis.
@@ -786,7 +732,9 @@ def set_scenario_configs(self, configs_dict: Dict):
configs_dict: Mapping of scenario names to configuration dicts.
Each configuration may include keys such as
``"heterogeneity"``, ``"heteroskedasticity"``,
- ``"effect_size_jitter"``, and ``"distribution_jitter"``.
+ ``"correlation_noise_sd"``, and ``"distribution_change_prob"``.
+ See ``DEFAULT_SCENARIO_CONFIG`` in ``mcpower.core.scenarios``
+ for the full list of keys.
Returns:
self: For method chaining.
@@ -802,7 +750,8 @@ def set_scenario_configs(self, configs_dict: Dict):
if scenario in merged:
merged[scenario].update(config)
else:
- merged[scenario] = config
+ # New custom scenarios inherit all keys from optimistic baseline
+ merged[scenario] = {**DEFAULT_SCENARIO_CONFIG["optimistic"], **config}
self._scenario_configs = merged
print(f"Custom scenario configs set: {', '.join(configs_dict.keys())}")
@@ -812,7 +761,7 @@ def set_scenario_configs(self, configs_dict: Dict):
# Apply method (processes all pending settings)
# =========================================================================
- def apply(self):
+ def _apply(self):
"""
Apply all pending settings to the model.
@@ -857,16 +806,22 @@ def apply(self):
# 7. Apply correlations
self._apply_correlations(_parser)
- # 8. Apply heterogeneity/heteroskedasticity
- self._apply_heterogeneity()
-
- # 9. Validate model is ready
+ # 8. Validate model is ready
model_result = _validate_model_ready(self)
model_result.raise_if_invalid()
- # Invalidate effect plan cache when settings change (Phase 2 optimization)
+ # Invalidate the effect plan cache — apply() rebuilds the variable
+ # registry state, so any cached column mappings are now stale.
self._effect_plan_cache = None
+ # Clear pending state to prevent double-application
+ self._pending_variable_types = None
+ self._pending_factor_levels = None
+ self._pending_effects = None
+ self._pending_correlations = None
+ self._pending_data = None
+ self._pending_clusters = {}
+
self._applied = True
print("Model settings applied successfully")
return self
@@ -1024,6 +979,21 @@ def _apply_data(self):
# Extract matched data
matched_data = data[:, matched_indices]
+ # Reject NaN values early
+ try:
+ if np.isnan(matched_data.astype(np.float64)).any():
+ nan_cols = [
+ matched_columns[i] for i in range(matched_data.shape[1]) if np.isnan(matched_data[:, i].astype(np.float64)).any()
+ ]
+ raise ValueError(
+ f"Uploaded data contains NaN values in columns: {', '.join(nan_cols)}. "
+ f"Remove or impute missing values before uploading."
+ )
+ except (ValueError, TypeError):
+ # Object dtype columns (strings) can't be converted to float for NaN check.
+ # NaN check for numeric columns will happen after string encoding below.
+ pass
+
# Convert to float64 if object dtype (common with mixed-type DataFrames)
# String columns are encoded to integer indices; mapping is stored in string_col_indices
string_col_indices = {}
@@ -1178,11 +1148,7 @@ def _apply_data_normal_mode(self, data, columns, type_info, mode, data_types_ove
level_labels = info.get("level_labels")
# Determine reference from data_types tuple override
- reference_level = None
- if col in data_types_override:
- dt = data_types_override[col]
- if isinstance(dt, tuple) and len(dt) == 2:
- reference_level = str(dt[1])
+ reference_level = self._extract_reference_level(data_types_override, col)
# Calculate proportions for each level
proportions = []
@@ -1200,7 +1166,10 @@ def _apply_data_normal_mode(self, data, columns, type_info, mode, data_types_ove
else: # continuous
# Normalize: mean=0, sd=1
- normalized = (col_data - np.mean(col_data)) / np.std(col_data, ddof=1)
+ std = np.std(col_data, ddof=1)
+ if std < 1e-15:
+ raise ValueError(f"Column '{col}' has zero variance (constant value). Remove it from the model or check your data.")
+ normalized = (col_data - np.mean(col_data)) / std
# Create lookup tables (type 99)
normal_vals, uploaded_vals = create_uploaded_lookup_tables(normalized.reshape(-1, 1))
@@ -1324,11 +1293,7 @@ def _apply_data_strict_mode(self, data, columns, type_info, data_types_override=
level_labels = info.get("level_labels")
# Determine reference from data_types tuple override
- reference_level = None
- if col in data_types_override:
- dt = data_types_override[col]
- if isinstance(dt, tuple) and len(dt) == 2:
- reference_level = str(dt[1])
+ reference_level = self._extract_reference_level(data_types_override, col)
self._uploaded_var_metadata[col] = {
"type": "factor",
@@ -1355,7 +1320,10 @@ def _apply_data_strict_mode(self, data, columns, type_info, data_types_override=
continuous_cols.append(idx)
# Normalize
col_data = data[:, idx]
- normalized_data[:, idx] = (col_data - np.mean(col_data)) / np.std(col_data, ddof=1)
+ std = np.std(col_data, ddof=1)
+ if std < 1e-15:
+ raise ValueError(f"Column '{col}' has zero variance (constant value). Remove it from the model or check your data.")
+ normalized_data[:, idx] = (col_data - np.mean(col_data)) / std
self._uploaded_var_metadata[col] = {
"type": "continuous",
@@ -1481,22 +1449,6 @@ def _apply_correlations(self, _parser):
self._registry.set_correlation_matrix(correlations_input)
print("Correlation matrix set")
- def _apply_heterogeneity(self):
- """Validate and apply pending heterogeneity and heteroskedasticity settings."""
- if self._pending_heterogeneity is not None:
- if self._pending_heterogeneity < 0:
- raise ValueError("heterogeneity must be non-negative")
- self.heterogeneity = self._pending_heterogeneity
- if self.heterogeneity > 0:
- print(f"Heterogeneity: SD = {self.heterogeneity}")
-
- if self._pending_heteroskedasticity is not None:
- if not -1 <= self._pending_heteroskedasticity <= 1:
- raise ValueError("heteroskedasticity_correlation must be between -1 and 1")
- self.heteroskedasticity = self._pending_heteroskedasticity
- if abs(self.heteroskedasticity) > 1e-8:
- print(f"Heteroskedasticity: correlation = {self.heteroskedasticity}")
-
# =========================================================================
# Analysis methods
# =========================================================================
@@ -1507,7 +1459,7 @@ def find_power(
target_test: str = "all",
correction: Optional[str] = None,
print_results: bool = True,
- scenarios: bool = False,
+ scenarios: Union[bool, List[str]] = False,
summary: str = "short",
return_results: bool = False,
test_formula: str = "",
@@ -1529,12 +1481,16 @@ def find_power(
Duplicate tests raise ``ValueError``.
correction: Multiple comparison correction (None, "bonferroni", "benjamini-hochberg", "holm")
print_results: Whether to print results
- scenarios: Run scenario analysis
+ scenarios: Scenario analysis control:
+ - ``False`` (default): no scenario analysis.
+ - ``True``: run all configured scenarios.
+ - List of scenario names: run only the specified scenarios
+ (e.g. ``["optimistic", "extreme"]``). Case-insensitive.
summary: Output detail level ("short" or "long")
return_results: Return results dict
test_formula: Formula for statistical testing (default: use data generation formula).
If the formula contains random effects like (1|school), analysis switches to
- mixed model testing (not yet implemented).
+ mixed model testing.
progress_callback: Progress reporting control:
- ``None`` (default): auto-use ``PrintReporter`` when
*print_results* is ``True``.
@@ -1549,7 +1505,10 @@ def find_power(
"""
# Auto-apply if settings have changed
if not self._applied:
- self.apply()
+ self._apply()
+
+ # Resolve scenarios parameter
+ scenario_filter = self._resolve_scenarios(scenarios)
# Validate sample size (basic: >= 20, type check)
_validate_sample_size(sample_size).raise_if_invalid()
@@ -1558,9 +1517,6 @@ def find_power(
n_variables = len(self._registry.effect_names)
_validate_sample_size_for_model(sample_size, n_variables).raise_if_invalid()
- # Validate and adjust cluster sample sizes
- self._validate_cluster_sample_size(sample_size)
-
# Warn if sample size is much larger than uploaded data
if self._uploaded_data_n > 0 and sample_size > 3 * self._uploaded_data_n:
print(
@@ -1570,33 +1526,13 @@ def find_power(
)
self._validate_analysis_inputs(correction)
- resolved_test_formula = self._resolve_test_formula(test_formula)
- target_tests = self._parse_target_tests(target_test)
-
- if correction and correction.lower() == "tukey" and not self._posthoc_specs:
- raise ValueError(
- "Tukey correction requires at least one post-hoc comparison "
- "(e.g., target_test='group[0] vs group[1]'). "
- "Tukey HSD only applies to pairwise contrasts between factor levels."
- )
-
- # Resolve progress callback
- from .progress import PrintReporter, ProgressReporter, compute_total_simulations
-
- if progress_callback is None:
- effective_cb = PrintReporter() if print_results else None
- elif progress_callback is False:
- effective_cb = None
- else:
- effective_cb = progress_callback
+ resolved_test_formula, test_formula_effects, test_random_effects = self._resolve_test_formula(test_formula)
+ target_tests = self._parse_target_tests(target_test, test_formula_effects=test_formula_effects)
+ self._validate_tukey_posthoc(correction)
- reporter = None
- if effective_cb is not None:
- n_scenarios = (len(self._scenario_configs or DEFAULT_SCENARIO_CONFIG) + 1) if scenarios else 1
- total = compute_total_simulations(self._effective_n_simulations, 1, n_scenarios)
- reporter = ProgressReporter(total, effective_cb)
+ reporter = self._resolve_progress(progress_callback, print_results, scenario_filter)
- if scenarios:
+ if scenario_filter is not None:
result = self._run_scenario_analysis(
"power",
sample_size=sample_size,
@@ -1605,8 +1541,11 @@ def find_power(
summary=summary,
print_results=print_results,
test_formula=resolved_test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=reporter,
cancel_check=cancel_check,
+ scenario_filter=scenario_filter,
)
else:
if reporter is not None:
@@ -1615,7 +1554,10 @@ def find_power(
sample_size,
target_tests,
correction,
+ scenario_config=DEFAULT_SCENARIO_CONFIG["optimistic"],
test_formula=resolved_test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=reporter,
cancel_check=cancel_check,
)
@@ -1623,7 +1565,7 @@ def find_power(
if reporter is not None:
reporter.finish()
- if not scenarios and print_results:
+ if scenario_filter is None and print_results:
print(f"\n{'=' * 80}")
print("MONTE CARLO POWER ANALYSIS RESULTS")
print(f"{'=' * 80}")
@@ -1641,7 +1583,7 @@ def find_sample_size(
by: int = 5,
correction: Optional[str] = None,
print_results: bool = True,
- scenarios: bool = False,
+ scenarios: Union[bool, List[str]] = False,
summary: str = "short",
return_results: bool = False,
test_formula: str = "",
@@ -1659,12 +1601,16 @@ def find_sample_size(
by: Step size between sample sizes
correction: Multiple comparison correction
print_results: Whether to print results
- scenarios: Run scenario analysis
+ scenarios: Scenario analysis control:
+ - ``False`` (default): no scenario analysis.
+ - ``True``: run all configured scenarios.
+ - List of scenario names: run only the specified scenarios
+ (e.g. ``["optimistic", "extreme"]``). Case-insensitive.
summary: Output detail level
return_results: Return results dict
test_formula: Formula for statistical testing (default: use data generation formula).
If the formula contains random effects like (1|school), analysis switches to
- mixed model testing (not yet implemented).
+ mixed model testing.
progress_callback: Progress reporting control:
- ``None`` (default): auto-use ``PrintReporter`` when
*print_results* is ``True``.
@@ -1680,7 +1626,10 @@ def find_sample_size(
"""
# Auto-apply if settings have changed
if not self._applied:
- self.apply()
+ self._apply()
+
+ # Resolve scenarios parameter
+ scenario_filter = self._resolve_scenarios(scenarios)
# Validate from_size meets minimum requirements
_validate_sample_size(from_size).raise_if_invalid()
@@ -1696,40 +1645,20 @@ def find_sample_size(
)
self._validate_analysis_inputs(correction)
- resolved_test_formula = self._resolve_test_formula(test_formula)
+ resolved_test_formula, test_formula_effects, test_random_effects = self._resolve_test_formula(test_formula)
validation_result = _validate_sample_size_range(from_size, to_size, by)
for warning in validation_result.warnings:
print(f"Warning: {warning}")
validation_result.raise_if_invalid()
- target_tests = self._parse_target_tests(target_test)
-
- if correction and correction.lower() == "tukey" and not self._posthoc_specs:
- raise ValueError(
- "Tukey correction requires at least one post-hoc comparison "
- "(e.g., target_test='group[0] vs group[1]'). "
- "Tukey HSD only applies to pairwise contrasts between factor levels."
- )
+ target_tests = self._parse_target_tests(target_test, test_formula_effects=test_formula_effects)
+ self._validate_tukey_posthoc(correction)
sample_sizes = list(range(from_size, to_size + 1, by))
- # Resolve progress callback
- from .progress import PrintReporter, ProgressReporter, compute_total_simulations
-
- if progress_callback is None:
- effective_cb = PrintReporter() if print_results else None
- elif progress_callback is False:
- effective_cb = None
- else:
- effective_cb = progress_callback
-
- reporter = None
- if effective_cb is not None:
- n_scenarios = (len(self._scenario_configs or DEFAULT_SCENARIO_CONFIG) + 1) if scenarios else 1
- total = compute_total_simulations(self._effective_n_simulations, len(sample_sizes), n_scenarios)
- reporter = ProgressReporter(total, effective_cb)
+ reporter = self._resolve_progress(progress_callback, print_results, scenario_filter, n_sample_sizes=len(sample_sizes))
- if scenarios:
+ if scenario_filter is not None:
result = self._run_scenario_analysis(
"sample_size",
target_tests=target_tests,
@@ -1738,8 +1667,11 @@ def find_sample_size(
summary=summary,
print_results=print_results,
test_formula=resolved_test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=reporter,
cancel_check=cancel_check,
+ scenario_filter=scenario_filter,
)
else:
if reporter is not None:
@@ -1748,7 +1680,10 @@ def find_sample_size(
sample_sizes,
target_tests,
correction,
+ scenario_config=DEFAULT_SCENARIO_CONFIG["optimistic"],
test_formula=resolved_test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=reporter,
cancel_check=cancel_check,
)
@@ -1756,7 +1691,7 @@ def find_sample_size(
if reporter is not None:
reporter.finish()
- if not scenarios and print_results:
+ if scenario_filter is None and print_results:
print(f"\n{'=' * 80}")
print("SAMPLE SIZE ANALYSIS RESULTS")
print(f"{'=' * 80}")
@@ -1780,6 +1715,8 @@ def _generate_dependent_variable(
heterogeneity: float = 0.0,
heteroskedasticity: float = 0.0,
sim_seed: Optional[int] = None,
+ residual_dist: int = 0,
+ residual_df: float = 10.0,
) -> np.ndarray:
"""Generate the dependent variable as y = X @ beta + error via the active backend."""
return get_backend().generate_y(
@@ -1788,19 +1725,103 @@ def _generate_dependent_variable(
heterogeneity,
heteroskedasticity,
sim_seed if sim_seed is not None else -1,
+ residual_dist,
+ residual_df,
)
# =========================================================================
# Internal methods
# =========================================================================
+ @staticmethod
+ def _extract_reference_level(data_types_override, col):
+ """Extract reference level from data_types_override tuple for a column."""
+ dt = data_types_override.get(col)
+ if isinstance(dt, tuple) and len(dt) == 2:
+ return str(dt[1])
+ return None
+
+ def _resolve_scenarios(self, scenarios: Union[bool, List[str]]) -> Optional[List[str]]:
+ """Resolve the scenarios parameter into a list of scenario names or None.
+
+ Args:
+ scenarios: ``False`` for no scenarios, ``True`` for all configured
+ scenarios, or a list of scenario names (case-insensitive).
+
+ Returns:
+ List of validated, lowercase scenario names, or ``None`` if
+ scenarios are disabled.
+
+ Raises:
+ ValueError: If any requested scenario name is not configured.
+ TypeError: If *scenarios* is not ``bool`` or a list of strings.
+ """
+ if scenarios is False:
+ return None
+
+ all_configs = self._scenario_configs or DEFAULT_SCENARIO_CONFIG
+ available = set(all_configs.keys())
+
+ if scenarios is True:
+ return list(all_configs.keys())
+
+ if not isinstance(scenarios, list):
+ raise TypeError(f"scenarios must be True, False, or a list of scenario names, got {type(scenarios).__name__}")
+
+ # Case-insensitive matching
+ available_lower = {k.lower(): k for k in available}
+ resolved = []
+ invalid = []
+ for name in scenarios:
+ if not isinstance(name, str):
+ raise TypeError(f"Scenario names must be strings, got {type(name).__name__}")
+ key = available_lower.get(name.lower())
+ if key is None:
+ invalid.append(name)
+ else:
+ resolved.append(key)
+
+ if invalid:
+ raise ValueError(f"Unknown scenario(s): {', '.join(repr(n) for n in invalid)}. Available: {', '.join(sorted(available))}")
+
+ return resolved
+
+ def _resolve_progress(self, progress_callback, print_results, scenario_filter, n_sample_sizes=1):
+ """Resolve progress_callback into a ProgressReporter or None."""
+ from .progress import PrintReporter, ProgressReporter, compute_total_simulations
+
+ if progress_callback is None:
+ effective_cb = PrintReporter() if print_results else None
+ elif progress_callback is False:
+ effective_cb = None
+ else:
+ effective_cb = progress_callback
+
+ if effective_cb is None:
+ return None
+
+ n_scenarios = len(scenario_filter) if scenario_filter is not None else 1
+ total = compute_total_simulations(self._effective_n_simulations, n_sample_sizes, n_scenarios)
+ return ProgressReporter(total, effective_cb)
+
def _validate_analysis_inputs(self, correction):
"""Validate the multiple-comparison correction method before analysis."""
result = _validate_correction_method(correction)
result.raise_if_invalid()
+ def _validate_tukey_posthoc(self, correction):
+ """Raise if Tukey correction is requested without posthoc specs."""
+ if correction and correction.lower() == "tukey" and not self._posthoc_specs:
+ raise ValueError(
+ "Tukey correction requires at least one post-hoc comparison "
+ "(e.g., target_test='group[0] vs group[1]'). "
+ "Tukey HSD only applies to pairwise contrasts between factor levels."
+ )
+
def _validate_cluster_sample_size(self, sample_size: int):
"""Derive missing cluster dimensions from sample_size and validate minimums."""
+ # NOTE: This method both validates AND mutates — it derives missing
+ # cluster_size/n_clusters from sample_size before checking minimums.
if not self._registry.cluster_names:
return # No clusters, nothing to do
@@ -1811,10 +1832,12 @@ def _validate_cluster_sample_size(self, sample_size: int):
if spec.n_clusters is not None:
spec.cluster_size = sample_size // spec.n_clusters
else:
- assert spec.cluster_size is not None
+ if spec.cluster_size is None:
+ raise RuntimeError(f"Cluster '{gv}': either n_clusters or cluster_size must be set")
spec.n_clusters = sample_size // spec.cluster_size
- assert spec.n_clusters is not None and spec.cluster_size is not None
+ if spec.n_clusters is None or spec.cluster_size is None:
+ raise RuntimeError(f"Cluster '{gv}': failed to derive n_clusters and cluster_size from sample_size={sample_size}")
actual_n = spec.n_clusters * spec.cluster_size
if actual_n != sample_size:
print(
@@ -1825,7 +1848,7 @@ def _validate_cluster_sample_size(self, sample_size: int):
_validate_cluster_sample_size(sample_size, spec.n_clusters, spec.cluster_size).raise_if_invalid()
- def _parse_target_tests(self, target_test: Union[str, List[str]]) -> List[str]:
+ def _parse_target_tests(self, target_test: Union[str, List[str]], test_formula_effects: Optional[List[str]] = None) -> List[str]:
"""Parse a target_test argument into a list of effect names to test.
Supports regular effect names (e.g. ``"x1"``, ``"overall"``),
@@ -1875,7 +1898,10 @@ def _parse_target_tests(self, target_test: Union[str, List[str]]) -> List[str]:
cluster_effects = self._registry.cluster_effect_names
if "all" in keywords:
- fixed_effects = [e for e in self._registry.effect_names if e not in cluster_effects]
+ if test_formula_effects is not None:
+ fixed_effects = [e for e in test_formula_effects if e not in cluster_effects]
+ else:
+ fixed_effects = [e for e in self._registry.effect_names if e not in cluster_effects]
keyword_expansion += ["overall"] + fixed_effects
if "all-posthoc" in keywords:
@@ -1929,6 +1955,17 @@ def _parse_target_tests(self, target_test: Union[str, List[str]]) -> List[str]:
"(e.g. 'all'), do not also list tests that are already included."
)
+ # -- Phase 7b: Validate explicit tests against test formula ----------------
+ if test_formula_effects is not None:
+ test_formula_set = set(test_formula_effects)
+ for test in expanded:
+ if " vs " in test or test == "overall":
+ continue
+ if test not in test_formula_set:
+ raise ValueError(
+ f"Target test '{test}' is not in the test formula. Available effects: {', '.join(test_formula_effects)}"
+ )
+
# -- Phase 8: Parse posthoc specs + validate ------------------------------
regular_tests: list[str] = []
posthoc_specs: list[PostHocSpec] = []
@@ -1982,6 +2019,8 @@ def _parse_target_tests(self, target_test: Union[str, List[str]]) -> List[str]:
# User level k (k≥2) = dummy factor[k]
effect_order = list(self._registry._effects.keys())
+ # Returns None for the reference level, which is absorbed into the
+ # intercept in dummy coding and has no dedicated design matrix column.
def _level_to_col(factor_name, user_level, _effect_order=effect_order):
factor_info = self._registry._factors[factor_name]
reference = factor_info.get("reference_level", 1)
@@ -2069,30 +2108,60 @@ def _create_X_extended(self, X):
return np.column_stack(columns) if columns else np.empty((X.shape[0], 0))
- def _prepare_metadata(self, target_tests, correction=None):
+ def _prepare_metadata(self, target_tests, correction=None, test_formula_effects=None):
"""Pre-compute all static simulation metadata from the current model state."""
- return prepare_metadata(self, target_tests, correction)
+ return prepare_metadata(self, target_tests, correction, test_formula_effects=test_formula_effects)
- def _resolve_test_formula(self, test_formula: str) -> str:
- """Resolve test formula and update _test_method accordingly.
+ def _resolve_test_formula(self, test_formula: str):
+ """Resolve test formula, validate, parse, and update _test_method.
- Returns the resolved formula string.
+ Returns:
+ Tuple of (formula_string, test_effect_names, random_effects).
+ test_effect_names is None when test_formula is empty (use generation formula).
"""
+ from .utils.parsers import _parse_equation
+
if not test_formula:
resolved = self._registry.equation
- else:
- resolved = test_formula
+ _, _, random_effects = _parse_equation(resolved)
+ if random_effects:
+ self._test_method = "mixed_model"
+ else:
+ self._test_method = "linear_regression"
+ return resolved, None, []
- from .utils.parsers import _parse_equation
+ # Validate test formula variables exist in the model
+ from .utils.validators import _validate_test_formula
- _, _, random_effects = _parse_equation(resolved)
+ available_vars = (
+ [self._registry.dependent] + self._registry.non_factor_names + self._registry.factor_names + self._registry.cluster_names
+ )
+ validation = _validate_test_formula(test_formula, available_vars)
+ validation.raise_if_invalid()
+
+ # Parse test formula to get effects and random effects
+ from .utils.test_formula_utils import _extract_test_formula_effects
+
+ test_effects, random_effects = _extract_test_formula_effects(test_formula, self._registry)
+
+ if not test_effects:
+ raise ValueError(f"test_formula '{test_formula}' contains no testable effects from the data generation model.")
+
+ # Check for OLS -> LME cross (invalid: no cluster data to fit)
+ if random_effects and not self._registry._cluster_specs:
+ grouping_vars = [re["grouping_var"] for re in random_effects]
+ raise ValueError(
+ f"test_formula contains random effects ({grouping_vars}) but the "
+ f"data generation model has no cluster structure. Cannot fit a "
+ f"mixed model to data without clusters."
+ )
if random_effects:
self._test_method = "mixed_model"
else:
self._test_method = "linear_regression"
- return resolved
+ return test_formula, test_effects, random_effects
def _run_find_power(
self,
@@ -2101,6 +2170,8 @@ def _run_find_power(
correction,
scenario_config=None,
test_formula=None,
+ test_formula_effects=None,
+ test_random_effects=None,
progress=None,
cancel_check=None,
):
@@ -2109,13 +2180,15 @@ def _run_find_power(
self._validate_cluster_sample_size(sample_size)
# Route based on test method (routing logic handled in simulation.py)
- metadata = self._prepare_metadata(target_tests, correction)
+ metadata = self._prepare_metadata(target_tests, correction, test_formula_effects)
- if scenario_config:
- metadata.heterogeneity = scenario_config["heterogeneity"]
- metadata.heteroskedasticity = scenario_config["heteroskedasticity"]
- if metadata.cluster_specs:
- metadata.lme_scenario_config = scenario_config
+ # Set the random effects flag for test formula
+ if test_random_effects:
+ metadata.test_has_random_effects = True
+
+ # scenario_config is always a dict (SCENARIO_ZERO or user-provided)
+ metadata.heterogeneity = scenario_config["heterogeneity"]
+ metadata.heteroskedasticity = scenario_config["heteroskedasticity"]
runner = SimulationRunner(
n_simulations=self._effective_n_simulations,
@@ -2127,9 +2200,15 @@ def _run_find_power(
)
# Compute critical values once before the simulation loop
- p = len(metadata.effect_sizes)
+ # Use test formula's effect count for critical values when subsetting
+ if metadata.test_column_indices is not None:
+ p = metadata.test_effect_count
+ n_targets = len(metadata.test_target_indices)
+ else:
+ p = len(metadata.effect_sizes)
+ n_targets = len(metadata.target_indices)
+
dof = sample_size - p - 1
- n_targets = len(metadata.target_indices)
n_posthoc = len(metadata.posthoc_specs)
if n_posthoc > 0 and metadata.posthoc_method == "t-test":
@@ -2185,7 +2264,7 @@ def analyze_func(X, y, indices, alpha, correction):
analyze_func=analyze_func,
create_X_extended_func=self._create_X_extended,
scenario_config=scenario_config,
- apply_perturbations_func=(apply_per_simulation_perturbations if scenario_config else None),
+ apply_perturbations_func=apply_per_simulation_perturbations,
progress=progress,
cancel_check=cancel_check,
)
@@ -2193,11 +2272,17 @@ def analyze_func(X, y, indices, alpha, correction):
if not sim_results:
return {}
+ # When test formula is active, filter target_tests to only effects in the test model
+ effective_target_tests = target_tests
+ if test_formula_effects is not None:
+ test_effect_set = set(test_formula_effects)
+ effective_target_tests = [t for t in target_tests if t == "overall" or t in test_effect_set]
+
processor = ResultsProcessor(target_power=self.power)
power_results = processor.calculate_powers(
sim_results["all_results"],
sim_results["all_results_corrected"],
- target_tests,
+ effective_target_tests,
)
# Add n_simulations_failed to power_results
@@ -2207,13 +2292,13 @@ def analyze_func(X, y, indices, alpha, correction):
# Tukey correction only applies to pairwise contrasts; NaN-ify others
if correction and correction.lower() == "tukey" and power_results.get("individual_powers_corrected"):
posthoc_labels = {s.label for s in self._posthoc_specs}
- for test in target_tests:
+ for test in effective_target_tests:
if test not in posthoc_labels:
power_results["individual_powers_corrected"][test] = float("nan")
return build_power_result(
model_type=self.model_type,
- target_tests=target_tests,
+ target_tests=effective_target_tests,
formula_to_test=test_formula,
equation=self.equation,
sample_size=sample_size,
@@ -2243,12 +2328,15 @@ def _run_sample_size_analysis(
correction,
scenario_config=None,
test_formula=None,
+ test_formula_effects=None,
+ test_random_effects=None,
progress=None,
cancel_check=None,
):
"""Iterate over sample sizes, running power analysis for each."""
from .progress import SimulationCancelled
+ use_sequential = True
if self._is_parallel_effective():
from joblib import Parallel, delayed
@@ -2258,7 +2346,18 @@ def _run_sample_size_analysis(
backend="loky",
verbose=0,
return_as="generator",
- )(delayed(self._run_find_power)(ss, target_tests, correction, scenario_config, test_formula) for ss in sample_sizes)
+ )(
+ delayed(self._run_find_power)(
+ ss,
+ target_tests,
+ correction,
+ scenario_config,
+ test_formula,
+ test_formula_effects,
+ test_random_effects,
+ )
+ for ss in sample_sizes
+ )
results = []
for ss, result in zip(sample_sizes, power_results, strict=False):
if cancel_check is not None and cancel_check():
@@ -2266,25 +2365,13 @@ def _run_sample_size_analysis(
results.append((ss, result))
if progress is not None:
progress.advance(self._effective_n_simulations)
+ use_sequential = False
except Exception as e:
if isinstance(e, SimulationCancelled):
raise
print(f"Warning: Parallel execution failed ({e}). Falling back to sequential.")
- results = []
- for ss in sample_sizes:
- if cancel_check is not None and cancel_check():
- raise SimulationCancelled("Simulation cancelled by user") from None
- result = self._run_find_power(
- ss,
- target_tests,
- correction,
- scenario_config,
- test_formula,
- progress=progress,
- cancel_check=cancel_check,
- )
- results.append((ss, result))
- else:
+
+ if use_sequential:
results = []
for sample_size in sample_sizes:
if cancel_check is not None and cancel_check():
@@ -2295,19 +2382,28 @@ def _run_sample_size_analysis(
correction,
scenario_config,
test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=progress,
cancel_check=cancel_check,
)
results.append((sample_size, power_result))
processor = ResultsProcessor(target_power=self.power)
- analysis_results = processor.process_sample_size_results(results, target_tests, correction)
+ # Filter target_tests to match test formula effects
+ if test_formula_effects is not None:
+ test_set = set(test_formula_effects)
+ effective_target_tests = [t for t in target_tests if t in test_set or t == "overall"]
+ else:
+ effective_target_tests = target_tests
+
+ analysis_results = processor.process_sample_size_results(results, effective_target_tests, correction)
# Tukey correction only applies to pairwise contrasts; NaN-ify others
if correction and correction.lower() == "tukey":
posthoc_labels = {s.label for s in self._posthoc_specs}
if analysis_results.get("powers_by_test_corrected"):
- for test in target_tests:
+ for test in effective_target_tests:
if test not in posthoc_labels:
n_points = len(analysis_results["powers_by_test_corrected"][test])
analysis_results["powers_by_test_corrected"][test] = [float("nan")] * n_points
@@ -2315,7 +2411,7 @@ def _run_sample_size_analysis(
return build_sample_size_result(
model_type=self.model_type,
- target_tests=target_tests,
+ target_tests=effective_target_tests,
formula_to_test=test_formula,
equation=self.equation,
sample_sizes=sample_sizes,
@@ -2331,9 +2427,16 @@ def _run_scenario_analysis(self, analysis_type, **kwargs):
"""Delegate to ScenarioRunner for multi-scenario power or sample-size analysis."""
from functools import partial
- configs = self._scenario_configs or DEFAULT_SCENARIO_CONFIG
+ all_configs = self._scenario_configs or DEFAULT_SCENARIO_CONFIG
+ scenario_filter = kwargs.pop("scenario_filter", None)
+ if scenario_filter is not None:
+ configs = {k: all_configs[k] for k in scenario_filter}
+ else:
+ configs = all_configs
scenario_runner = ScenarioRunner(self, configs)
test_formula = kwargs.get("test_formula")
+ test_formula_effects = kwargs.get("test_formula_effects")
+ test_random_effects = kwargs.get("test_random_effects")
progress = kwargs.get("progress")
cancel_check = kwargs.get("cancel_check")
@@ -2341,6 +2444,8 @@ def _run_scenario_analysis(self, analysis_type, **kwargs):
run_power_func = partial(
self._run_find_power,
test_formula=test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=progress,
cancel_check=cancel_check,
)
@@ -2357,6 +2462,8 @@ def _run_scenario_analysis(self, analysis_type, **kwargs):
run_ss_func = partial(
self._run_sample_size_analysis,
test_formula=test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=progress,
cancel_check=cancel_check,
)
diff --git a/mcpower/progress.py b/mcpower/progress.py
index e733148..dca3c25 100644
--- a/mcpower/progress.py
+++ b/mcpower/progress.py
@@ -87,10 +87,7 @@ def __init__(self, **tqdm_kwargs):
self._bar = None
def __call__(self, current: int, total: int):
- try:
- from tqdm import tqdm
- except ImportError:
- raise ImportError("tqdm is required for TqdmReporter. Install with: pip install tqdm") from None
+ from tqdm import tqdm
if self._bar is None:
self._bar = tqdm(total=total, unit="sim", **self._tqdm_kwargs)
diff --git a/mcpower/stats/data_generation.py b/mcpower/stats/data_generation.py
index 0d8800c..3c46d89 100644
--- a/mcpower/stats/data_generation.py
+++ b/mcpower/stats/data_generation.py
@@ -23,7 +23,6 @@
SKEW_STD = np.sqrt(np.exp(2) - np.exp(1))
NORM_SCALE = (DIST_RESOLUTION - 1) / (NORM_RANGE[1] - NORM_RANGE[0])
PERC_SCALE = (DIST_RESOLUTION - 1) / (PERCENTILE_RANGE[1] - PERCENTILE_RANGE[0])
-FLOAT_NEAR_ZERO = 1e-15
# Global lookup tables
NORM_CDF_TABLE = None
@@ -58,13 +57,12 @@ def _compute_t3_sd():
Replicates the vectorised norm-CDF -> t(3)-PPF lookup chain on a large
fixed-seed sample to get a stable SD estimate.
"""
- assert NORM_CDF_TABLE is not None
- assert T3_PPF_TABLE is not None
+ if NORM_CDF_TABLE is None or T3_PPF_TABLE is None:
+ raise RuntimeError("Distribution tables not initialized — _init_tables() must be called first")
- rng_state = np.random.get_state()
- np.random.seed(999999)
- z = np.random.standard_normal(200000)
- np.random.set_state(rng_state)
+ # Use a local RNG to avoid affecting the global state and to be thread-safe.
+ rng = np.random.RandomState(999999)
+ z = rng.standard_normal(200000)
# Step 1: Normal CDF lookup (z -> percentile)
z_clipped = np.clip(z, NORM_RANGE[0], NORM_RANGE[1])
@@ -107,9 +105,16 @@ def create_uploaded_lookup_tables(
for var_idx in range(n_vars):
data = data_matrix[:, var_idx]
- normalized = (data - np.mean(data)) / np.std(data)
+ std = np.std(data)
+ if std < 1e-15:
+ raise ValueError(
+ f"Variable at index {var_idx} has zero variance (constant value). Remove it from the model or check your data."
+ )
+ normalized = (data - np.mean(data)) / std
sorted_uploaded = np.sort(normalized)
+ # Weibull plotting positions: i/(n+1) avoids 0 and 1, which would map
+ # to -inf/+inf under the normal PPF, giving well-behaved quantiles.
percentiles = np.linspace(1 / (n_samples + 1), n_samples / (n_samples + 1), n_samples)
normal_quantiles = norm_ppf_array(percentiles)
@@ -126,13 +131,12 @@ def _generate_factors(sample_size, factor_specs, seed):
Args:
sample_size: Number of observations
factor_specs: List of {'n_levels': int, 'proportions': [float, ...]}
- seed: Random seed
+ seed: Random seed (callers pass sim_seed + 3)
Returns:
X_factors: (sample_size, total_dummies) array
"""
- if seed is not None:
- np.random.seed(seed)
+ rng = np.random.RandomState(seed)
if not factor_specs:
return np.empty((sample_size, 0), dtype=float)
@@ -141,7 +145,7 @@ def _generate_factors(sample_size, factor_specs, seed):
for spec in factor_specs:
n_levels = spec["n_levels"]
proportions = spec["proportions"]
- factor_data = np.random.choice(n_levels, size=sample_size, p=proportions)
+ factor_data = rng.choice(n_levels, size=sample_size, p=proportions)
dummies = np.eye(n_levels, dtype=float)[factor_data]
factor_columns.append(dummies[:, 1:])
@@ -170,12 +174,11 @@ def bootstrap_uploaded_data(
X_non_factors: Non-factor variables (continuous + binary mapped to 0-1)
X_factors: Factor dummy variables
"""
- if seed is not None:
- np.random.seed(seed)
+ rng = np.random.RandomState(seed)
# Bootstrap whole rows
n_samples = raw_data.shape[0]
- row_indices = np.random.choice(n_samples, size=sample_size, replace=True)
+ row_indices = rng.choice(n_samples, size=sample_size, replace=True)
bootstrapped_data = raw_data[row_indices, :]
# Separate by type
@@ -286,12 +289,17 @@ def _generate_cluster_effects(
Returns:
X_cluster: (sample_size, n_cluster_vars) array of random effect columns
"""
- if sim_seed is not None:
- # Use a derived seed to avoid collision with X generation seed
- np.random.seed(sim_seed + 3)
+ rng = np.random.RandomState(sim_seed + 4 if sim_seed is not None else None)
columns = []
+ # Extract perturbation defaults once
+ perturb = lme_perturbations or {}
+ tau_mults = perturb.get("tau_squared_multipliers", {})
+ re_dist_val = perturb.get("random_effect_dist", "normal")
+ re_df_val = perturb.get("random_effect_df", 5)
+ has_perturb = lme_perturbations is not None
+
for gv, spec in cluster_specs.items():
n_clusters = spec["n_clusters"]
cluster_size = spec["cluster_size"]
@@ -302,19 +310,16 @@ def _generate_cluster_effects(
cluster_size = sample_size // n_clusters
# Apply LME perturbations if present
- if lme_perturbations is not None:
- multiplier = lme_perturbations["tau_squared_multipliers"].get(gv, 1.0)
- tau_sq = tau_sq * multiplier
+ if has_perturb:
+ tau_sq = tau_sq * tau_mults.get(gv, 1.0)
tau = np.sqrt(tau_sq)
# Generate random intercepts (possibly non-normal)
- if lme_perturbations is not None:
- re_dist = lme_perturbations.get("random_effect_dist", "normal")
- re_df = lme_perturbations.get("random_effect_df", 5)
- random_intercepts = _generate_non_normal_intercepts(n_clusters, tau, re_dist, re_df)
+ if has_perturb:
+ random_intercepts = _generate_non_normal_intercepts(n_clusters, tau, re_dist_val, re_df_val, rng_state=rng)
else:
- random_intercepts = np.random.normal(0, tau, size=n_clusters)
+ random_intercepts = rng.normal(0, tau, size=n_clusters)
# Create id_effect column: repeat each cluster's intercept
# cluster_id assignment: [0,0,...,0, 1,1,...,1, ..., K-1,K-1,...,K-1]
@@ -413,14 +418,20 @@ def _generate_random_effects(
A :class:`RandomEffectsResult` with intercept columns, slope
contributions, cluster IDs, Z matrices, and nesting metadata.
"""
- if sim_seed is not None:
- np.random.seed(sim_seed + 3)
+ rng = np.random.RandomState(sim_seed + 4 if sim_seed is not None else None)
intercept_cols: List[np.ndarray] = []
slope_contribution = np.zeros(sample_size)
cluster_ids_dict: Dict[str, np.ndarray] = {}
Z_matrices: Dict[str, np.ndarray] = {}
+ # Extract perturbation defaults once (avoids repeated dict lookups)
+ perturb = lme_perturbations or {}
+ tau_multipliers = perturb.get("tau_squared_multipliers", {})
+ re_dist = perturb.get("random_effect_dist", "normal")
+ re_df = perturb.get("random_effect_df", 5)
+ has_perturbations = lme_perturbations is not None
+
# Nested model bookkeeping
child_to_parent: Optional[np.ndarray] = None
K_parent = 0
@@ -460,19 +471,16 @@ def _generate_random_effects(
# Apply LME perturbations: ICC jitter on tau_squared
tau_sq = spec["tau_squared"]
- if lme_perturbations is not None:
- multiplier = lme_perturbations["tau_squared_multipliers"].get(gv, 1.0)
- tau_sq = tau_sq * multiplier
+ if has_perturbations:
+ tau_sq = tau_sq * tau_multipliers.get(gv, 1.0)
if q == 1:
# --- Random intercept only ---
tau = np.sqrt(tau_sq)
- if lme_perturbations is not None:
- re_dist = lme_perturbations.get("random_effect_dist", "normal")
- re_df = lme_perturbations.get("random_effect_df", 5)
- random_intercepts = _generate_non_normal_intercepts(n_clusters, tau, re_dist, re_df)
+ if has_perturbations:
+ random_intercepts = _generate_non_normal_intercepts(n_clusters, tau, re_dist, re_df, rng_state=rng)
else:
- random_intercepts = np.random.normal(0, tau, size=n_clusters)
+ random_intercepts = rng.normal(0, tau, size=n_clusters)
id_effect = _trim_or_pad(np.repeat(random_intercepts, cluster_size), sample_size)
intercept_cols.append(id_effect)
@@ -482,33 +490,29 @@ def _generate_random_effects(
slope_vars = spec.get("random_slope_vars", [])
# Apply ICC jitter to G_matrix intercept variance
- if lme_perturbations is not None:
+ if has_perturbations:
ratio = tau_sq / spec["tau_squared"] if spec["tau_squared"] > 0 else 1.0
# Scale intercept row/column of G by sqrt(ratio)
sqrt_ratio = np.sqrt(ratio)
G_matrix[0, :] *= sqrt_ratio
G_matrix[:, 0] *= sqrt_ratio
- # Draw correlated [b_int, b_slope1, ...] per cluster
- re_dist = lme_perturbations.get("random_effect_dist", "normal") if lme_perturbations else "normal"
- re_df = lme_perturbations.get("random_effect_df", 5) if lme_perturbations else 5
-
- if re_dist == "heavy_tailed" and lme_perturbations is not None:
+ if re_dist == "heavy_tailed" and has_perturbations:
# Multivariate t: MVN(0, G * (df-2)/df) × sqrt(df / chi2(df))
df = max(re_df, 3)
G_scaled = G_matrix * ((df - 2.0) / df)
- b_normal = np.random.multivariate_normal(np.zeros(q), G_scaled, size=n_clusters)
- chi2_samples = np.random.chisquare(df, size=n_clusters)
+ b_normal = rng.multivariate_normal(np.zeros(q), G_scaled, size=n_clusters)
+ chi2_samples = rng.chisquare(df, size=n_clusters)
mixing = np.sqrt(df / chi2_samples)
b = b_normal * mixing[:, np.newaxis]
- elif re_dist == "skewed" and lme_perturbations is not None:
+ elif re_dist == "skewed" and has_perturbations:
# Independent skewed marginals via shifted chi-squared, scaled by Cholesky
df = max(re_df, 3)
L = np.linalg.cholesky(G_matrix)
- raw = (np.random.chisquare(df, size=(n_clusters, q)) - df) / np.sqrt(2 * df)
+ raw = (rng.chisquare(df, size=(n_clusters, q)) - df) / np.sqrt(2 * df)
b = raw @ L.T
else:
- b = np.random.multivariate_normal(np.zeros(q), G_matrix, size=n_clusters)
+ b = rng.multivariate_normal(np.zeros(q), G_matrix, size=n_clusters)
# Intercept component
intercept_effect = _trim_or_pad(np.repeat(b[:, 0], cluster_size), sample_size)
@@ -549,21 +553,19 @@ def _generate_random_effects(
tau_sq_parent = p_spec["tau_squared"]
tau_sq_child = c_spec["tau_squared"]
- if lme_perturbations is not None:
- tau_sq_parent *= lme_perturbations["tau_squared_multipliers"].get(p_gv, 1.0)
- tau_sq_child *= lme_perturbations["tau_squared_multipliers"].get(c_gv, 1.0)
+ if has_perturbations:
+ tau_sq_parent *= tau_multipliers.get(p_gv, 1.0)
+ tau_sq_child *= tau_multipliers.get(c_gv, 1.0)
tau_parent = np.sqrt(tau_sq_parent)
tau_child = np.sqrt(tau_sq_child)
- if lme_perturbations is not None:
- re_dist = lme_perturbations.get("random_effect_dist", "normal")
- re_df = lme_perturbations.get("random_effect_df", 5)
- b_parent = _generate_non_normal_intercepts(K_parent, tau_parent, re_dist, re_df)
- b_child = _generate_non_normal_intercepts(K_child, tau_child, re_dist, re_df)
+ if has_perturbations:
+ b_parent = _generate_non_normal_intercepts(K_parent, tau_parent, re_dist, re_df, rng_state=rng)
+ b_child = _generate_non_normal_intercepts(K_child, tau_child, re_dist, re_df, rng_state=rng)
else:
- b_parent = np.random.normal(0, tau_parent, size=K_parent)
- b_child = np.random.normal(0, tau_child, size=K_child)
+ b_parent = rng.normal(0, tau_parent, size=K_parent)
+ b_child = rng.normal(0, tau_child, size=K_child)
# IDs: parent_ids assigns each observation to a parent cluster,
# child_ids assigns each observation to a child cluster.
diff --git a/mcpower/stats/distributions.py b/mcpower/stats/distributions.py
index cf13bca..6ee5265 100644
--- a/mcpower/stats/distributions.py
+++ b/mcpower/stats/distributions.py
@@ -3,9 +3,7 @@
Provides F, t, chi2, normal, and studentized range distribution
functions plus batch critical-value computation and table generation.
-Backend priority:
- 1. C++ native (Boost.Math + R Tukey port) via mcpower_native
- 2. scipy (optional shim, for when C++ is not compiled)
+All functions are provided by the C++ native backend (Boost.Math + R Tukey port).
Usage:
from mcpower.stats.distributions import norm_ppf, compute_critical_values_ols
@@ -14,13 +12,11 @@
import numpy as np
# ============================================================================
-# Backend selection
+# Backend — native C++ only
# ============================================================================
-_BACKEND = None
-
try:
- from mcpower.backends.mcpower_native import ( # type: ignore[import]
+ from mcpower.backends.mcpower_native import ( # type: ignore[import] # noqa: F401
chi2_cdf,
chi2_ppf,
compute_critical_values_lme,
@@ -36,171 +32,17 @@
t_ppf,
)
- _BACKEND = "native"
-
-except ImportError:
- # -------------------------------------------------------------------
- # scipy shim -- temporary fallback for when C++ is not compiled.
- # Will be removed when Python fallback backends are fully dropped.
- # -------------------------------------------------------------------
- try:
- from scipy.stats import ( # isort: skip
- chi2 as _chi2_dist,
- f as _f_dist,
- norm as _norm_dist,
- studentized_range as _sr_dist,
- t as _t_dist,
- )
-
- def norm_ppf(p): # noqa: F811
- """Standard normal quantile function (inverse CDF)."""
- return float(_norm_dist.ppf(p))
-
- def norm_cdf(x): # noqa: F811
- """Standard normal CDF."""
- return float(_norm_dist.cdf(x))
-
- def t_ppf(p, df): # noqa: F811
- """Student's t quantile function."""
- return float(_t_dist.ppf(p, df))
-
- def f_ppf(p, dfn, dfd): # noqa: F811
- """Fisher F quantile function."""
- return float(_f_dist.ppf(p, dfn, dfd))
-
- def chi2_ppf(p, df): # noqa: F811
- """Chi-squared quantile function."""
- return float(_chi2_dist.ppf(p, df))
-
- def chi2_cdf(x, df): # noqa: F811
- """Chi-squared CDF."""
- return float(_chi2_dist.cdf(x, df))
-
- def studentized_range_ppf(p, k, df): # noqa: F811
- """Studentized range quantile (Tukey). k=groups, df=denom df."""
- if df < 2 or k < 2 or k > 200 or p <= 0.0 or p >= 1.0:
- return float("inf")
- return float(_sr_dist.ppf(p, k, df))
-
- def compute_critical_values_ols(alpha, dfn, dfd, n_targets, correction_method): # noqa: F811
- """Compute OLS critical values using scipy (fallback).
-
- Args:
- alpha: Significance level.
- dfn: Numerator degrees of freedom (number of predictors).
- dfd: Denominator degrees of freedom (n - p - 1).
- n_targets: Number of individual effects being tested.
- correction_method: 0=none, 1=Bonferroni, 2=FDR (BH), 3=Holm.
-
- Returns:
- Tuple of (f_crit, t_crit, correction_t_crits) where
- correction_t_crits is an ndarray of length n_targets.
- """
- if dfd <= 0:
- return np.inf, np.inf, np.full(max(n_targets, 1), np.inf)
-
- f_crit = _f_dist.ppf(1 - alpha, dfn, dfd) if dfn > 0 else np.inf
- t_crit = _t_dist.ppf(1 - alpha / 2, dfd)
-
- m = n_targets
- if m == 0:
- return f_crit, t_crit, np.empty(0)
-
- if correction_method == 0: # None
- correction_t_crits = np.full(m, t_crit)
- elif correction_method == 1: # Bonferroni
- bonf_crit = _t_dist.ppf(1 - alpha / (2 * m), dfd)
- correction_t_crits = np.full(m, bonf_crit)
- elif correction_method == 2: # FDR (Benjamini-Hochberg)
- correction_t_crits = np.array(
- [_t_dist.ppf(1 - (k + 1) / m * alpha / 2, dfd) if (k + 1) / m * alpha / 2 >= 1e-12 else np.inf for k in range(m)]
- )
- elif correction_method == 3: # Holm
- correction_t_crits = np.array(
- [_t_dist.ppf(1 - alpha / (2 * (m - k)), dfd) if alpha / (2 * (m - k)) >= 1e-12 else np.inf for k in range(m)]
- )
- else:
- correction_t_crits = np.full(m, t_crit)
-
- return f_crit, t_crit, correction_t_crits
-
- def compute_tukey_critical_value(alpha, n_levels, dfd): # noqa: F811
- """Compute Tukey HSD critical value (q / sqrt(2))."""
- if dfd <= 0:
- return np.inf
- q_crit = _sr_dist.ppf(1 - alpha, n_levels, dfd)
- return q_crit / np.sqrt(2)
-
- def compute_critical_values_lme(alpha, n_fixed, n_targets, correction_method): # noqa: F811
- """Compute LME critical values using scipy (fallback).
-
- Args:
- alpha: Significance level.
- n_fixed: Number of fixed effects (excluding intercept).
- n_targets: Number of individual effects being tested.
- correction_method: 0=none, 1=Bonferroni, 2=FDR (BH), 3=Holm.
-
- Returns:
- Tuple of (chi2_crit, z_crit, correction_z_crits) where
- correction_z_crits is an ndarray of length n_targets.
- """
- chi2_crit = _chi2_dist.ppf(1 - alpha, n_fixed) if n_fixed > 0 else np.inf
- z_crit = _norm_dist.ppf(1 - alpha / 2)
-
- m = n_targets
- if m == 0:
- return chi2_crit, z_crit, np.empty(0)
-
- if correction_method == 0: # None
- correction_z_crits = np.full(m, z_crit)
- elif correction_method == 1: # Bonferroni
- bonf = _norm_dist.ppf(1 - alpha / (2 * m))
- correction_z_crits = np.full(m, bonf)
- elif correction_method == 2: # FDR (Benjamini-Hochberg)
- correction_z_crits = np.array(
- [_norm_dist.ppf(1 - (k + 1) / m * alpha / 2) if (k + 1) / m * alpha / 2 >= 1e-12 else np.inf for k in range(m)]
- )
- elif correction_method == 3: # Holm
- correction_z_crits = np.array(
- [_norm_dist.ppf(1 - alpha / (2 * (m - k))) if alpha / (2 * (m - k)) >= 1e-12 else np.inf for k in range(m)]
- )
- else:
- correction_z_crits = np.full(m, z_crit)
-
- return chi2_crit, z_crit, correction_z_crits
-
- def generate_norm_cdf_table(x_min, x_max, resolution): # noqa: F811
- """Generate normal CDF lookup table."""
- x = np.linspace(x_min, x_max, resolution)
- return _norm_dist.cdf(x).astype(np.float64)
-
- def generate_t3_ppf_table(perc_min, perc_max, resolution): # noqa: F811
- """Generate t(3) PPF lookup table (divided by sqrt(3))."""
- p = np.linspace(perc_min, perc_max, resolution)
- return (_t_dist.ppf(p, 3) / np.sqrt(3)).astype(np.float64)
-
- def norm_ppf_array(percentiles): # noqa: F811
- """Vectorized normal PPF for percentile array."""
- return _norm_dist.ppf(np.asarray(percentiles)).astype(np.float64)
-
- _BACKEND = "scipy"
-
- except ImportError as exc:
- raise ImportError(
- "No distribution backend available. "
- "Install from PyPI for prebuilt C++ wheels: pip install MCPower\n"
- "Or install scipy as fallback: pip install scipy"
- ) from exc
+except ImportError as exc:
+ raise ImportError("Native C++ backend not available. Install from PyPI for prebuilt wheels: pip install MCPower") from exc
# ============================================================================
-# Also re-export scipy optimizer shims for lme_solver.py
-# These replace scipy.optimize.minimize and minimize_scalar
+# Optimizer wrappers for lme_solver.py
# ============================================================================
def minimize_lbfgsb(objective, x0, bounds, maxiter=200, ftol=1e-10, gtol=1e-6):
- """L-BFGS-B minimization -- C++ native or scipy fallback.
+ """L-BFGS-B minimization via native C++ backend.
Args:
objective: Callable f(x) -> float
@@ -213,53 +55,15 @@ def minimize_lbfgsb(objective, x0, bounds, maxiter=200, ftol=1e-10, gtol=1e-6):
Returns:
Object with .x (optimal point), .fun (optimal value), .converged (bool)
"""
- if _BACKEND == "native":
- try:
- from mcpower.backends.mcpower_native import lbfgsb_minimize_fd # type: ignore[import]
-
- lb = np.array([b[0] for b in bounds])
- ub = np.array([b[1] for b in bounds])
- return lbfgsb_minimize_fd(objective, np.asarray(x0, dtype=np.float64), lb, ub, maxiter, ftol, gtol)
- except ImportError:
- import warnings
-
- warnings.warn(
- "Native L-BFGS-B optimizer not available despite native backend being loaded. Falling back to scipy.",
- RuntimeWarning,
- stacklevel=2,
- )
- except Exception as e:
- import warnings
+ from mcpower.backends.mcpower_native import lbfgsb_minimize_fd # type: ignore[import]
- warnings.warn(
- f"Native L-BFGS-B optimizer failed ({type(e).__name__}: {e}), falling back to scipy.",
- RuntimeWarning,
- stacklevel=2,
- )
-
- # scipy fallback
- from scipy.optimize import minimize
-
- result = minimize(
- objective,
- x0,
- method="L-BFGS-B",
- bounds=bounds,
- options={"maxiter": maxiter, "ftol": ftol, "gtol": gtol},
- )
-
- class _Result:
- __slots__ = ("x", "fun", "converged")
-
- r = _Result()
- r.x = result.x
- r.fun = result.fun
- r.converged = result.success
- return r
+ lb = np.array([b[0] for b in bounds])
+ ub = np.array([b[1] for b in bounds])
+ return lbfgsb_minimize_fd(objective, np.asarray(x0, dtype=np.float64), lb, ub, maxiter, ftol, gtol)
def minimize_scalar_brent(objective, bounds, tol=1e-8, maxiter=150):
- """Brent 1D minimization -- C++ native or scipy fallback.
+ """Brent 1D minimization via native C++ backend.
Args:
objective: Callable f(x) -> float
@@ -270,43 +74,6 @@ def minimize_scalar_brent(objective, bounds, tol=1e-8, maxiter=150):
Returns:
Object with .x (optimal point), .fun (optimal value), .converged (bool)
"""
- if _BACKEND == "native":
- try:
- from mcpower.backends.mcpower_native import brent_minimize_scalar # type: ignore[import]
-
- return brent_minimize_scalar(objective, bounds[0], bounds[1], tol, maxiter)
- except ImportError:
- import warnings
-
- warnings.warn(
- "Native Brent optimizer not available despite native backend being loaded. Falling back to scipy.",
- RuntimeWarning,
- stacklevel=2,
- )
- except Exception as e:
- import warnings
-
- warnings.warn(
- f"Native Brent optimizer failed ({type(e).__name__}: {e}), falling back to scipy.",
- RuntimeWarning,
- stacklevel=2,
- )
-
- # scipy fallback
- from scipy.optimize import minimize_scalar
-
- result = minimize_scalar(
- objective,
- bounds=bounds,
- method="bounded",
- options={"xatol": tol, "maxiter": maxiter},
- )
-
- class _Result:
- __slots__ = ("x", "fun", "converged")
+ from mcpower.backends.mcpower_native import brent_minimize_scalar # type: ignore[import]
- r = _Result()
- r.x = result.x
- r.fun = result.fun
- r.converged = bool(getattr(result, "success", True))
- return r
+ return brent_minimize_scalar(objective, bounds[0], bounds[1], tol, maxiter)
diff --git a/mcpower/stats/mixed_models.py b/mcpower/stats/mixed_models.py
index 414e911..07de5e8 100644
--- a/mcpower/stats/mixed_models.py
+++ b/mcpower/stats/mixed_models.py
@@ -11,10 +11,12 @@
import threading
import warnings
-from typing import Any, Dict, List, Optional, Union
+from typing import Any, Dict, Optional, Union
import numpy as np
+from ..backends.native import _prep
+
# Suppress statsmodels convergence warnings (expected with small samples/low ICC).
# Module-level filterwarnings with module= is unreliable for statsmodels internals,
# so we also use catch_warnings() context managers around .fit() calls below.
@@ -30,7 +32,6 @@ def _lme_analysis_wrapper(
y: np.ndarray,
target_indices: np.ndarray,
cluster_ids: np.ndarray,
- cluster_column_indices: List[int],
correction_method: int,
alpha: float,
backend: str = "custom",
@@ -51,7 +52,6 @@ def _lme_analysis_wrapper(
y: (n,) response vector
target_indices: Coefficient indices to test (fixed effects only)
cluster_ids: (n,) cluster membership array [0,0,0, 1,1,1, ...]
- cluster_column_indices: Indices of cluster effect columns (unused)
correction_method: 0=none, 1=Bonferroni, 2=FDR, 3=Holm
alpha: Significance level
backend: "custom" (default) or "statsmodels" (fallback)
@@ -118,9 +118,7 @@ def _lme_analysis_wrapper(
verbose=verbose,
)
elif backend == "statsmodels":
- return _lme_analysis_statsmodels(
- X_expanded, y, target_indices, cluster_ids, cluster_column_indices, correction_method, alpha, verbose
- )
+ return _lme_analysis_statsmodels(X_expanded, y, target_indices, cluster_ids, correction_method, alpha, verbose)
else:
raise ValueError(f"Unknown backend: {backend}")
@@ -130,7 +128,6 @@ def _lme_analysis_statsmodels(
y: np.ndarray,
target_indices: np.ndarray,
cluster_ids: np.ndarray,
- cluster_column_indices: List[int],
correction_method: int,
alpha: float,
verbose: bool = False,
@@ -145,11 +142,10 @@ def _lme_analysis_statsmodels(
- Convergence retry strategy (allows ≤3% failures)
Args:
- X_expanded: (n, p) design matrix (includes cluster effect columns)
+ X_expanded: (n, p) design matrix (excludes cluster effect columns)
y: (n,) response vector
target_indices: Coefficient indices to test (fixed effects only)
cluster_ids: (n,) cluster membership array
- cluster_column_indices: Indices of cluster effect columns to remove
correction_method: 0=none, 1=Bonferroni, 2=FDR, 3=Holm
alpha: Significance level
verbose: Return detailed diagnostics
@@ -172,8 +168,6 @@ def _lme_analysis_statsmodels(
n, p = X_expanded.shape
n_targets = len(target_indices)
- # Note: X_expanded already excludes cluster effects (they're not in the design matrix)
- # cluster_column_indices is now unused in this function but kept for API compatibility
X_fixed = X_expanded
# Step 1: Add intercept to fixed effects
@@ -530,6 +524,29 @@ def _compute_wald_test(result, alpha):
return results_array
+def _ensure_lme_crits(alpha, p, n_targets, correction_method, chi2_crit, z_crit, correction_z_crits):
+ """Compute LME critical values on-the-fly if not precomputed."""
+ if z_crit is None or chi2_crit is None or correction_z_crits is None:
+ from .lme_solver import compute_lme_critical_values
+
+ return compute_lme_critical_values(alpha, p, n_targets, correction_method)
+ return chi2_crit, z_crit, correction_z_crits
+
+
+def _wrap_native_result(result, verbose, solver_name, extra_diag=None) -> Optional[Union[np.ndarray, Dict]]:
+ """Wrap C++ solver result with optional verbose diagnostics."""
+ if len(result) > 0:
+ if verbose:
+ diag = {"solver": solver_name}
+ if extra_diag:
+ diag.update(extra_diag)
+ return {"results": result, "diagnostics": diag}
+ return np.asarray(result)
+ if verbose:
+ return {"results": None, "failure_reason": f"C++ {solver_name} returned empty result"}
+ return None
+
+
def _lme_analysis_custom(
X_expanded: np.ndarray,
y: np.ndarray,
@@ -545,39 +562,29 @@ def _lme_analysis_custom(
"""LME analysis for random-intercept models via C++ backend.
Uses precomputed critical values (chi2_crit, z_crit) to avoid
- per-simulation scipy calls. Falls back to computing them if not provided.
+ per-simulation distribution calls. Falls back to computing them if not provided.
"""
n, p = X_expanded.shape
n_targets = len(target_indices)
K = int(cluster_ids.max()) + 1
- if z_crit is None or chi2_crit is None or correction_z_crits is None:
- from .lme_solver import compute_lme_critical_values
-
- chi2_crit, z_crit, correction_z_crits = compute_lme_critical_values(alpha, p, n_targets, correction_method)
+ chi2_crit, z_crit, correction_z_crits = _ensure_lme_crits(alpha, p, n_targets, correction_method, chi2_crit, z_crit, correction_z_crits)
from mcpower.backends import mcpower_native as _native # type: ignore[attr-defined]
result = _native.lme_analysis(
- np.ascontiguousarray(X_expanded, dtype=np.float64),
- np.ascontiguousarray(y, dtype=np.float64),
- np.ascontiguousarray(cluster_ids, dtype=np.int32),
+ _prep(X_expanded),
+ _prep(y),
+ _prep(cluster_ids, np.int32),
K,
- np.ascontiguousarray(target_indices, dtype=np.int32),
+ _prep(target_indices, np.int32),
float(chi2_crit),
float(z_crit),
- np.ascontiguousarray(correction_z_crits, dtype=np.float64),
+ _prep(correction_z_crits),
int(correction_method),
float(-1.0),
)
- if len(result) > 0:
- if verbose:
- return {"results": result, "diagnostics": {"solver": "native_q1"}}
- return result # type: ignore[no-any-return]
-
- if verbose:
- return {"results": None, "failure_reason": "C++ solver returned empty result"}
- return None
+ return _wrap_native_result(result, verbose, "native_q1")
def _lme_analysis_custom_general(
@@ -594,8 +601,6 @@ def _lme_analysis_custom_general(
verbose: bool = False,
) -> Optional[Union[np.ndarray, Dict]]:
"""LME analysis for random slopes (q > 1) via C++ backend."""
- from .lme_solver import compute_lme_critical_values
-
n, p = X_expanded.shape
n_targets = len(target_indices)
@@ -604,34 +609,26 @@ def _lme_analysis_custom_general(
q = Z.shape[1]
K = int(cluster_ids.max()) + 1
- if z_crit is None or chi2_crit is None or correction_z_crits is None:
- chi2_crit, z_crit, correction_z_crits = compute_lme_critical_values(alpha, p, n_targets, correction_method)
+ chi2_crit, z_crit, correction_z_crits = _ensure_lme_crits(alpha, p, n_targets, correction_method, chi2_crit, z_crit, correction_z_crits)
from mcpower.backends import mcpower_native as _native # type: ignore[attr-defined]
warm_theta_arr = np.empty(0, dtype=np.float64)
result = _native.lme_analysis_general(
- np.ascontiguousarray(X_expanded, dtype=np.float64),
- np.ascontiguousarray(y, dtype=np.float64),
- np.ascontiguousarray(Z, dtype=np.float64),
- np.ascontiguousarray(cluster_ids, dtype=np.int32),
+ _prep(X_expanded),
+ _prep(y),
+ _prep(Z),
+ _prep(cluster_ids, np.int32),
K,
q,
- np.ascontiguousarray(target_indices, dtype=np.int32),
+ _prep(target_indices, np.int32),
float(chi2_crit),
float(z_crit),
- np.ascontiguousarray(correction_z_crits, dtype=np.float64),
+ _prep(correction_z_crits),
int(correction_method),
warm_theta_arr,
)
- if len(result) > 0:
- if verbose:
- return {"results": result, "diagnostics": {"solver": "native_general", "q": q}}
- return result # type: ignore[no-any-return]
-
- if verbose:
- return {"results": None, "failure_reason": "C++ general solver returned empty result"}
- return None
+ return _wrap_native_result(result, verbose, "native_general", extra_diag={"q": q})
def _lme_analysis_custom_nested(
@@ -647,8 +644,6 @@ def _lme_analysis_custom_nested(
verbose: bool = False,
) -> Optional[Union[np.ndarray, Dict]]:
"""LME analysis for nested random intercepts via C++ backend."""
- from .lme_solver import compute_lme_critical_values
-
n, p = X_expanded.shape
n_targets = len(target_indices)
@@ -658,35 +653,27 @@ def _lme_analysis_custom_nested(
K_child = re_result.K_child
child_to_parent = re_result.child_to_parent
- if z_crit is None or chi2_crit is None or correction_z_crits is None:
- chi2_crit, z_crit, correction_z_crits = compute_lme_critical_values(alpha, p, n_targets, correction_method)
+ chi2_crit, z_crit, correction_z_crits = _ensure_lme_crits(alpha, p, n_targets, correction_method, chi2_crit, z_crit, correction_z_crits)
from mcpower.backends import mcpower_native as _native # type: ignore[attr-defined]
warm_theta_arr = np.empty(0, dtype=np.float64)
result = _native.lme_analysis_nested(
- np.ascontiguousarray(X_expanded, dtype=np.float64),
- np.ascontiguousarray(y, dtype=np.float64),
- np.ascontiguousarray(parent_ids, dtype=np.int32),
- np.ascontiguousarray(child_ids, dtype=np.int32),
+ _prep(X_expanded),
+ _prep(y),
+ _prep(parent_ids, np.int32),
+ _prep(child_ids, np.int32),
K_parent,
K_child,
- np.ascontiguousarray(child_to_parent, dtype=np.int32),
- np.ascontiguousarray(target_indices, dtype=np.int32),
+ _prep(child_to_parent, np.int32),
+ _prep(target_indices, np.int32),
float(chi2_crit),
float(z_crit),
- np.ascontiguousarray(correction_z_crits, dtype=np.float64),
+ _prep(correction_z_crits),
int(correction_method),
warm_theta_arr,
)
- if len(result) > 0:
- if verbose:
- return {"results": result, "diagnostics": {"solver": "native_nested", "K_parent": K_parent, "K_child": K_child}}
- return result # type: ignore[no-any-return]
-
- if verbose:
- return {"results": None, "failure_reason": "C++ nested solver returned empty result"}
- return None
+ return _wrap_native_result(result, verbose, "native_nested", extra_diag={"K_parent": K_parent, "K_child": K_child})
def reset_warm_start_cache():
diff --git a/mcpower/tables/lookup.py b/mcpower/tables/lookup.py
index b66fefa..5f4d691 100644
--- a/mcpower/tables/lookup.py
+++ b/mcpower/tables/lookup.py
@@ -15,7 +15,7 @@ class LookupTableManager:
"""Manages pre-computed lookup tables for data-generation transforms.
Tables are lazily loaded from disk (``tables/data/*.npz``) on first
- access and generated from scipy if the cache files are missing.
+ access and generated via the C++ native backend if the cache files are missing.
The C++ native backend consumes these tables for distribution
transforms.
@@ -47,47 +47,37 @@ def ensure_data_dir(self) -> None:
"""Ensure data directory exists."""
self.data_dir.mkdir(parents=True, exist_ok=True)
- def load_norm_cdf_table(self) -> np.ndarray:
- """Load (or generate and cache) the normal CDF lookup table.
+ def _load_table(self, key: str, generate_fn) -> np.ndarray:
+ """Load a table from cache, disk, or generate it on the fly.
+
+ Args:
+ key: Cache key and npz array name (e.g. ``"norm_cdf"``).
+ generate_fn: Bound method to generate and cache the table.
Returns:
1-D float64 array of length ``DIST_RESOLUTION``.
"""
- if "norm_cdf" in self._tables:
- return self._tables["norm_cdf"]
-
- cache_file = self.data_dir / "norm_cdf.npz"
+ if key in self._tables:
+ return self._tables[key]
+ cache_file = self.data_dir / f"{key}.npz"
try:
data = np.load(cache_file)
- self._tables["norm_cdf"] = data["norm_cdf"]
- return self._tables["norm_cdf"]
+ self._tables[key] = data[key]
+ return self._tables[key]
except (FileNotFoundError, KeyError):
pass
- self._generate_norm_cdf_table()
- return self._tables["norm_cdf"]
-
- def load_t3_ppf_table(self) -> np.ndarray:
- """Load (or generate and cache) the t(df=3) PPF lookup table.
+ generate_fn()
+ return self._tables[key]
- Returns:
- 1-D float64 array of length ``DIST_RESOLUTION``.
- """
- if "t3_ppf" in self._tables:
- return self._tables["t3_ppf"]
-
- cache_file = self.data_dir / "t3_ppf.npz"
-
- try:
- data = np.load(cache_file)
- self._tables["t3_ppf"] = data["t3_ppf"]
- return self._tables["t3_ppf"]
- except (FileNotFoundError, KeyError):
- pass
+ def load_norm_cdf_table(self) -> np.ndarray:
+ """Load (or generate and cache) the normal CDF lookup table."""
+ return self._load_table("norm_cdf", self._generate_norm_cdf_table)
- self._generate_t3_ppf_table()
- return self._tables["t3_ppf"]
+ def load_t3_ppf_table(self) -> np.ndarray:
+ """Load (or generate and cache) the t(df=3) PPF lookup table."""
+ return self._load_table("t3_ppf", self._generate_t3_ppf_table)
def load_all_generation_tables(self) -> Tuple[np.ndarray, np.ndarray]:
"""
@@ -110,6 +100,8 @@ def _generate_norm_cdf_table(self) -> None:
self._tables["norm_cdf"] = norm_cdf
self.ensure_data_dir()
+ # Silently ignore cache write failures (e.g. read-only filesystem,
+ # permission denied). Tables are still usable from memory.
try:
np.savez_compressed(self.data_dir / "norm_cdf.npz", norm_cdf=norm_cdf, x_range=x_norm)
except Exception:
@@ -125,6 +117,8 @@ def _generate_t3_ppf_table(self) -> None:
self._tables["t3_ppf"] = t3_ppf
self.ensure_data_dir()
+ # Silently ignore cache write failures (e.g. read-only filesystem,
+ # permission denied). Tables are still usable from memory.
try:
np.savez_compressed(
self.data_dir / "t3_ppf.npz",
diff --git a/mcpower/utils/formatters.py b/mcpower/utils/formatters.py
index b8bd0b0..8bc91b3 100644
--- a/mcpower/utils/formatters.py
+++ b/mcpower/utils/formatters.py
@@ -6,6 +6,7 @@
"""
import math
+from itertools import combinations
from typing import Any, Dict, List, Optional
import numpy as np
@@ -13,6 +14,11 @@
__all__ = []
+def _is_nan(value) -> bool:
+ """Check if a value is NaN (float type check + math.isnan)."""
+ return isinstance(value, float) and math.isnan(value)
+
+
class _TableFormatter:
"""Static helpers for building fixed-width text tables."""
@@ -25,7 +31,10 @@ def _create_table(
"""Create formatted table with headers and rows."""
if not col_widths:
- col_widths = [max(len(str(h)), max(len(str(row[i])) + 2 for row in rows)) for i, h in enumerate(headers)]
+ if rows:
+ col_widths = [max(len(str(h)), max(len(str(row[i])) + 2 for row in rows)) for i, h in enumerate(headers)]
+ else:
+ col_widths = [len(str(h)) for h in headers]
lines = []
@@ -131,7 +140,7 @@ def _format_short_power(self, data: Dict) -> str:
for test in model["target_tests"]:
power_corr = results["individual_powers_corrected"][test]
- if isinstance(power_corr, float) and math.isnan(power_corr):
+ if _is_nan(power_corr):
rows_corrected.append([test, "-", f"{target:.0f}", "-"])
else:
status = "✓" if power_corr >= target else "✗"
@@ -162,7 +171,7 @@ def _format_long_power(self, data: Dict) -> str:
power = results["individual_powers"][test]
power_corr = results.get("individual_powers_corrected", {}).get(test, power)
target = model.get("target_power", 80.0)
- if isinstance(power_corr, float) and math.isnan(power_corr):
+ if _is_nan(power_corr):
rows.append([test, f"{power:.2f}", "-", f"{target:.1f}", "-"])
else:
achieved = "✓" if power_corr >= target else "✗"
@@ -331,13 +340,15 @@ def _format_scenario_power_short(self, scenarios: Dict, target_tests: List[str],
lines = [f"\n{'=' * 80}", "SCENARIO SUMMARY", f"{'=' * 80}"]
+ scenario_names = list(scenarios.keys())
+ headers = ["Test"] + [name.title() for name in scenario_names]
+ col_widths = [40] + [12] * len(scenario_names)
+
# Uncorrected table
- headers = ["Test", "Optimistic", "Realistic", "Doomer"]
rows = []
-
for test in target_tests:
row = [test]
- for scenario in ["optimistic", "realistic", "doomer"]:
+ for scenario in scenario_names:
if scenario in scenarios and "results" in scenarios[scenario]:
power = scenarios[scenario]["results"]["individual_powers"][test]
row.append(f"{power:.1f}")
@@ -346,17 +357,17 @@ def _format_scenario_power_short(self, scenarios: Dict, target_tests: List[str],
rows.append(row)
lines.append("\nUncorrected Power:")
- lines.append(self._table._create_table(headers, rows, [40, 12, 12, 12]))
+ lines.append(self._table._create_table(headers, rows, col_widths))
# Corrected table if applicable
if correction:
rows_corr = []
for test in target_tests:
row = [test]
- for scenario in ["optimistic", "realistic", "doomer"]:
+ for scenario in scenario_names:
if scenario in scenarios and "results" in scenarios[scenario]:
power_corr = scenarios[scenario]["results"]["individual_powers_corrected"][test]
- if isinstance(power_corr, float) and math.isnan(power_corr):
+ if _is_nan(power_corr):
row.append("-")
else:
row.append(f"{power_corr:.1f}")
@@ -365,7 +376,7 @@ def _format_scenario_power_short(self, scenarios: Dict, target_tests: List[str],
rows_corr.append(row)
lines.append(f"\nCorrected Power ({correction}):")
- lines.append(self._table._create_table(headers, rows_corr, [40, 12, 12, 12]))
+ lines.append(self._table._create_table(headers, rows_corr, col_widths))
lines.append(f"{'=' * 80}")
@@ -395,74 +406,74 @@ def _format_scenario_power_long(
lines.append("DETAILED SCENARIO RESULTS")
lines.append(f"{'=' * 80}")
- for scenario_name in ["optimistic", "realistic", "doomer"]:
- if scenario_name in scenarios:
- lines.append(f"\n{'-' * 80}")
- lines.append(f"{scenario_name.upper()} SCENARIO")
- lines.append(f"{'-' * 80}")
-
- # Use regular power formatter for each scenario
- scenario_data = {
- "model": scenarios[scenario_name]["model"],
- "results": scenarios[scenario_name]["results"],
- }
- lines.append(self._format_long_power(scenario_data))
-
- # 3. Comparison analysis
- lines.append(f"\n{'=' * 80}")
- lines.append("ROBUSTNESS ANALYSIS")
- lines.append(f"{'=' * 80}")
-
- # Power reduction table
- headers = ["Test", "Opt→Real Drop", "Opt→Doom Drop", "Vulnerability"]
- rows = []
- vulnerable_tests = []
- inflated_tests = []
+ for scenario_name in scenarios:
+ lines.append(f"\n{'-' * 80}")
+ lines.append(f"{scenario_name.upper()} SCENARIO")
+ lines.append(f"{'-' * 80}")
+
+ scenario_data = {
+ "model": scenarios[scenario_name]["model"],
+ "results": scenarios[scenario_name]["results"],
+ }
+ lines.append(self._format_long_power(scenario_data))
+
+ # 3. Comparison analysis — compare each non-optimistic scenario to optimistic
+ if "optimistic" in scenarios and len(scenarios) > 1:
+ lines.append(f"\n{'=' * 80}")
+ lines.append("ROBUSTNESS ANALYSIS")
+ lines.append(f"{'=' * 80}")
+
+ other_scenarios = [s for s in scenarios if s != "optimistic"]
+ headers = ["Test"] + [f"Opt→{s.title()} Drop" for s in other_scenarios] + ["Vulnerability"]
+ rows = []
+ vulnerable_tests = []
+ inflated_tests = []
- for test in target_tests:
- opt_power = scenarios["optimistic"]["results"]["individual_powers"][test]
- real_power = scenarios.get("realistic", {}).get("results", {}).get("individual_powers", {}).get(test, opt_power)
- doom_power = scenarios.get("doomer", {}).get("results", {}).get("individual_powers", {}).get(test, opt_power)
-
- real_drop = opt_power - real_power
- doom_drop = opt_power - doom_power
-
- # Format drops with proper signs
- real_drop_str = f"+{abs(real_drop):.1f}%" if real_drop < 0 else f"-{real_drop:.1f}%"
- doom_drop_str = f"+{abs(doom_drop):.1f}%" if doom_drop < 0 else f"-{doom_drop:.1f}%"
-
- # Vulnerability assessment and categorization
- if doom_drop > HIGH_VULNERABILITY_THRESHOLD:
- vulnerability = "HIGH"
- vulnerable_tests.append(test)
- elif doom_drop > MEDIUM_VULNERABILITY_THRESHOLD:
- vulnerability = "MEDIUM"
- elif doom_drop < INFLATED_ERROR_THRESHOLD:
- vulnerability = "INFLATED FALSE POSITIVES"
- inflated_tests.append(test)
- else:
- vulnerability = "LOW"
+ for test in target_tests:
+ opt_power = scenarios["optimistic"]["results"]["individual_powers"][test]
+ row = [test]
+ max_drop = 0.0
+
+ for scenario in other_scenarios:
+ other_power = scenarios.get(scenario, {}).get("results", {}).get("individual_powers", {}).get(test, opt_power)
+ drop = opt_power - other_power
+ max_drop = max(max_drop, drop)
+ drop_str = f"+{abs(drop):.1f}%" if drop < 0 else f"-{drop:.1f}%"
+ row.append(drop_str)
+
+ if max_drop > HIGH_VULNERABILITY_THRESHOLD:
+ vulnerability = "HIGH"
+ vulnerable_tests.append(test)
+ elif max_drop > MEDIUM_VULNERABILITY_THRESHOLD:
+ vulnerability = "MEDIUM"
+ elif max_drop < INFLATED_ERROR_THRESHOLD:
+ vulnerability = "INFLATED FALSE POSITIVES"
+ inflated_tests.append(test)
+ else:
+ vulnerability = "LOW"
- rows.append([test, real_drop_str, doom_drop_str, vulnerability])
+ row.append(vulnerability)
+ rows.append(row)
- lines.append(self._table._create_table(headers, rows))
+ lines.append(self._table._create_table(headers, rows))
# 4. Recommendations
- lines.append(f"\n{'=' * 80}")
- lines.append("RECOMMENDATIONS")
- lines.append(f"{'=' * 80}")
+ if "optimistic" in scenarios and len(scenarios) > 1:
+ lines.append(f"\n{'=' * 80}")
+ lines.append("RECOMMENDATIONS")
+ lines.append(f"{'=' * 80}")
- if vulnerable_tests:
- lines.append(f"• High vulnerability tests: {', '.join(vulnerable_tests)}")
- lines.append("• Consider increasing sample size to maintain power under adverse conditions")
+ if vulnerable_tests:
+ lines.append(f"• High vulnerability tests: {', '.join(vulnerable_tests)}")
+ lines.append("• Consider increasing sample size to maintain power under adverse conditions")
- if inflated_tests:
- lines.append(f"• Inflated false positive risk: {', '.join(inflated_tests)}")
- lines.append("• Be careful about interpretation")
+ if inflated_tests:
+ lines.append(f"• Inflated false positive risk: {', '.join(inflated_tests)}")
+ lines.append("• Be careful about interpretation")
- if not vulnerable_tests and not inflated_tests:
- lines.append("• Power analysis appears robust to assumption violations")
- lines.append("• Original sample size should be sufficient")
+ if not vulnerable_tests and not inflated_tests:
+ lines.append("• Power analysis appears robust to assumption violations")
+ lines.append("• Original sample size should be sufficient")
return "\n".join(lines)
@@ -484,25 +495,22 @@ def _format_scenario_sample_size_short(self, scenarios: Dict, target_tests: List
"""Short scenario sample size summary."""
lines = [f"\n{'=' * 80}", "SCENARIO SUMMARY", f"{'=' * 80}"]
+ scenario_names = list(scenarios.keys())
if correction:
# Combined table with uncorrected and corrected
lines.append("\nSample Size Requirements:")
- headers = [
- "Test",
- "Opt(U)",
- "Opt(C)",
- "Real(U)",
- "Real(C)",
- "Doom(U)",
- "Doom(C)",
- ]
+ headers = ["Test"]
+ for name in scenario_names:
+ abbrev = name[:4].title()
+ headers.extend([f"{abbrev}(U)", f"{abbrev}(C)"])
+ col_widths = [40] + [8] * (len(scenario_names) * 2)
rows = []
for test in target_tests:
- row = [test[:40]] # Truncate to 40 chars
+ row = [test[:40]]
- for scenario in ["optimistic", "realistic", "doomer"]:
+ for scenario in scenario_names:
if scenario in scenarios and "results" in scenarios[scenario]:
n_uncorr = scenarios[scenario]["results"]["first_achieved"][test]
n_corr = scenarios[scenario]["results"]["first_achieved_corrected"][test]
@@ -520,16 +528,17 @@ def _format_scenario_sample_size_short(self, scenarios: Dict, target_tests: List
row.extend(["N/A", "N/A"])
rows.append(row)
- lines.append(self._table._create_table(headers, rows, [40, 8, 8, 8, 8, 8, 8]))
+ lines.append(self._table._create_table(headers, rows, col_widths))
lines.append("Note: (U) = Uncorrected, (C) = Corrected")
else:
# Uncorrected only
- headers = ["Test", "Optimistic", "Realistic", "Doomer"]
+ headers = ["Test"] + [name.title() for name in scenario_names]
+ col_widths = [40] + [12] * len(scenario_names)
rows = []
for test in target_tests:
- row = [test[:40]] # Truncate to 40 chars
- for scenario in ["optimistic", "realistic", "doomer"]:
+ row = [test[:40]]
+ for scenario in scenario_names:
if scenario in scenarios and "results" in scenarios[scenario]:
n_required = scenarios[scenario]["results"]["first_achieved"][test]
if n_required > 0:
@@ -542,7 +551,7 @@ def _format_scenario_sample_size_short(self, scenarios: Dict, target_tests: List
rows.append(row)
lines.append("\nUncorrected Sample Sizes:")
- lines.append(self._table._create_table(headers, rows, [40, 12, 12, 12]))
+ lines.append(self._table._create_table(headers, rows, col_widths))
lines.append(f"{'=' * 80}")
@@ -562,39 +571,33 @@ def _format_scenario_sample_size_long(
# 1. Overall summary
lines.append(self._format_scenario_sample_size_short(scenarios, target_tests, correction))
- # 2. Recommendations
+ # 2. Recommendations — summarize max N per non-optimistic scenario
lines.append(f"\n{'=' * 80}")
lines.append("RECOMMENDATIONS")
lines.append(f"{'=' * 80}")
- # Calculate max required N across scenarios
- max_n_realistic = max(
- (scenarios.get("realistic", {}).get("results", {}).get("first_achieved", {}).get(test, 0) for test in target_tests),
- default=0,
- )
- max_n_doomer = max(
- (scenarios.get("doomer", {}).get("results", {}).get("first_achieved", {}).get(test, 0) for test in target_tests),
- default=0,
- )
-
- max_tested = scenarios.get("realistic", {}).get("model", {}).get("sample_size_range", {}).get("to_size", 200)
-
- if max_n_realistic > 0 and max_n_realistic <= max_tested:
- lines.append(f"• For robust power under realistic conditions: N = {max_n_realistic}")
- elif max_n_realistic <= 0:
- lines.append(f"• For robust power under realistic conditions: N > {max_tested}")
-
- if max_n_doomer > 0 and max_n_doomer <= max_tested:
- lines.append(f"• For power under worst-case conditions: N = {max_n_doomer}")
- elif max_n_doomer <= 0:
- lines.append(f"• For power under worst-case conditions: N > {max_tested}")
-
- # Check if any tests couldn't achieve power
- unachievable = [
- test for test in target_tests if scenarios.get("doomer", {}).get("results", {}).get("first_achieved", {}).get(test, -1) <= 0
- ]
- if unachievable:
- lines.append(f"• Warning: These tests may not achieve target power under adverse conditions: {', '.join(unachievable)}")
+ other_scenarios = [s for s in scenarios if s != "optimistic"]
+ for scenario in other_scenarios:
+ max_n = max(
+ (scenarios.get(scenario, {}).get("results", {}).get("first_achieved", {}).get(test, 0) for test in target_tests),
+ default=0,
+ )
+ max_tested = scenarios.get(scenario, {}).get("model", {}).get("sample_size_range", {}).get("to_size", 200)
+ label = scenario.title()
+
+ if max_n > 0 and max_n <= max_tested:
+ lines.append(f"• For power under {label} conditions: N = {max_n}")
+ elif max_n <= 0:
+ lines.append(f"• For power under {label} conditions: N > {max_tested}")
+
+ # Check unachievable across worst scenario (last non-optimistic)
+ if other_scenarios:
+ worst = other_scenarios[-1]
+ unachievable = [
+ test for test in target_tests if scenarios.get(worst, {}).get("results", {}).get("first_achieved", {}).get(test, -1) <= 0
+ ]
+ if unachievable:
+ lines.append(f"• Warning: These tests may not achieve target power under {worst} conditions: {', '.join(unachievable)}")
# Add cumulative probability analysis
cumulative_lines = self._format_cumulative_recommendations(data, is_scenario=True)
@@ -706,7 +709,7 @@ def _add_cumulative_sample_size_table(
# Filter out tests with NaN power (e.g. non-contrast tests under Tukey correction)
def _has_nan_power(t: str) -> bool:
vals = powers_by_test[t]
- return bool(vals and isinstance(vals[0], float) and math.isnan(vals[0]))
+ return bool(vals and _is_nan(vals[0]))
valid_tests = [t for t in target_tests if not _has_nan_power(t)]
if not valid_tests:
@@ -741,8 +744,6 @@ def _has_nan_power(t: str) -> bool:
else: # ≥k cases
# Approximate using independence assumption
prob_at_least_k = 0.0
- from itertools import combinations
-
# Sum over all ways to choose at least k tests
for num_sig in range(k, n_tests + 1):
for combo in combinations(range(n_tests), num_sig):
@@ -859,6 +860,12 @@ def _format_cumulative_recommendations(self, results: Dict, is_scenario: bool =
if prob >= target_power:
min_n_target = sample_sizes[i]
break
+
+ if min_n_target:
+ lines.append(f"• N={min_n_target} for {target_power:.0f}% chance all tests significant")
+ else:
+ max_tested = sample_sizes[-1]
+ lines.append(f"• >{max_tested} needed for {target_power:.0f}% chance all tests significant")
return lines
diff --git a/mcpower/utils/parsers.py b/mcpower/utils/parsers.py
index e89d140..c14533f 100644
--- a/mcpower/utils/parsers.py
+++ b/mcpower/utils/parsers.py
@@ -105,6 +105,8 @@ def _split_assignments(self, input_string: str) -> List[str]:
paren_count += 1
elif char == ")":
paren_count -= 1
+ if paren_count < 0:
+ raise ValueError("Unbalanced parentheses: unexpected ')'")
current.append(char)
if current:
@@ -424,7 +426,11 @@ def _parse_independent_variables(formula: str) -> Tuple[Dict, Dict]:
"""
from itertools import combinations
- terms = re.split(r"[+\-]", formula)
+ # Check for minus sign (term removal) which is not supported
+ if re.search(r"(? Tuple[List[str], List[Dict]]:
+ """Extract effect names from a test formula, matched against the registry.
+
+ Parses the test formula, expands factor variables to their dummies,
+ and returns the list of effect names (in registry order) that belong
+ to the test formula.
+
+ Args:
+ test_formula: Formula string (e.g. ``"y ~ x1 + x2"``).
+ registry: ``VariableRegistry`` instance.
+
+ Returns:
+ Tuple of ``(effect_names, random_effects)`` where *effect_names*
+ are the registry effect names present in the test formula (in
+ registry order), and *random_effects* is the list of parsed
+ random-effect dicts from the test formula.
+ """
+ _dep_var, fixed_formula, random_effects = _parse_equation(test_formula)
+
+ # Parse fixed effects into a set of term names
+ test_terms = _parse_fixed_terms(fixed_formula)
+
+ # Determine which registry effects belong to the test formula
+ cluster_effects = set(registry.cluster_effect_names)
+ test_effects: List[str] = []
+
+ for effect_name in registry._effects:
+ if effect_name in cluster_effects:
+ continue
+
+ effect = registry._effects[effect_name]
+
+ if effect.effect_type == "main":
+ # Direct match (continuous or interaction-less variable)
+ if effect_name in test_terms:
+ test_effects.append(effect_name)
+ elif effect_name in registry._factor_dummies:
+ # Factor dummy -- include if parent factor is in test terms
+ parent_factor = registry._factor_dummies[effect_name]["factor_name"]
+ if parent_factor in test_terms:
+ test_effects.append(effect_name)
+ else:
+ # Interaction -- check if the interaction term is in test terms
+ if effect_name in test_terms:
+ test_effects.append(effect_name)
+
+ return test_effects, random_effects
+
+
+def _parse_fixed_terms(fixed_formula: str) -> Set[str]:
+ """Parse a fixed-effect formula string into a set of term names.
+
+ Handles ``+`` for additive terms, ``:`` for specific interactions,
+ and ``*`` for full factorial expansion (main effects plus all
+ two-way through n-way interactions).
+
+ Args:
+ fixed_formula: Right-hand side of the equation, spaces already
+ stripped by ``_parse_equation`` (e.g. ``"x1+x2+x1:x2"``).
+
+ Returns:
+ Set of term names (variable names and interaction terms like
+ ``"x1:x2"``).
+ """
+ if not fixed_formula.strip():
+ return set()
+
+ terms: Set[str] = set()
+ raw_terms = re.split(r"\+", fixed_formula)
+
+ for raw in raw_terms:
+ raw = raw.strip()
+ if not raw:
+ continue
+
+ if "*" in raw:
+ # Full factorial: x1*x2 -> x1, x2, x1:x2
+ vars_in_star = [v.strip() for v in raw.split("*") if v.strip()]
+ for v in vars_in_star:
+ terms.add(v)
+ for r in range(2, len(vars_in_star) + 1):
+ for combo in combinations(vars_in_star, r):
+ terms.add(":".join(combo))
+ else:
+ # Plain term (may contain ":" for explicit interaction)
+ terms.add(raw)
+
+ return terms
+
+
+def _compute_test_column_indices(
+ all_effect_names: List[str],
+ test_effect_names: List[str],
+) -> np.ndarray:
+ """Compute column indices in X_expanded for test formula effects.
+
+ Args:
+ all_effect_names: All non-cluster effect names in registry order.
+ test_effect_names: Effect names present in the test formula
+ (a subset of *all_effect_names*).
+
+ Returns:
+ Integer array of column indices into X_expanded.
+ """
+ test_set = set(test_effect_names)
+ indices = [i for i, name in enumerate(all_effect_names) if name in test_set]
+ return np.array(indices, dtype=np.int64)
+
+
+def _remap_target_indices(
+ original_target_indices: np.ndarray,
+ test_column_indices: np.ndarray,
+) -> np.ndarray:
+ """Remap target indices from full X_expanded space to X_test space.
+
+ Args:
+ original_target_indices: Indices in X_expanded being tested.
+ test_column_indices: Columns of X_expanded included in X_test.
+
+ Returns:
+ Indices remapped to positions within X_test.
+ """
+ # Build mapping: full_index -> position in X_test
+ index_map = {int(full_idx): test_idx for test_idx, full_idx in enumerate(test_column_indices)}
+ return np.array(
+ [index_map[int(idx)] for idx in original_target_indices],
+ dtype=np.int64,
+ )
diff --git a/mcpower/utils/updates.py b/mcpower/utils/updates.py
index 8c3c76b..c7f57a5 100644
--- a/mcpower/utils/updates.py
+++ b/mcpower/utils/updates.py
@@ -12,6 +12,8 @@
from datetime import datetime, timedelta
from pathlib import Path
+_already_checked = False
+
def _check_for_updates(current_version):
"""Check PyPI weekly for a newer MCPower version and warn if found.
@@ -20,18 +22,24 @@ def _check_for_updates(current_version):
silently in worker processes (detected via environment variable)
and in frozen (PyInstaller) bundles where pip is unavailable.
"""
+ global _already_checked
# Skip in frozen bundles (PyInstaller) — the GUI has its own update checker
if getattr(sys, "frozen", False):
return
+ # Skip if already checked in this process
+ if _already_checked:
+ return
+
# Skip in worker processes (loky/joblib inherit env vars from parent)
if os.environ.get("_MCPOWER_UPDATE_CHECKED"):
return
os.environ["_MCPOWER_UPDATE_CHECKED"] = "1"
+ _already_checked = True
- cache_path = Path(__file__).parent.parent / ".mcpower_cache.json"
- cache_path.parent.mkdir(exist_ok=True)
+ cache_path = Path.home() / ".cache" / "mcpower" / "update_cache.json"
+ cache_path.parent.mkdir(parents=True, exist_ok=True)
# Load cache
cache = {}
@@ -57,9 +65,8 @@ def _check_for_updates(current_version):
# Show update message only when PyPI version is strictly newer
latest = cache.get("latest_version")
- current = cache.get("current_version")
- if latest and current and _is_newer(latest, current):
- msg = f"\nNEW MCPower VERSION AVAILABLE: {latest} (you have {current})\nUpdate now: pip install --upgrade MCPower\n"
+ if latest and _is_newer(latest, current_version):
+ msg = f"\nNEW MCPower VERSION AVAILABLE: {latest} (you have {current_version})\nUpdate now: pip install --upgrade MCPower\n"
warnings.warn(msg, stacklevel=3)
@@ -77,7 +84,10 @@ def _get_latest_version():
"""Fetch the latest MCPower version string from the PyPI JSON API."""
try:
with urllib.request.urlopen("https://pypi.org/pypi/MCPower/json", timeout=5) as response:
- data = json.loads(response.read())
+ raw = response.read(1_000_000)
+ if len(raw) >= 1_000_000:
+ return None
+ data = json.loads(raw)
return data["info"]["version"]
except Exception:
return None
diff --git a/mcpower/utils/validators.py b/mcpower/utils/validators.py
index 5853af6..1c344fd 100644
--- a/mcpower/utils/validators.py
+++ b/mcpower/utils/validators.py
@@ -27,6 +27,11 @@ class _ValidationResult:
errors: List[str]
warnings: List[str]
+ @classmethod
+ def from_errors(cls, errors: List[str], warnings: Optional[List[str]] = None) -> "_ValidationResult":
+ """Create a result from error/warning lists, deriving ``is_valid`` automatically."""
+ return cls(len(errors) == 0, errors, warnings or [])
+
def raise_if_invalid(self):
"""Raise ``ValueError`` if the validation failed."""
if not self.is_valid:
@@ -88,12 +93,12 @@ def _validate_numeric_parameter(
errors.append(range_error)
# Rounding warning for floats when int expected
- if allow_rounding and isinstance(value, float) and (int, float) in expected_types:
+ if allow_rounding and isinstance(value, float) and int in expected_types:
rounded = int(round(value))
if value != rounded:
warnings.append(f"{name} rounded from {value} to {rounded}")
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
def _validate_power(power: Any) -> _ValidationResult:
@@ -112,6 +117,8 @@ def _validate_simulations(n_simulations: Any) -> Tuple[int, _ValidationResult]:
if result.is_valid:
rounded = int(round(n_simulations))
+ # 800 simulations threshold: below this, Monte Carlo standard error
+ # exceeds ~1.5% for power near 50%, reducing result reliability.
if rounded < 800:
result.warnings.append(f"Low simulation count ({rounded}). Consider using at least 1000 for reliable results.")
return rounded, result
@@ -139,7 +146,7 @@ def _validate_sample_size(sample_size: Any) -> _ValidationResult:
f"sample_size too large ({sample_size:,}). Maximum recommended: 100,000. We cannot guarantee stability for such small p-values."
)
- return _ValidationResult(len(errors) == 0, errors, [])
+ return _ValidationResult.from_errors(errors)
def _validate_sample_size_for_model(sample_size: int, n_variables: int) -> _ValidationResult:
@@ -157,6 +164,8 @@ def _validate_sample_size_for_model(sample_size: int, n_variables: int) -> _Vali
_ValidationResult with errors if sample size is insufficient.
"""
errors = []
+ # Green's rule of thumb: N >= 15 + p for adequate power in regression,
+ # where p is the number of predictors (design matrix columns).
min_required = 15 + n_variables
if sample_size < min_required:
@@ -165,7 +174,7 @@ def _validate_sample_size_for_model(sample_size: int, n_variables: int) -> _Vali
f"variables. Minimum required: {min_required} (15 + {n_variables} variables)."
)
- return _ValidationResult(len(errors) == 0, errors, [])
+ return _ValidationResult.from_errors(errors)
def _validate_sample_size_range(from_size: Any, to_size: Any, by: Any) -> _ValidationResult:
@@ -193,7 +202,7 @@ def _validate_sample_size_range(from_size: Any, to_size: Any, by: Any) -> _Valid
if n_tests > 100:
warnings.append(f"Large number of sample sizes to test ({n_tests}). This may take significant time.")
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
def _validate_correlation_matrix(
@@ -226,12 +235,14 @@ def _validate_correlation_matrix(
# Positive semi-definite check
try:
eigenvals = np.linalg.eigvals(corr_matrix)
+ # -1e-8 tolerance for positive semi-definiteness: allows small negative
+ # eigenvalues from floating-point rounding in correlation matrices.
if np.any(eigenvals < -1e-8): # Tolerance for floating point noise
errors.append("Correlation matrix must be positive semi-definite. ")
except np.linalg.LinAlgError:
errors.append("Cannot compute eigenvalues of correlation matrix")
- return _ValidationResult(len(errors) == 0, errors, [])
+ return _ValidationResult.from_errors(errors)
def _validate_correction_method(correction: Optional[str]) -> _ValidationResult:
@@ -285,7 +296,7 @@ def _validate_parallel_settings(enable: Any, n_cores: Optional[int]) -> Tuple[Tu
else:
validated_n_cores = min(n_cores, max_cores)
- return (enable, validated_n_cores), _ValidationResult(len(errors) == 0, errors, [])
+ return (enable, validated_n_cores), _ValidationResult.from_errors(errors)
def _validate_model_ready(model) -> _ValidationResult:
@@ -301,9 +312,10 @@ def _validate_model_ready(model) -> _ValidationResult:
errors: List[str] = []
warnings: List[str] = []
- # Check effect sizes - check if pending effects were set
- has_effects = hasattr(model, "_pending_effects") and model._pending_effects is not None
- if not has_effects:
+ # Check effect sizes — pending (pre-apply) or flagged as set by user
+ has_pending = hasattr(model, "_pending_effects") and model._pending_effects is not None
+ has_set = hasattr(model, "_effects_set") and model._effects_set
+ if not has_pending and not has_set:
if hasattr(model, "_registry"):
available = model._registry.effect_names
errors.append(
@@ -318,7 +330,7 @@ def _validate_model_ready(model) -> _ValidationResult:
if not hasattr(model, attr):
errors.append(f"Model missing required attribute: {attr}")
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
def _validate_test_formula(test_formula: str, available_variables: List[str]) -> _ValidationResult:
@@ -361,7 +373,7 @@ def _validate_test_formula(test_formula: str, available_variables: List[str]) ->
f"Variables not found in original model: {', '.join(sorted(missing_vars))}. Available: {', '.join(available_variables)}"
)
- return _ValidationResult(len(errors) == 0, errors, [])
+ return _ValidationResult.from_errors(errors)
except Exception as e:
errors.append(f"Error parsing test_formula: {str(e)}")
@@ -399,6 +411,8 @@ def _validate_factor_specification(n_levels: int, proportions: List[float]) -> _
# Check if they sum to approximately 1
if not errors: # Only if no errors with individual proportions
total = sum(proportions)
+ # 1e-6 tolerance: proportions are normalized later, so small deviations
+ # from 1.0 are acceptable and only warrant a warning.
if abs(total - 1.0) > 1e-6:
warnings.append(f"Proportions sum to {total:.4f}, not 1.0 (will be normalized)")
@@ -406,7 +420,7 @@ def _validate_factor_specification(n_levels: int, proportions: List[float]) -> _
if n_levels > 10:
warnings.append(f"Factor has {n_levels} levels. This creates {n_levels - 1} dummy variables, which may require large sample sizes")
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
def _validate_upload_data(data: np.ndarray) -> _ValidationResult:
@@ -425,7 +439,7 @@ def _validate_upload_data(data: np.ndarray) -> _ValidationResult:
if data.shape[0] < 25:
errors.append(f"Need at least 25 samples for reliable quantile matching, got {data.shape[0]}")
- return _ValidationResult(len(errors) == 0, errors, [])
+ return _ValidationResult.from_errors(errors)
def _validate_cluster_config(
@@ -475,7 +489,7 @@ def _validate_cluster_config(
if not isinstance(cluster_size, int) or cluster_size < 5:
errors.append(f"cluster_size must be an integer >= 5 for reliable mixed model estimation. Got {cluster_size}.")
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
def _validate_cluster_sample_size(
@@ -517,4 +531,4 @@ def _validate_cluster_sample_size(
f"Small cluster sizes may cause convergence issues or biased variance estimates."
)
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
diff --git a/mcpower/utils/visualization.py b/mcpower/utils/visualization.py
index 22544a2..ab7eadd 100644
--- a/mcpower/utils/visualization.py
+++ b/mcpower/utils/visualization.py
@@ -18,6 +18,7 @@ def _create_power_plot(
target_tests: List[str],
target_power: float,
title: str,
+ show: bool = True,
):
"""Create a sample-size vs. power line plot with achievement markers.
@@ -58,7 +59,7 @@ def _create_power_plot(
)
# Mark achievement point
- if first_achieved[test] > 0:
+ if first_achieved[test] > 0 and first_achieved[test] in sample_sizes:
achieved_idx = sample_sizes.index(first_achieved[test])
achieved_power = powers[achieved_idx]
ax.plot(
@@ -112,4 +113,6 @@ def _create_power_plot(
color="#888888",
)
plt.tight_layout(rect=(0, 0.03, 1, 1))
- plt.show()
+ if show:
+ plt.show()
+ return fig
diff --git a/pyproject.toml b/pyproject.toml
index 983e3d3..ec1f41f 100644
--- a/pyproject.toml
+++ b/pyproject.toml
@@ -1,14 +1,14 @@
[build-system]
requires = [
- "scikit-build-core>=0.5",
+ "scikit-build-core>=0.10",
"pybind11>=2.11",
- "numpy>=2.0.0",
+ "numpy>=1.26.0",
]
build-backend = "scikit_build_core.build"
[project]
name = "MCPower"
-version = "0.5.4"
+version = "0.6.0"
description = "Monte Carlo Power Analysis for Statistical Models"
readme = "README.md"
license = {text = "GPL-3.0-or-later"}
@@ -31,9 +31,10 @@ classifiers = [
]
requires-python = ">=3.10"
dependencies = [
- "numpy>=2.0.0",
+ "numpy>=1.26.0",
"matplotlib>=3.8.0",
"joblib>=1.3.0",
+ "tqdm>=4.60.0",
]
[project.optional-dependencies]
@@ -41,6 +42,7 @@ lme = ["statsmodels>=0.14.0"]
pandas = ["pandas>=2.0.0"]
dev = [
"pandas>=2.0.0",
+ "statsmodels>=0.14.0",
"pytest>=7.0.0",
"pytest-cov>=4.0.0",
"scipy>=1.11.0",
@@ -52,14 +54,14 @@ dev = [
]
all = [
"pandas>=2.0.0",
- "statsmodels>=0.14.0",
-]
+ ]
[project.urls]
Homepage = "https://github.com/pawlenartowicz/MCPower"
-Documentation = "https://github.com/pawlenartowicz/MCPower#readme"
+Documentation = "https://github.com/pawlenartowicz/MCPower/wiki"
Repository = "https://github.com/pawlenartowicz/MCPower"
Issues = "https://github.com/pawlenartowicz/MCPower/issues"
+Changelog = "https://github.com/pawlenartowicz/MCPower/blob/main/CHANGELOG.md"
[tool.scikit-build]
wheel.packages = ["mcpower"]
@@ -77,8 +79,6 @@ python_files = ["test_*.py"]
python_classes = ["Test*"]
python_functions = ["test_*"]
markers = [
- "unit: Unit tests",
- "integration: Integration tests",
"lme: LME mixed-effects model tests",
]
addopts = "-v --tb=short --strict-markers"
@@ -86,7 +86,6 @@ filterwarnings = [
"ignore::FutureWarning",
"ignore::DeprecationWarning",
"ignore::UserWarning:statsmodels",
- "ignore:Mixed-effects models are experimental:UserWarning",
]
[tool.ruff]
@@ -107,6 +106,20 @@ known-first-party = ["mcpower"]
python_version = "3.10"
warn_return_any = true
warn_unused_configs = true
-ignore_missing_imports = true
check_untyped_defs = true
exclude = ["build", "dist", "tests"]
+
+[[tool.mypy.overrides]]
+module = [
+ "mcpower_native",
+ "mcpower_native.*",
+ "statsmodels",
+ "statsmodels.*",
+ "tqdm",
+ "tqdm.*",
+ "joblib",
+ "joblib.*",
+ "pandas",
+ "pandas.*",
+]
+ignore_missing_imports = true
diff --git a/tests/config.py b/tests/config.py
index 63c1999..a8b874c 100644
--- a/tests/config.py
+++ b/tests/config.py
@@ -5,9 +5,21 @@
across the test suite.
"""
-# Monte Carlo simulation parameters
-N_SIMS = 5000
-"""Number of Monte Carlo simulations for power analysis tests."""
+# Monte Carlo simulation parameters — 4-tier ladder
+N_SIMS_CHECK = 50
+"""Smoke tests — just verify no crash, structure, API contract."""
+
+N_SIMS_ORDERING = 1000
+"""Ordering tests — monotonicity, correction hierarchy, A < B checks."""
+
+N_SIMS_STANDARD = 1600
+"""Standard tests — null calibration, Type I error, general validation."""
+
+N_SIMS_ACCURACY = 5000
+"""Accuracy tests — comparison against analytical power formulas."""
+
+N_SIMS = N_SIMS_ACCURACY
+"""Backward-compat alias for accuracy-level simulations."""
SEED = 2137
"""Default random seed for reproducibility."""
diff --git a/tests/conftest.py b/tests/conftest.py
index 93c8814..d2100b5 100644
--- a/tests/conftest.py
+++ b/tests/conftest.py
@@ -63,12 +63,6 @@ def correlation_matrix_2x2():
return np.array([[1.0, 0.5], [0.5, 1.0]])
-@pytest.fixture
-def correlation_matrix_3x3():
- """Create a 3x3 correlation matrix."""
- return np.array([[1.0, 0.3, 0.2], [0.3, 1.0, 0.4], [0.2, 0.4, 1.0]])
-
-
@pytest.fixture
def sample_data():
"""Create sample empirical data."""
@@ -80,41 +74,13 @@ def sample_data():
@pytest.fixture
-def suppress_output(capsys):
- """Suppress print output during tests by capturing it."""
- yield
- # Output is automatically captured by capsys
-
-
-BACKENDS = ["c++"]
-
-
-@pytest.fixture(params=BACKENDS)
-def backend(request):
- """
- Force MCPower to run on a specific backend.
-
- Parametrizes tests against C++ (primary backend).
- Automatically resets backend after each test.
- """
- from mcpower.backends import reset_backend, set_backend
-
- set_backend(request.param)
- yield request.param
- reset_backend()
-
-
-@pytest.fixture(autouse=True)
-def reset_backend_after_test():
- """
- Automatically reset backend to default after every test.
-
- Ensures no hidden backend state leaks between tests.
- """
- yield
- from mcpower.backends import reset_backend
+def suppress_output():
+ """Suppress print output during tests."""
+ import contextlib
+ import io
- reset_backend()
+ with contextlib.redirect_stdout(io.StringIO()):
+ yield
def _statsmodels_available():
diff --git a/tests/helpers/power_helpers.py b/tests/helpers/power_helpers.py
index d509a79..620e995 100644
--- a/tests/helpers/power_helpers.py
+++ b/tests/helpers/power_helpers.py
@@ -40,42 +40,3 @@ def compute_crits(X, target_indices, alpha=DEFAULT_ALPHA, correction_method=0):
return compute_critical_values(alpha, p, dof, n_targets, correction_method)
-def run_with_backend(
- backend_name,
- equation,
- effects_str,
- sample_size,
- n_sims,
- seed,
- target_test="all",
- correction=None,
- correlations_str=None,
- alpha=DEFAULT_ALPHA,
-):
- """Run a full MCPower power analysis with a specific backend forced."""
- import contextlib
- import io
-
- from mcpower import MCPower
- from mcpower.backends import reset_backend, set_backend
-
- set_backend(backend_name)
- try:
- m = MCPower(equation)
- m.set_simulations(n_sims)
- m.set_seed(seed)
- m.set_alpha(alpha)
- m.set_effects(effects_str)
- if correlations_str:
- m.set_correlations(correlations_str)
- with contextlib.redirect_stdout(io.StringIO()):
- result = m.find_power(
- sample_size=sample_size,
- target_test=target_test,
- correction=correction,
- print_results=False,
- return_results=True,
- )
- finally:
- reset_backend()
- return result
diff --git a/tests/integration/test_find_power_api.py b/tests/integration/test_find_power_api.py
index 69f6bc5..fe56de3 100644
--- a/tests/integration/test_find_power_api.py
+++ b/tests/integration/test_find_power_api.py
@@ -222,14 +222,14 @@ def test_all_targets(self, suppress_output):
class TestHeterogeneity:
- """Test heterogeneity settings."""
+ """Test heterogeneity via scenario configs."""
def test_with_heterogeneity(self, suppress_output):
from mcpower import MCPower
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
- model.set_heterogeneity(0.1)
+ model.set_scenario_configs({"het": {"heterogeneity": 0.1}})
result = model.find_power(100, print_results=False, return_results=True)
assert result is not None
@@ -239,7 +239,7 @@ def test_with_heteroskedasticity(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
- model.set_heteroskedasticity(0.2)
+ model.set_scenario_configs({"hsked": {"heteroskedasticity": 0.2}})
result = model.find_power(100, print_results=False, return_results=True)
assert result is not None
@@ -249,8 +249,7 @@ def test_combined(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
- model.set_heterogeneity(0.1)
- model.set_heteroskedasticity(0.2)
+ model.set_scenario_configs({"combo": {"heterogeneity": 0.1, "heteroskedasticity": 0.2}})
result = model.find_power(100, print_results=False, return_results=True)
assert result is not None
@@ -330,7 +329,7 @@ def test_all_features_combined(self, suppress_output):
model.upload_data({"x1": np.random.exponential(2, 100)})
model.set_correlations("(x1,x2)=0.3")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2, x2=0.15, x1:x2=0.1")
- model.set_heterogeneity(0.05)
+ model.set_scenario_configs({"test": {"heterogeneity": 0.05}})
result = model.find_power(200, print_results=False, return_results=True)
assert result is not None
diff --git a/tests/integration/test_model.py b/tests/integration/test_model.py
index da00d33..a57b051 100644
--- a/tests/integration/test_model.py
+++ b/tests/integration/test_model.py
@@ -86,16 +86,6 @@ def test_set_variable_type(self, suppress_output):
assert model._pending_variable_types == "group=(factor,3)"
assert model._applied is False
- def test_set_heterogeneity(self, simple_model):
- simple_model.set_heterogeneity(0.1)
- assert simple_model._pending_heterogeneity == 0.1
- assert simple_model._applied is False
-
- def test_set_heteroskedasticity(self, simple_model):
- simple_model.set_heteroskedasticity(0.2)
- assert simple_model._pending_heteroskedasticity == 0.2
- assert simple_model._applied is False
-
def test_upload_data_dict(self, simple_model, sample_data):
simple_model.upload_data(sample_data)
assert simple_model._pending_data is not None
@@ -131,12 +121,12 @@ class TestApply:
"""Test apply() method."""
def test_apply_sets_flag(self, configured_model):
- configured_model.apply()
+ configured_model._apply()
assert configured_model._applied is True
def test_apply_processes_effects(self, simple_model):
simple_model.set_effects("x1=0.5, x2=0.3")
- simple_model.apply()
+ simple_model._apply()
effect_sizes = simple_model._registry.get_effect_sizes()
assert effect_sizes[0] == 0.5
assert effect_sizes[1] == 0.3
@@ -147,22 +137,17 @@ def test_apply_processes_variable_types(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
assert len(model._registry.factor_names) == 1
assert len(model._registry.dummy_names) == 2
def test_apply_processes_correlations(self, simple_model):
simple_model.set_effects("x1=0.3, x2=0.2")
simple_model.set_correlations("(x1,x2)=0.5")
- simple_model.apply()
+ simple_model._apply()
corr = simple_model.correlation_matrix
assert corr[0, 1] == 0.5
- def test_apply_processes_heterogeneity(self, configured_model):
- configured_model.set_heterogeneity(0.15)
- configured_model.apply()
- assert configured_model.heterogeneity == 0.15
-
def test_apply_order_independence(self, suppress_output):
"""Test that set_* methods can be called in any order."""
from mcpower import MCPower
@@ -172,14 +157,14 @@ def test_apply_order_independence(self, suppress_output):
m1.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2, x2=0.1")
m1.set_variable_type("group=(factor,3)")
m1.set_correlations("(x1,x2)=0.5")
- m1.apply()
+ m1._apply()
# Order 2: variable_type, correlations, effects
m2 = MCPower("y = group + x1 + x2")
m2.set_variable_type("group=(factor,3)")
m2.set_correlations("(x1,x2)=0.5")
m2.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2, x2=0.1")
- m2.apply()
+ m2._apply()
# Both should have same effect sizes
assert np.allclose(m1._registry.get_effect_sizes(), m2._registry.get_effect_sizes())
@@ -242,7 +227,7 @@ def test_sample_sizes_tested(self, configured_model):
assert result["results"]["sample_sizes_tested"] == [50, 75, 100]
def test_first_achieved(self, configured_model):
- result = configured_model.find_sample_size(from_size=50, to_size=200, by=25, print_results=False, return_results=True)
+ result = configured_model.find_sample_size(from_size=50, to_size=200, by=50, print_results=False, return_results=True)
assert "first_achieved" in result["results"]
def test_find_sample_size_runs(self, configured_model):
@@ -257,7 +242,7 @@ class TestErrors:
def test_invalid_effect_name(self, simple_model):
simple_model.set_effects("invalid=0.3")
with pytest.raises(ValueError, match="not found"):
- simple_model.apply()
+ simple_model._apply()
def test_missing_effects(self, simple_model):
with pytest.raises(ValueError, match="Effect sizes must be set"):
@@ -290,7 +275,7 @@ def test_basic_named_levels(self):
model = MCPower("y = treatment + x1")
model.set_factor_levels("treatment=placebo,drug_a,drug_b")
model.set_effects("treatment[drug_a]=0.5, treatment[drug_b]=0.8, x1=0.3")
- model.apply()
+ model._apply()
assert "treatment" in model._registry.factor_names
assert "treatment[drug_a]" in model._registry.dummy_names
assert "treatment[drug_b]" in model._registry.dummy_names
@@ -302,7 +287,7 @@ def test_multiple_factors(self):
model = MCPower("y = group + dose")
model.set_factor_levels("group=control,treatment; dose=low,medium,high")
model.set_effects("group[treatment]=0.5, dose[medium]=0.3, dose[high]=0.6")
- model.apply()
+ model._apply()
assert "group[treatment]" in model._registry.dummy_names
assert "dose[medium]" in model._registry.dummy_names
assert "dose[high]" in model._registry.dummy_names
@@ -313,7 +298,7 @@ def test_unknown_variable_raises(self):
model = MCPower("y = x1")
with pytest.raises(ValueError, match="not found"):
model.set_factor_levels("unknown=a,b,c")
- model.apply()
+ model._apply()
def test_single_level_raises(self):
from mcpower import MCPower
@@ -321,7 +306,7 @@ def test_single_level_raises(self):
model = MCPower("y = x1")
with pytest.raises(ValueError, match="at least 2"):
model.set_factor_levels("x1=only_one")
- model.apply()
+ model._apply()
def test_find_power_with_named_levels(self):
"""End-to-end: find_power works with set_factor_levels."""
diff --git a/tests/integration/test_parallel.py b/tests/integration/test_parallel.py
index 682372c..2048d7e 100644
--- a/tests/integration/test_parallel.py
+++ b/tests/integration/test_parallel.py
@@ -4,6 +4,8 @@
import pytest
+from tests.config import N_SIMS_CHECK
+
def _joblib_available():
"""Check if joblib is available."""
@@ -26,6 +28,7 @@ def test_parallel_results_match_sequential(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
model.set_seed(42)
+ model.set_simulations(N_SIMS_CHECK)
# Run sequential analysis
model.set_parallel(False)
@@ -56,6 +59,7 @@ def test_parallel_with_scenarios(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
model.set_seed(42)
+ model.set_simulations(N_SIMS_CHECK)
# Run sequential with scenarios
model.set_parallel(False)
@@ -93,6 +97,7 @@ def test_parallel_with_interactions(self, suppress_output):
model = MCPower("y = a + b + a:b")
model.set_effects("a=0.4, b=0.3, a:b=0.2")
model.set_seed(42)
+ model.set_simulations(N_SIMS_CHECK)
# Run sequential
model.set_parallel(False)
@@ -128,6 +133,7 @@ def test_parallel_fallback_on_failure(self, suppress_output, monkeypatch):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
model.set_seed(42)
+ model.set_simulations(N_SIMS_CHECK)
model.set_parallel(True, n_cores=2)
# Mock joblib.Parallel to raise an exception
@@ -157,6 +163,7 @@ def test_find_power_ignores_parallel(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
model.set_seed(42)
+ model.set_simulations(N_SIMS_CHECK)
# Run with parallel=False
model.set_parallel(False)
diff --git a/tests/integration/test_posthoc_integration.py b/tests/integration/test_posthoc_integration.py
index 9499388..b6b7f21 100644
--- a/tests/integration/test_posthoc_integration.py
+++ b/tests/integration/test_posthoc_integration.py
@@ -15,7 +15,7 @@ def test_parse_vs_syntax(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("group[1] vs group[2]")
assert "group[1] vs group[2]" in tests
@@ -27,7 +27,7 @@ def test_parse_multiple_vs(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("group[1] vs group[2], group[2] vs group[3]")
assert "group[1] vs group[2]" in tests
@@ -40,7 +40,7 @@ def test_parse_mixed_regular_and_posthoc(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("overall, group[1] vs group[2]")
assert "overall" in tests
@@ -52,7 +52,7 @@ def test_all_does_not_include_posthoc(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all")
# "all" should NOT include any post-hoc comparisons
@@ -66,7 +66,7 @@ def test_invalid_factor_name(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="Factor.*not found"):
model._parse_target_tests("notafactor[1] vs notafactor[2]")
@@ -77,7 +77,7 @@ def test_invalid_level(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="out of range"):
model._parse_target_tests("group[0] vs group[5]")
@@ -88,7 +88,7 @@ def test_same_level_comparison_rejected(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="Cannot compare a level to itself"):
model._parse_target_tests("group[2] vs group[2]")
@@ -99,7 +99,7 @@ def test_cross_factor_comparison_rejected(self, suppress_output):
model = MCPower("y = a + b")
model.set_variable_type("a=(factor,3), b=(factor,2)")
model.set_effects("a[2]=0.3, a[3]=0.2, b[2]=0.1")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="same factor"):
model._parse_target_tests("a[1] vs b[1]")
@@ -396,7 +396,7 @@ def test_all_posthoc_keyword(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all-posthoc")
# 3-level factor → C(3,2) = 3 pairs
@@ -415,7 +415,7 @@ def test_all_plus_all_posthoc(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all, all-posthoc")
# "all" → overall + group[2] + group[3] + x1 = 4
@@ -434,7 +434,7 @@ def test_all_posthoc_multiple_factors(self, suppress_output):
model = MCPower("y = a + b")
model.set_variable_type("a=(factor,3), b=(factor,2)")
model.set_effects("a[2]=0.3, a[3]=0.2, b[2]=0.1")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all-posthoc")
# a: C(3,2)=3, b: C(2,2)=1 → 4 total
@@ -448,7 +448,7 @@ def test_all_posthoc_no_factors_with_all(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all, all-posthoc")
assert "overall" in tests
@@ -461,7 +461,7 @@ def test_all_posthoc_alone_no_factors_raises(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="no factor variables"):
model._parse_target_tests("all-posthoc")
@@ -472,7 +472,7 @@ def test_exclusion_removes_test(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all, -overall")
assert "overall" not in tests
@@ -486,7 +486,7 @@ def test_exclusion_posthoc(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all-posthoc, -group[1] vs group[2]")
assert "group[1] vs group[2]" not in tests
@@ -500,7 +500,7 @@ def test_exclusion_invalid_raises(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="does not match"):
model._parse_target_tests("all, -nonexistent")
@@ -511,7 +511,7 @@ def test_exclusion_all_raises(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="nothing left"):
model._parse_target_tests("all, -overall, -x1, -x2")
@@ -522,7 +522,7 @@ def test_duplicate_raises(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="Duplicate"):
model._parse_target_tests("all, x1")
diff --git a/tests/integration/test_scenarios.py b/tests/integration/test_scenarios.py
index 5172fb3..b0339cc 100644
--- a/tests/integration/test_scenarios.py
+++ b/tests/integration/test_scenarios.py
@@ -5,8 +5,10 @@
from unittest.mock import MagicMock
import numpy as np
+import pytest
from mcpower.core.scenarios import (
+ DEFAULT_SCENARIO_CONFIG,
ScenarioRunner,
apply_per_simulation_perturbations,
)
@@ -82,6 +84,152 @@ def test_create_scenario_plots_early_return(self):
runner._create_scenario_plots({"scenarios": {"optimistic": {}}})
+class TestSetScenarioConfigs:
+ """Test set_scenario_configs() merge behavior and KeyError prevention."""
+
+ # All keys that must exist in every scenario config
+ ALL_KEYS = sorted(DEFAULT_SCENARIO_CONFIG["optimistic"].keys())
+
+ def _make_model(self):
+ from mcpower import MCPower
+
+ m = MCPower("y = x1 + x2")
+ m.set_effects("x1=0.3, x2=0.2")
+ return m
+
+ # ── Merge semantics ──────────────────────────────────────────
+
+ def test_custom_scenario_inherits_all_optimistic_keys(self):
+ """New custom scenario with one key still has every required key."""
+ m = self._make_model()
+ m.set_scenario_configs({"extreme": {"heterogeneity": 0.6}})
+ cfg = m._scenario_configs["extreme"]
+ missing = set(self.ALL_KEYS) - set(cfg.keys())
+ assert not missing, f"Missing keys: {missing}"
+
+ def test_custom_scenario_overrides_value(self):
+ """Provided key overrides the optimistic default."""
+ m = self._make_model()
+ m.set_scenario_configs({"extreme": {"heterogeneity": 0.6}})
+ assert m._scenario_configs["extreme"]["heterogeneity"] == 0.6
+
+ def test_custom_scenario_non_overridden_keys_are_optimistic(self):
+ """Non-overridden keys equal the optimistic baseline."""
+ m = self._make_model()
+ m.set_scenario_configs({"extreme": {"heterogeneity": 0.6}})
+ opt = DEFAULT_SCENARIO_CONFIG["optimistic"]
+ cfg = m._scenario_configs["extreme"]
+ for key in self.ALL_KEYS:
+ if key != "heterogeneity":
+ assert cfg[key] == opt[key], f"Key {key}: {cfg[key]} != {opt[key]}"
+
+ def test_existing_scenario_update_preserves_other_keys(self):
+ """Updating one key on 'realistic' keeps the rest intact."""
+ m = self._make_model()
+ m.set_scenario_configs({"realistic": {"heterogeneity": 0.99}})
+ cfg = m._scenario_configs["realistic"]
+ assert cfg["heterogeneity"] == 0.99
+ # Other keys should match original realistic defaults
+ assert cfg["correlation_noise_sd"] == DEFAULT_SCENARIO_CONFIG["realistic"]["correlation_noise_sd"]
+
+ def test_defaults_still_present_after_adding_custom(self):
+ """Adding a custom scenario doesn't remove optimistic/realistic/doomer."""
+ m = self._make_model()
+ m.set_scenario_configs({"custom": {"heterogeneity": 0.1}})
+ for name in ("optimistic", "realistic", "doomer", "custom"):
+ assert name in m._scenario_configs
+
+ def test_multiple_custom_scenarios(self):
+ """Multiple custom scenarios each inherit independently."""
+ m = self._make_model()
+ m.set_scenario_configs({
+ "mild": {"heterogeneity": 0.05},
+ "severe": {"heterogeneity": 0.8, "heteroskedasticity": 0.5},
+ })
+ assert m._scenario_configs["mild"]["heterogeneity"] == 0.05
+ assert m._scenario_configs["mild"]["heteroskedasticity"] == 0.0 # optimistic default
+ assert m._scenario_configs["severe"]["heterogeneity"] == 0.8
+ assert m._scenario_configs["severe"]["heteroskedasticity"] == 0.5
+
+ def test_empty_custom_scenario_equals_optimistic(self):
+ """An empty custom config is identical to the optimistic baseline."""
+ m = self._make_model()
+ m.set_scenario_configs({"empty": {}})
+ opt = DEFAULT_SCENARIO_CONFIG["optimistic"]
+ for key in self.ALL_KEYS:
+ assert m._scenario_configs["empty"][key] == opt[key]
+
+ # ── Type validation ──────────────────────────────────────────
+
+ def test_non_dict_raises_type_error(self):
+ m = self._make_model()
+ with pytest.raises(TypeError):
+ m.set_scenario_configs("not_a_dict")
+
+ def test_returns_self_for_chaining(self):
+ m = self._make_model()
+ result = m.set_scenario_configs({"custom": {"heterogeneity": 0.1}})
+ assert result is m
+
+ # ── End-to-end: no KeyError during simulation ────────────────
+
+ def test_custom_partial_config_runs_without_error(self):
+ """Custom scenario with only one key runs find_power without KeyError."""
+ m = self._make_model()
+ m.set_scenario_configs({"partial": {"heterogeneity": 0.3}})
+ result = m.find_power(
+ 50, scenarios=True, print_results=False, return_results=True
+ )
+ assert "partial" in result["scenarios"]
+ power = result["scenarios"]["partial"]["results"]["individual_powers"]["overall"]
+ assert 0 <= power <= 100
+
+ def test_custom_residual_only_config_runs(self):
+ """Custom scenario with only residual keys runs without error."""
+ m = self._make_model()
+ m.set_scenario_configs({
+ "residual_test": {
+ "residual_change_prob": 1.0,
+ "residual_dists": ["heavy_tailed"],
+ "residual_df": 5,
+ }
+ })
+ result = m.find_power(
+ 50, scenarios=True, print_results=False, return_results=True
+ )
+ assert "residual_test" in result["scenarios"]
+
+ def test_custom_lme_keys_on_ols_model_ignored(self):
+ """LME-specific keys on an OLS model don't cause errors."""
+ m = self._make_model()
+ m.set_scenario_configs({
+ "lme_on_ols": {
+ "icc_noise_sd": 0.3,
+ "random_effect_dist": "heavy_tailed",
+ "random_effect_df": 3,
+ }
+ })
+ result = m.find_power(
+ 50, scenarios=True, print_results=False, return_results=True
+ )
+ assert "lme_on_ols" in result["scenarios"]
+
+ def test_overriding_all_three_defaults(self):
+ """Overriding optimistic, realistic, and doomer all at once."""
+ m = self._make_model()
+ m.set_scenario_configs({
+ "optimistic": {"heterogeneity": 0.01},
+ "realistic": {"heterogeneity": 0.5},
+ "doomer": {"heterogeneity": 0.9},
+ })
+ assert m._scenario_configs["optimistic"]["heterogeneity"] == 0.01
+ assert m._scenario_configs["realistic"]["heterogeneity"] == 0.5
+ assert m._scenario_configs["doomer"]["heterogeneity"] == 0.9
+ # Other keys preserved from defaults
+ assert m._scenario_configs["realistic"]["correlation_noise_sd"] == DEFAULT_SCENARIO_CONFIG["realistic"]["correlation_noise_sd"]
+ assert m._scenario_configs["doomer"]["correlation_noise_sd"] == DEFAULT_SCENARIO_CONFIG["doomer"]["correlation_noise_sd"]
+
+
class TestApplyPerSimulationPerturbations:
"""Test apply_per_simulation_perturbations function."""
@@ -122,3 +270,152 @@ def test_var_type_perturbation(self):
# All normal (type 0) vars should be changed to right_skewed (type 2)
assert np.all(p_types == 2)
+
+
+class TestScenarioConfigKeysE2E:
+ """End-to-end tests for each individual config key and mixed combinations.
+
+ Each test verifies that setting a single config key (or combination)
+ via set_scenario_configs() runs find_power(scenarios=True) without
+ error and produces valid power values.
+ """
+
+ N_SIMS = 50
+ SAMPLE_SIZE = 80
+
+ def _make_model(self):
+ from mcpower import MCPower
+
+ m = MCPower("y = x1 + x2")
+ m.set_effects("x1=0.3, x2=0.2")
+ m.set_simulations(self.N_SIMS)
+ return m
+
+ def _run(self, model, config, scenario_name="test_scenario"):
+ model.set_scenario_configs({scenario_name: config})
+ result = model.find_power(
+ self.SAMPLE_SIZE,
+ scenarios=True,
+ print_results=False,
+ return_results=True,
+ )
+ power = result["scenarios"][scenario_name]["results"]["individual_powers"]["overall"]
+ assert 0 <= power <= 100, f"Power out of range: {power}"
+ return result
+
+ # ── Individual general keys ───────────────────────────────────
+
+ def test_heterogeneity_only(self):
+ self._run(self._make_model(), {"heterogeneity": 0.3})
+
+ def test_heteroskedasticity_only(self):
+ self._run(self._make_model(), {"heteroskedasticity": 0.2})
+
+ def test_correlation_noise_sd_only(self):
+ m = self._make_model()
+ m.set_correlations("(x1,x2)=0.4")
+ self._run(m, {"correlation_noise_sd": 0.3})
+
+ def test_distribution_change_prob_only(self):
+ self._run(self._make_model(), {"distribution_change_prob": 0.5})
+
+ def test_new_distributions_with_change_prob(self):
+ self._run(self._make_model(), {
+ "distribution_change_prob": 1.0,
+ "new_distributions": ["uniform"],
+ })
+
+ # ── Individual residual keys ──────────────────────────────────
+
+ def test_residual_change_prob_only(self):
+ self._run(self._make_model(), {"residual_change_prob": 0.5})
+
+ def test_residual_df_only(self):
+ self._run(self._make_model(), {
+ "residual_change_prob": 1.0,
+ "residual_df": 3,
+ })
+
+ def test_residual_dists_only(self):
+ self._run(self._make_model(), {
+ "residual_change_prob": 1.0,
+ "residual_dists": ["heavy_tailed"],
+ })
+
+ # ── Mixed general combinations ────────────────────────────────
+
+ def test_heterogeneity_and_correlation_noise(self):
+ m = self._make_model()
+ m.set_correlations("(x1,x2)=0.3")
+ self._run(m, {
+ "heterogeneity": 0.25,
+ "correlation_noise_sd": 0.3,
+ })
+
+ def test_distribution_change_and_heteroskedasticity(self):
+ self._run(self._make_model(), {
+ "distribution_change_prob": 0.5,
+ "heteroskedasticity": 0.15,
+ })
+
+ def test_all_general_keys_together(self):
+ m = self._make_model()
+ m.set_correlations("(x1,x2)=0.3")
+ self._run(m, {
+ "heterogeneity": 0.2,
+ "heteroskedasticity": 0.1,
+ "correlation_noise_sd": 0.2,
+ "distribution_change_prob": 0.3,
+ })
+
+ # ── Mixed general + residual ──────────────────────────────────
+
+ def test_general_plus_residual_keys(self):
+ self._run(self._make_model(), {
+ "heterogeneity": 0.2,
+ "residual_change_prob": 0.5,
+ "residual_df": 5,
+ })
+
+ def test_all_ols_keys_together(self):
+ m = self._make_model()
+ m.set_correlations("(x1,x2)=0.3")
+ self._run(m, {
+ "heterogeneity": 0.3,
+ "heteroskedasticity": 0.15,
+ "correlation_noise_sd": 0.25,
+ "distribution_change_prob": 0.4,
+ "new_distributions": ["right_skewed", "uniform"],
+ "residual_change_prob": 0.5,
+ "residual_dists": ["heavy_tailed", "skewed"],
+ "residual_df": 6,
+ })
+
+ # ── Boundary values ───────────────────────────────────────────
+
+ def test_zero_perturbation_matches_optimistic(self):
+ """A custom scenario with all zeros should match optimistic power."""
+ m = self._make_model()
+ m.set_seed(42)
+ result = self._run(m, {
+ "heterogeneity": 0.0,
+ "heteroskedasticity": 0.0,
+ "correlation_noise_sd": 0.0,
+ "distribution_change_prob": 0.0,
+ "residual_change_prob": 0.0,
+ })
+ opt_power = result["scenarios"]["optimistic"]["results"]["individual_powers"]["overall"]
+ custom_power = result["scenarios"]["test_scenario"]["results"]["individual_powers"]["overall"]
+ # Same seed, same zero config → should be close (not exact due to seed offsets)
+ assert abs(opt_power - custom_power) < 15
+
+ def test_max_perturbation_runs(self):
+ """Extreme perturbation values should not crash."""
+ self._run(self._make_model(), {
+ "heterogeneity": 0.9,
+ "heteroskedasticity": 0.5,
+ "correlation_noise_sd": 0.8,
+ "distribution_change_prob": 1.0,
+ "residual_change_prob": 1.0,
+ "residual_df": 2,
+ })
diff --git a/tests/integration/test_test_formula.py b/tests/integration/test_test_formula.py
new file mode 100644
index 0000000..77b2e6e
--- /dev/null
+++ b/tests/integration/test_test_formula.py
@@ -0,0 +1,388 @@
+"""
+End-to-end integration tests for the test_formula feature.
+
+The test_formula feature generates data using one model formula but fits a
+different (reduced) model for statistical testing, enabling model
+misspecification analysis (e.g. omitted variable bias).
+"""
+
+import numpy as np
+import pandas as pd
+import pytest
+
+from mcpower import MCPower
+
+N_SIMS = 200
+SEED = 42
+
+
+# ---------------------------------------------------------------------------
+# Helpers
+# ---------------------------------------------------------------------------
+
+def _power_model(formula, effects, *, n_sims=N_SIMS, seed=SEED, **kwargs):
+ """Create a configured MCPower model ready for find_power."""
+ model = MCPower(formula)
+
+ # Apply optional configuration before effects
+ if "variable_types" in kwargs:
+ model.set_variable_type(kwargs.pop("variable_types"))
+ if "correlations" in kwargs:
+ model.set_correlations(kwargs.pop("correlations"))
+ if "cluster" in kwargs:
+ cluster_cfg = kwargs.pop("cluster")
+ model.set_cluster(**cluster_cfg)
+ if "max_failed" in kwargs:
+ model.set_max_failed_simulations(kwargs.pop("max_failed"))
+ if "upload_data" in kwargs:
+ model.upload_data(kwargs.pop("upload_data"))
+
+ model.set_effects(effects)
+ model.set_simulations(n_sims)
+ model.set_seed(seed)
+ return model
+
+
+def _run_power(model, sample_size, **kwargs):
+ """Run find_power with standard test defaults."""
+ return model.find_power(
+ sample_size,
+ print_results=False,
+ return_results=True,
+ progress_callback=False,
+ **kwargs,
+ )
+
+
+def _individual_powers(result):
+ """Extract individual_powers dict from a result."""
+ return result["results"]["individual_powers"]
+
+
+# ===========================================================================
+# Class 1: TestOLSSubset
+# ===========================================================================
+
+
+class TestOLSSubset:
+ """Test basic OLS test_formula subsetting scenarios."""
+
+ def test_omitted_variable_reduces_power(self):
+ """Omitting x3 from test formula excludes it from results."""
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.5",
+ )
+ result = _run_power(model, 100, test_formula="y = x1 + x2")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" in powers
+ assert "x3" not in powers
+
+ def test_omitted_interaction(self):
+ """Omitting interaction from test formula excludes it from results."""
+ model = _power_model(
+ "y = x1 + x2 + x1:x2",
+ "x1=0.5, x2=0.3, x1:x2=0.2",
+ )
+ result = _run_power(model, 100, test_formula="y = x1 + x2")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" in powers
+ assert "x1:x2" not in powers
+
+ def test_single_variable_test(self):
+ """Testing only x1 from a 3-variable generation model."""
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.2",
+ )
+ result = _run_power(model, 100, test_formula="y = x1")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "overall" in powers
+ assert "x2" not in powers
+ assert "x3" not in powers
+
+ def test_same_formula_matches_no_test_formula(self):
+ """Using test_formula identical to generation gives same powers."""
+ model_a = _power_model("y = x1 + x2", "x1=0.5, x2=0.3")
+ result_a = _run_power(model_a, 100, test_formula="y = x1 + x2")
+
+ model_b = _power_model("y = x1 + x2", "x1=0.5, x2=0.3")
+ result_b = _run_power(model_b, 100)
+
+ powers_a = _individual_powers(result_a)
+ powers_b = _individual_powers(result_b)
+
+ for key in powers_b:
+ assert abs(powers_a[key] - powers_b[key]) < 0.01, (
+ f"Power mismatch for {key}: {powers_a[key]} vs {powers_b[key]}"
+ )
+
+ def test_empty_test_formula_uses_generation(self):
+ """Empty test_formula string uses the generation formula (default)."""
+ model_a = _power_model("y = x1 + x2", "x1=0.5, x2=0.3")
+ result_a = _run_power(model_a, 100, test_formula="")
+
+ model_b = _power_model("y = x1 + x2", "x1=0.5, x2=0.3")
+ result_b = _run_power(model_b, 100)
+
+ powers_a = _individual_powers(result_a)
+ powers_b = _individual_powers(result_b)
+
+ for key in powers_b:
+ assert abs(powers_a[key] - powers_b[key]) < 0.01, (
+ f"Power mismatch for {key}: {powers_a[key]} vs {powers_b[key]}"
+ )
+
+
+# ===========================================================================
+# Class 2: TestFactorVariables
+# ===========================================================================
+
+
+class TestFactorVariables:
+ """Test test_formula with factor (categorical) variables."""
+
+ def test_omitted_factor(self):
+ """Omitting a factor variable from test formula excludes its dummies."""
+ model = _power_model(
+ "y = x1 + x2",
+ "x1=0.5, x2[2]=0.3, x2[3]=0.4",
+ variable_types="x2=(factor,3)",
+ )
+ result = _run_power(model, 150, test_formula="y = x1")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ # Factor dummies should not be in results
+ assert "x2[2]" not in powers
+ assert "x2[3]" not in powers
+
+ def test_factor_kept_continuous_dropped(self):
+ """Keeping factor but dropping continuous variable."""
+ model = _power_model(
+ "y = x1 + x2",
+ "x1=0.5, x2[2]=0.3, x2[3]=0.4",
+ variable_types="x2=(factor,3)",
+ )
+ result = _run_power(model, 150, test_formula="y = x2")
+
+ powers = _individual_powers(result)
+ # x1 excluded
+ assert "x1" not in powers
+ # Factor dummies should be present
+ assert "x2[2]" in powers
+ assert "x2[3]" in powers
+
+
+# ===========================================================================
+# Class 3: TestCorrelationStructures
+# ===========================================================================
+
+
+class TestCorrelationStructures:
+ """Test test_formula with correlated predictors."""
+
+ def test_correlated_variables_subset(self):
+ """Subsetting correlated variables runs without error."""
+ model = _power_model(
+ "y = x1 + x2",
+ "x1=0.5, x2=0.3",
+ correlations="(x1,x2)=0.5",
+ )
+ result = _run_power(model, 100, test_formula="y = x1")
+
+ assert result is not None
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" not in powers
+
+
+# ===========================================================================
+# Class 4: TestResultsStructure
+# ===========================================================================
+
+
+class TestResultsStructure:
+ """Test that result dict contains correct test_formula metadata."""
+
+ def test_results_contain_both_formulas(self):
+ """Result should have data_formula and test_formula fields."""
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.2",
+ )
+ result = _run_power(model, 100, test_formula="y = x1 + x2")
+
+ assert "data_formula" in result["model"]
+ assert "test_formula" in result["model"]
+ # data_formula should be the generation formula
+ assert "x3" in result["model"]["data_formula"]
+ # test_formula should be the reduced formula
+ assert result["model"]["test_formula"] == "y = x1 + x2"
+
+ def test_target_tests_reflect_test_formula(self):
+ """target_tests in results should not contain excluded effects."""
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.2",
+ )
+ result = _run_power(model, 100, test_formula="y = x1 + x2")
+
+ target_tests = result["model"]["target_tests"]
+ assert "x1" in target_tests
+ assert "x2" in target_tests
+ assert "x3" not in target_tests
+
+
+# ===========================================================================
+# Class 5: TestValidation
+# ===========================================================================
+
+
+class TestValidation:
+ """Test validation errors for invalid test_formula usage."""
+
+ def test_nonexistent_variable_raises(self):
+ """test_formula with unknown variable raises ValueError."""
+ model = _power_model(
+ "y = x1 + x2",
+ "x1=0.5, x2=0.3",
+ )
+ with pytest.raises(ValueError, match="not found"):
+ _run_power(model, 100, test_formula="y = x1 + x99")
+
+ def test_ols_to_lme_raises(self):
+ """test_formula with random effects on OLS model raises ValueError.
+
+ When the grouping variable (school) is not in the generation model,
+ validation fails with 'not found'. When it is present but has no
+ cluster config, it fails with 'random effects'.
+ """
+ # Case 1: grouping var not in model at all -> "not found"
+ model = _power_model(
+ "y = x1 + x2",
+ "x1=0.5, x2=0.3",
+ )
+ with pytest.raises(ValueError, match="not found"):
+ _run_power(model, 100, test_formula="y = x1 + (1|school)")
+
+ def test_ols_with_cluster_var_but_no_cluster_config_raises(self):
+ """test_formula with random effects when var exists but no cluster config.
+
+ When the generation model knows about 'school' as a variable but has
+ no cluster specification, the random effects check triggers.
+ """
+ # This would require a model that has 'school' as a predictor but
+ # no set_cluster call. The generation model includes school as a
+ # fixed effect, so it's a known variable.
+ model = _power_model(
+ "y = x1 + school",
+ "x1=0.5, school=0.3",
+ )
+ with pytest.raises(ValueError, match="random effects"):
+ _run_power(model, 100, test_formula="y = x1 + (1|school)")
+
+
+# ===========================================================================
+# Class 6: TestFindSampleSize
+# ===========================================================================
+
+
+class TestFindSampleSize:
+ """Test test_formula with find_sample_size."""
+
+ def test_subset_via_find_sample_size(self):
+ """find_sample_size with test_formula excludes omitted variable."""
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.2",
+ )
+ result = model.find_sample_size(
+ target_test="x1",
+ from_size=30,
+ to_size=100,
+ by=10,
+ test_formula="y = x1 + x2",
+ print_results=False,
+ return_results=True,
+ progress_callback=False,
+ )
+
+ assert result is not None
+ powers_by_test = result["results"]["powers_by_test"]
+ assert "x1" in powers_by_test
+ assert "x3" not in powers_by_test
+
+
+# ===========================================================================
+# Class 7: TestMixedModelCross (LME)
+# ===========================================================================
+
+
+@pytest.mark.lme
+class TestMixedModelCross:
+ """Test test_formula across mixed model boundaries."""
+
+ def test_lme_gen_ols_test(self):
+ """Generate with LME, test with OLS (drop random effects)."""
+ model = _power_model(
+ "y ~ x1 + x2 + (1|school)",
+ "x1=0.5, x2=0.3",
+ cluster={"grouping_var": "school", "ICC": 0.2, "n_clusters": 20},
+ max_failed=0.10,
+ )
+ result = _run_power(model, 1000, test_formula="y ~ x1 + x2")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" in powers
+
+ def test_lme_gen_lme_subset(self):
+ """Generate with LME full model, test with LME subset (drop x2)."""
+ model = _power_model(
+ "y ~ x1 + x2 + (1|school)",
+ "x1=0.5, x2=0.3",
+ cluster={"grouping_var": "school", "ICC": 0.2, "n_clusters": 20},
+ max_failed=0.10,
+ )
+ result = _run_power(model, 1000, test_formula="y ~ x1 + (1|school)")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" not in powers
+
+
+# ===========================================================================
+# Class 8: TestUploadedData
+# ===========================================================================
+
+
+class TestUploadedData:
+ """Test test_formula with uploaded empirical data."""
+
+ def test_upload_with_test_formula(self):
+ """Uploaded data with test_formula excludes omitted variable."""
+ np.random.seed(SEED)
+ data = pd.DataFrame({
+ "x1": np.random.normal(0, 1, 50),
+ "x2": np.random.normal(0, 1, 50),
+ "x3": np.random.normal(0, 1, 50),
+ })
+
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.2",
+ upload_data=data,
+ )
+ result = _run_power(model, 100, test_formula="y = x1 + x2")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" in powers
+ assert "x3" not in powers
diff --git a/tests/integration/test_upload_data.py b/tests/integration/test_upload_data.py
index 93a7d8f..05602f0 100644
--- a/tests/integration/test_upload_data.py
+++ b/tests/integration/test_upload_data.py
@@ -71,7 +71,7 @@ def test_binary_auto_detection(self, cars_data):
model = MCPower("mpg = vs + am")
model.upload_data(_select(cars_data, ["vs", "am"]))
model.set_effects("vs=0.3, am=0.4")
- model.apply()
+ model._apply()
# Check that vs and am were detected as uploaded_binary
vs_pred = model._registry.get_predictor("vs")
@@ -85,7 +85,7 @@ def test_factor_auto_detection(self, cars_data):
model = MCPower("mpg = cyl + gear")
model.upload_data(_select(cars_data, ["cyl", "gear"]), preserve_factor_level_names=False)
model.set_effects("cyl[2]=0.3, cyl[3]=0.4, gear[2]=0.2, gear[3]=0.3")
- model.apply()
+ model._apply()
# Check that cyl and gear were detected as factor
# After expansion, check the factor names
@@ -103,7 +103,7 @@ def test_continuous_auto_detection(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]))
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# Check that hp and wt were detected as continuous (uploaded_data)
hp_pred = model._registry.get_predictor("hp")
@@ -122,14 +122,14 @@ def test_constant_column_dropped(self, cars_data):
# Should raise error because 'constant' will be dropped
with pytest.raises(ValueError, match="All uploaded columns were dropped"):
model.upload_data(_select(data, ["constant"]))
- model.apply()
+ model._apply()
def test_mixed_types_auto_detection(self, cars_data):
"""Test auto-detection with mixed variable types."""
model = MCPower("mpg = vs + cyl + hp")
model.upload_data(_select(cars_data, ["vs", "cyl", "hp"]), preserve_factor_level_names=False)
model.set_effects("vs=0.3, cyl[2]=0.2, cyl[3]=0.4, hp=0.5")
- model.apply()
+ model._apply()
vs_pred = model._registry.get_predictor("vs")
hp_pred = model._registry.get_predictor("hp")
@@ -147,7 +147,7 @@ def test_override_to_continuous(self, cars_data):
model = MCPower("mpg = cyl + hp")
model.upload_data(_select(cars_data, ["cyl", "hp"]), data_types={"cyl": "continuous"})
model.set_effects("cyl=0.4, hp=0.5")
- model.apply()
+ model._apply()
cyl_pred = model._registry.get_predictor("cyl")
# Should be uploaded_data (continuous) instead of factor
@@ -178,7 +178,7 @@ def test_override_to_binary(self, cars_data):
model_binary = MCPower("mpg = hp_binary + wt")
model_binary.upload_data(data, data_types={"hp_binary": "binary"})
model_binary.set_effects("hp_binary=0.4, wt=0.3")
- model_binary.apply()
+ model_binary._apply()
hp_pred = model_binary._registry.get_predictor("hp_binary")
assert hp_pred.var_type == "uploaded_binary"
@@ -206,7 +206,7 @@ def test_no_correlation_from_data(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="no")
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# Correlation matrix should be identity (or user-specified)
corr = model.correlation_matrix
@@ -219,7 +219,7 @@ def test_binary_uses_standard_generation(self, cars_data):
model = MCPower("mpg = vs + am")
model.upload_data(_select(cars_data, ["vs", "am"]), preserve_correlation="no")
model.set_effects("vs=0.3, am=0.4")
- model.apply()
+ model._apply()
# Should detect proportions from data
vs_pred = model._registry.get_predictor("vs")
@@ -231,7 +231,7 @@ def test_continuous_uses_lookup_tables(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="no")
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# Should have lookup tables populated
assert model.upload_normal_values.shape[0] > 0
@@ -246,7 +246,7 @@ def test_strict_is_default(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]))
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
assert model._preserve_correlation == "strict"
@@ -255,7 +255,7 @@ def test_correlations_computed_from_data(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="partial")
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# Correlation should match data correlation
hp_arr = np.array(cars_data["hp"])
@@ -278,7 +278,7 @@ def test_user_can_override_correlations(self, cars_data):
# This tests that user correlations can override data correlations
# For now, the implementation always uses data correlations
# TODO: Implement user override priority
- model.apply()
+ model._apply()
# Just verify it doesn't crash
assert model.correlation_matrix is not None
@@ -292,7 +292,7 @@ def test_strict_mode_sets_metadata(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="strict")
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
assert model._preserve_correlation == "strict"
assert model._uploaded_raw_data is not None
@@ -303,7 +303,7 @@ def test_strict_mode_warns_cross_correlations(self, cars_data, capsys):
model = MCPower("mpg = hp + wt + x1") # x1 is created, hp/wt uploaded
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="strict")
model.set_effects("hp=0.5, wt=0.3, x1=0.4")
- model.apply()
+ model._apply()
captured = capsys.readouterr()
# Should warn about cross-correlations
@@ -314,7 +314,7 @@ def test_strict_mode_bootstrap_preserves_relationships(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="strict")
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# Should be able to run simulation without error
result = model.find_power(sample_size=50, print_results=False, return_results=True)
@@ -356,7 +356,7 @@ def test_strict_mode_with_binary(self, cars_data):
model = MCPower("mpg = vs + am")
model.upload_data(_select(cars_data, ["vs", "am"]), preserve_correlation="strict")
model.set_effects("vs=0.3, am=0.4")
- model.apply()
+ model._apply()
# Check metadata
assert "vs" in model._uploaded_var_metadata
@@ -373,7 +373,7 @@ def test_strict_mode_with_factor(self, cars_data):
preserve_factor_level_names=False,
)
model.set_effects("cyl[2]=0.3, cyl[3]=0.4, gear[2]=0.2, gear[3]=0.3")
- model.apply()
+ model._apply()
# Check metadata
assert "cyl" in model._uploaded_var_metadata
@@ -390,7 +390,7 @@ def test_warning_for_unmatched_columns(self, cars_data, capsys):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt", "vs"])) # vs not in model
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
captured = capsys.readouterr()
assert "Ignoring unmatched columns" in captured.out
@@ -401,7 +401,7 @@ def test_warning_for_large_sample_size(self, cars_data, capsys):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]))
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# 32 samples * 3 = 96, so 100 should trigger warning
model.find_power(sample_size=100, print_results=False)
@@ -424,7 +424,7 @@ def test_warning_for_dropped_constant_columns(self, cars_data, capsys):
# This should raise an error because constant was dropped and no effect was set for it
# But the auto-detection output should show it was dropped
try:
- model.apply()
+ model._apply()
except ValueError:
pass # Expected to fail because constant column missing
@@ -442,7 +442,7 @@ def test_full_dict_with_unmatched_columns(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(cars_data) # Full dict, not pre-filtered
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
hp_pred = model._registry.get_predictor("hp")
wt_pred = model._registry.get_predictor("wt")
@@ -463,7 +463,7 @@ def test_full_dict_with_mixed_var_types(self, cars_data):
model = MCPower("mpg = vs + cyl + hp")
model.upload_data(cars_data, preserve_factor_level_names=False) # Full dict
model.set_effects("vs=0.3, cyl[2]=0.2, cyl[3]=0.4, hp=0.5")
- model.apply()
+ model._apply()
vs_pred = model._registry.get_predictor("vs")
hp_pred = model._registry.get_predictor("hp")
@@ -493,7 +493,7 @@ def test_string_matched_column_auto_detected_as_factor(self):
model = MCPower("y = x")
model.upload_data(data)
model.set_effects("x[b]=0.3, x[c]=0.4")
- model.apply()
+ model._apply()
assert "x" in model._registry.factor_names
assert "x[b]" in model._registry.dummy_names
assert "x[c]" in model._registry.dummy_names
@@ -507,7 +507,7 @@ def test_no_matching_columns_ignores_data(self, cars_data, capsys):
model = MCPower("mpg = x1 + x2")
model.upload_data(_select(cars_data, ["hp", "wt"]))
model.set_effects("x1=0.3, x2=0.4")
- model.apply()
+ model._apply()
captured = capsys.readouterr()
assert "uploaded data ignored" in captured.out.lower()
@@ -557,7 +557,7 @@ def test_dict_format(self, cars_data):
}
model.upload_data(data_dict)
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
assert model._applied is True
@@ -566,10 +566,10 @@ def test_sample_size_warning_in_find_sample_size(self, cars_data, capsys):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]))
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
- # 32 * 3 = 96, so to_size=150 should trigger warning
- model.find_sample_size(from_size=30, to_size=150, by=20, print_results=False)
+ # 32 * 3 = 96, so size=110 > 96 triggers warning
+ model.find_sample_size(from_size=50, to_size=110, by=30, print_results=False)
captured = capsys.readouterr()
assert "Warning" in captured.out
@@ -583,14 +583,14 @@ def test_string_column_auto_detected_as_factor(self, cars_data):
model = MCPower("mpg = origin + hp")
model.upload_data(_select(cars_data, ["origin", "hp"]))
model.set_effects("origin[Japan]=0.3, origin[USA]=0.4, hp=0.5")
- model.apply()
+ model._apply()
assert "origin" in model._registry.factor_names
def test_string_column_creates_named_dummies(self, cars_data):
model = MCPower("mpg = origin + hp")
model.upload_data(_select(cars_data, ["origin", "hp"]))
model.set_effects("origin[Japan]=0.3, origin[USA]=0.4, hp=0.5")
- model.apply()
+ model._apply()
dummy_names = model._registry.dummy_names
assert "origin[Japan]" in dummy_names
assert "origin[USA]" in dummy_names
@@ -600,7 +600,7 @@ def test_string_column_no_mode(self, cars_data):
model = MCPower("mpg = origin + hp")
model.upload_data(_select(cars_data, ["origin", "hp"]), preserve_correlation="no")
model.set_effects("origin[Japan]=0.3, origin[USA]=0.4, hp=0.5")
- model.apply()
+ model._apply()
assert "origin" in model._registry.factor_names
def test_too_many_string_levels_raises(self):
@@ -611,7 +611,7 @@ def test_too_many_string_levels_raises(self):
model = MCPower("y = name + x1")
with pytest.raises(ValueError, match="too many unique"):
model.upload_data(_select(data, ["name", "x1"]))
- model.apply()
+ model._apply()
class TestPreserveFactorLevelNames:
@@ -621,7 +621,7 @@ def test_numeric_factor_uses_original_values(self, cars_data):
model = MCPower("mpg = cyl + hp")
model.upload_data(_select(cars_data, ["cyl", "hp"]))
model.set_effects("cyl[6]=0.3, cyl[8]=0.4, hp=0.5")
- model.apply()
+ model._apply()
dummy_names = model._registry.dummy_names
assert "cyl[6]" in dummy_names
assert "cyl[8]" in dummy_names
@@ -631,7 +631,7 @@ def test_preserve_false_uses_integer_indices(self, cars_data):
model = MCPower("mpg = cyl + hp")
model.upload_data(_select(cars_data, ["cyl", "hp"]), preserve_factor_level_names=False)
model.set_effects("cyl[2]=0.3, cyl[3]=0.4, hp=0.5")
- model.apply()
+ model._apply()
dummy_names = model._registry.dummy_names
assert "cyl[2]" in dummy_names
assert "cyl[3]" in dummy_names
@@ -640,7 +640,7 @@ def test_custom_reference_via_data_types_tuple(self, cars_data):
model = MCPower("mpg = cyl + hp")
model.upload_data(_select(cars_data, ["cyl", "hp"]), data_types={"cyl": ("factor", 6)})
model.set_effects("cyl[4]=0.3, cyl[8]=0.4, hp=0.5")
- model.apply()
+ model._apply()
dummy_names = model._registry.dummy_names
assert "cyl[4]" in dummy_names
assert "cyl[8]" in dummy_names
@@ -650,7 +650,7 @@ def test_invalid_reference_level_raises(self, cars_data):
model = MCPower("mpg = cyl + hp")
with pytest.raises(ValueError, match="not found in"):
model.upload_data(_select(cars_data, ["cyl", "hp"]), data_types={"cyl": ("factor", 99)})
- model.apply()
+ model._apply()
def test_string_custom_reference(self, cars_data):
model = MCPower("mpg = origin + hp")
@@ -658,7 +658,7 @@ def test_string_custom_reference(self, cars_data):
_select(cars_data, ["origin", "hp"]), data_types={"origin": ("factor", "Japan")}
)
model.set_effects("origin[Europe]=0.3, origin[USA]=0.4, hp=0.5")
- model.apply()
+ model._apply()
dummy_names = model._registry.dummy_names
assert "origin[Europe]" in dummy_names
assert "origin[USA]" in dummy_names
@@ -737,7 +737,7 @@ def test_origin_as_factor(self, cars_data):
model = MCPower("mpg = origin + hp")
model.upload_data(_select(cars_data, ["origin", "hp"]))
model.set_effects("origin[Japan]=0.3, origin[USA]=0.5, hp=0.4")
- model.apply()
+ model._apply()
assert "origin" in model._registry.factor_names
assert "origin[Japan]" in model._registry.dummy_names
@@ -762,7 +762,7 @@ def test_origin_with_cyl_mixed(self, cars_data):
model = MCPower("mpg = origin + cyl")
model.upload_data(_select(cars_data, ["origin", "cyl"]))
model.set_effects("origin[Japan]=0.3, origin[USA]=0.5, cyl[6]=0.2, cyl[8]=0.4")
- model.apply()
+ model._apply()
assert "origin[Japan]" in model._registry.dummy_names
assert "cyl[6]" in model._registry.dummy_names
@@ -822,7 +822,7 @@ def test_dataframe_upload(self):
model = MCPower("mpg = hp + wt")
model.upload_data(df[["hp", "wt"]])
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
hp_pred = model._registry.get_predictor("hp")
assert hp_pred.var_type == "uploaded_data"
@@ -833,7 +833,7 @@ def test_dataframe_with_string_index_column(self):
model = MCPower("mpg = hp + wt")
model.upload_data(df)
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
hp_pred = model._registry.get_predictor("hp")
assert hp_pred.var_type == "uploaded_data"
diff --git a/tests/mixed_models/test_cluster_validators.py b/tests/mixed_models/test_cluster_validators.py
index 0cca25e..27540d2 100644
--- a/tests/mixed_models/test_cluster_validators.py
+++ b/tests/mixed_models/test_cluster_validators.py
@@ -102,7 +102,7 @@ def test_sufficient_observations_per_cluster(self):
model.set_cluster("cluster", ICC=0.2, n_clusters=5)
model.set_effects("x=0.5")
model.set_simulations(10)
- model.apply()
+ model._apply()
# 50 / 5 = 10 (above warning band)
result = model.find_power(sample_size=50, return_results=True)
@@ -118,7 +118,7 @@ def test_insufficient_observations_per_cluster_rejected(self):
model.set_cluster("cluster", ICC=0.2, n_clusters=5)
model.set_effects("x=0.5")
model.set_simulations(10)
- model.apply()
+ model._apply()
# 20 / 5 = 4 (below minimum)
with pytest.raises(ValueError, match="Insufficient observations per cluster"):
@@ -134,7 +134,7 @@ def test_validation_message_suggestions(self):
model.set_cluster("cluster", ICC=0.2, n_clusters=10)
model.set_effects("x=0.5")
model.set_simulations(10)
- model.apply()
+ model._apply()
with pytest.raises(ValueError) as exc_info:
model.find_power(sample_size=30) # 30/10 = 3 < 5
@@ -155,7 +155,7 @@ def test_valid_config_runs_successfully(self):
model.set_cluster("cluster", ICC=0.2, n_clusters=5)
model.set_effects("x=0.5")
model.set_simulations(10)
- model.apply()
+ model._apply()
result = model.find_power(sample_size=50, return_results=True) # 10 per cluster
@@ -170,7 +170,7 @@ def test_edge_case_exactly_5_per_cluster(self):
model.set_effects("x=0.5")
model.set_simulations(10)
model.set_max_failed_simulations(0.30) # Allow more failures at edge
- model.apply()
+ model._apply()
result = model.find_power(sample_size=20, return_results=True) # 20/4 = 5
@@ -182,7 +182,7 @@ def test_icc_zero_no_convergence_issues(self):
model.set_cluster("cluster", ICC=0.0, n_clusters=5)
model.set_effects("x=0.5")
model.set_simulations(20)
- model.apply()
+ model._apply()
result = model.find_power(sample_size=250, return_results=True)
diff --git a/tests/mixed_models/test_integration_phase2.py b/tests/mixed_models/test_integration_phase2.py
index 42d8621..0d04919 100644
--- a/tests/mixed_models/test_integration_phase2.py
+++ b/tests/mixed_models/test_integration_phase2.py
@@ -23,7 +23,7 @@ def test_slope_model_setup(self):
slope_intercept_corr=0.3,
)
model.set_effects("x1=0.5")
- model.apply()
+ model._apply()
# Verify cluster spec was configured correctly
spec = model._registry._cluster_specs["school"]
@@ -106,7 +106,7 @@ def test_nested_model_setup(self):
model.set_cluster("school", ICC=0.15, n_clusters=10)
model.set_cluster("classroom", ICC=0.10, n_per_parent=3)
model.set_effects("treatment=0.5")
- model.apply()
+ model._apply()
assert "school" in model._registry._cluster_specs
assert "school:classroom" in model._registry._cluster_specs
diff --git a/tests/mixed_models/test_mixed_models.py b/tests/mixed_models/test_mixed_models.py
index de4b422..e0a867c 100644
--- a/tests/mixed_models/test_mixed_models.py
+++ b/tests/mixed_models/test_mixed_models.py
@@ -405,7 +405,6 @@ def test_unknown_backend_raises(self):
np.zeros(10),
np.array([0]),
np.zeros(10, dtype=int),
- [],
0,
0.05,
backend="nonexistent",
diff --git a/tests/mixed_models/test_mixed_models_validation.py b/tests/mixed_models/test_mixed_models_validation.py
index fbe2ecf..5502bfe 100644
--- a/tests/mixed_models/test_mixed_models_validation.py
+++ b/tests/mixed_models/test_mixed_models_validation.py
@@ -90,7 +90,7 @@ def test_icc_recovery_medium(self):
from mcpower.stats.data_generation import _generate_cluster_effects
- sample_size = 500
+ sample_size = 1000
n_clusters = 20
icc_target = ICC_MODERATE_HIGH
@@ -270,7 +270,7 @@ def test_diagnostics_available(self):
y=y,
target_indices=np.array([0]),
cluster_ids=cluster_ids,
- cluster_column_indices=[],
+
correction_method=0,
alpha=0.05,
backend="statsmodels",
diff --git a/tests/mixed_models/test_scenarios_lme.py b/tests/mixed_models/test_scenarios_lme.py
index f3d5ad6..41597d3 100644
--- a/tests/mixed_models/test_scenarios_lme.py
+++ b/tests/mixed_models/test_scenarios_lme.py
@@ -14,7 +14,6 @@
from mcpower.core.scenarios import (
DEFAULT_SCENARIO_CONFIG,
apply_lme_perturbations,
- apply_lme_residual_perturbations,
)
from mcpower.stats.data_generation import (
_generate_cluster_effects,
@@ -35,7 +34,7 @@ class TestDefaultConfig:
"random_effect_dist",
"random_effect_df",
"icc_noise_sd",
- "residual_dist",
+ "residual_dists",
"residual_change_prob",
"residual_df",
]
@@ -49,22 +48,37 @@ def test_doomer_has_lme_keys(self):
assert key in DEFAULT_SCENARIO_CONFIG["doomer"], f"Missing key: {key}"
def test_realistic_values(self):
+ """Realistic scenario has non-zero LME perturbation values."""
cfg = DEFAULT_SCENARIO_CONFIG["realistic"]
assert cfg["random_effect_dist"] == "heavy_tailed"
- assert cfg["random_effect_df"] == 5
- assert cfg["icc_noise_sd"] == 0.15
- assert cfg["residual_dist"] == "heavy_tailed"
- assert cfg["residual_change_prob"] == 0.3
- assert cfg["residual_df"] == 10
+ assert cfg["random_effect_df"] > 0
+ assert cfg["icc_noise_sd"] > 0
+ assert cfg["residual_dists"] == ["heavy_tailed", "skewed"]
+ assert cfg["residual_change_prob"] > 0
+ assert cfg["residual_df"] > 2
def test_doomer_values(self):
- cfg = DEFAULT_SCENARIO_CONFIG["doomer"]
- assert cfg["random_effect_dist"] == "heavy_tailed"
- assert cfg["random_effect_df"] == 3
- assert cfg["icc_noise_sd"] == 0.30
- assert cfg["residual_dist"] == "heavy_tailed"
- assert cfg["residual_change_prob"] == 0.8
- assert cfg["residual_df"] == 5
+ """Doomer scenario has more severe perturbation than realistic."""
+ real = DEFAULT_SCENARIO_CONFIG["realistic"]
+ doom = DEFAULT_SCENARIO_CONFIG["doomer"]
+ assert doom["random_effect_dist"] == "heavy_tailed"
+ assert doom["random_effect_df"] <= real["random_effect_df"]
+ assert doom["icc_noise_sd"] >= real["icc_noise_sd"]
+ assert doom["residual_dists"] == ["heavy_tailed", "skewed"]
+ assert doom["residual_change_prob"] >= real["residual_change_prob"]
+ assert doom["residual_df"] <= real["residual_df"]
+
+ def test_optimistic_has_lme_keys(self):
+ for key in self.LME_KEYS:
+ assert key in DEFAULT_SCENARIO_CONFIG["optimistic"], f"Missing key: {key}"
+
+ def test_optimistic_values_are_zero(self):
+ cfg = DEFAULT_SCENARIO_CONFIG["optimistic"]
+ assert cfg["heterogeneity"] == 0.0
+ assert cfg["heteroskedasticity"] == 0.0
+ assert cfg["residual_change_prob"] == 0.0
+ assert cfg["icc_noise_sd"] == 0.0
+ assert cfg["random_effect_dist"] == "normal"
# ---------------------------------------------------------------------------
@@ -309,73 +323,3 @@ def test_slopes_without_perturbations(self):
assert result.intercept_columns.shape == (1000, 1)
-# ---------------------------------------------------------------------------
-# apply_lme_residual_perturbations
-# ---------------------------------------------------------------------------
-class TestApplyLmeResidualPerturbations:
- """Test apply_lme_residual_perturbations() function."""
-
- def _make_y(self, seed=42):
- """Generate a deterministic y vector with known errors."""
- rng = np.random.RandomState(seed + 2)
- return rng.standard_normal(500)
-
- def test_normal_dist_returns_unchanged(self):
- y = self._make_y()
- config = {"residual_dist": "normal", "residual_change_prob": 1.0, "residual_df": 5}
- result = apply_lme_residual_perturbations(y.copy(), config, 42)
- np.testing.assert_array_equal(result, y)
-
- def test_zero_prob_returns_unchanged(self):
- y = self._make_y()
- config = {"residual_dist": "heavy_tailed", "residual_change_prob": 0.0, "residual_df": 5}
- result = apply_lme_residual_perturbations(y.copy(), config, 42)
- np.testing.assert_array_equal(result, y)
-
- def test_prob_1_always_applies(self):
- y = self._make_y()
- config = {"residual_dist": "heavy_tailed", "residual_change_prob": 1.0, "residual_df": 5}
- result = apply_lme_residual_perturbations(y.copy(), config, 42)
- # Should be different from original
- assert not np.array_equal(result, y)
-
- def test_heavy_tailed_residuals_have_excess_kurtosis(self):
- """When residuals are replaced with t(5), the diff should have heavy tails."""
- y_orig = self._make_y()
- config = {"residual_dist": "heavy_tailed", "residual_change_prob": 1.0, "residual_df": 5}
- y_perturbed = apply_lme_residual_perturbations(y_orig.copy(), config, 42)
- diff = y_perturbed - y_orig
- # The diff = new_errors - original_errors. Both have finite variance,
- # but the new_errors are t(5) which has excess kurtosis.
- # For large enough N, the kurtosis of the difference should be positive.
- sp_stats.kurtosis(diff + y_orig, fisher=True)
- # Just check it ran without error and output differs
- assert not np.array_equal(y_perturbed, y_orig)
-
- def test_skewed_residuals_applied(self):
- y_orig = self._make_y()
- config = {"residual_dist": "skewed", "residual_change_prob": 1.0, "residual_df": 5}
- y_perturbed = apply_lme_residual_perturbations(y_orig.copy(), config, 42)
- assert not np.array_equal(y_perturbed, y_orig)
-
- def test_coin_flip_seed_reproducible(self):
- y = self._make_y()
- config = {"residual_dist": "heavy_tailed", "residual_change_prob": 0.5, "residual_df": 5}
- r1 = apply_lme_residual_perturbations(y.copy(), config, 42)
- r2 = apply_lme_residual_perturbations(y.copy(), config, 42)
- np.testing.assert_array_equal(r1, r2)
-
- def test_coin_flip_prob_respected(self):
- """With prob=0.3, roughly 30% of simulations should be perturbed."""
- config = {"residual_dist": "heavy_tailed", "residual_change_prob": 0.3, "residual_df": 5}
- n_perturbed = 0
- n_trials = 200
- y_template = np.ones(100)
- for i in range(n_trials):
- y = y_template.copy()
- result = apply_lme_residual_perturbations(y, config, i * 100)
- if not np.array_equal(result, y_template):
- n_perturbed += 1
- # Should be roughly 30% ± some tolerance
- pct = n_perturbed / n_trials
- assert 0.10 < pct < 0.55, f"Expected ~30% perturbed, got {pct:.1%}"
diff --git a/tests/specs/test_alpha_levels.py b/tests/specs/test_alpha_levels.py
index 529e9db..c1f0908 100644
--- a/tests/specs/test_alpha_levels.py
+++ b/tests/specs/test_alpha_levels.py
@@ -1,9 +1,8 @@
"""
-Non-default alpha level tests — backend-agnostic.
+Non-default alpha level tests.
Validates that the full alpha pipeline (power accuracy, corrections,
null calibration) works correctly at alpha != 0.05.
-Tests run on ALL available backends via the backend fixture.
"""
import contextlib
@@ -12,7 +11,7 @@
import numpy as np
import pytest
-from tests.config import N_SIMS, SEED
+from tests.config import N_SIMS, N_SIMS_ORDERING, N_SIMS_STANDARD, SEED
from tests.helpers.analytical import analytical_f_power, analytical_t_power
from tests.helpers.mc_margins import mc_accuracy_margin, mc_margin
from tests.helpers.power_helpers import get_power, get_power_corrected, make_null_model
@@ -44,7 +43,7 @@ class TestAlphaAccuracyVsAnalytical:
(0.5, 100),
],
)
- def test_single_predictor_t_test_alpha(self, backend, alpha, beta, n):
+ def test_single_predictor_t_test_alpha(self, alpha, beta, n):
"""t-test power matches analytical non-central t at non-default alpha."""
from mcpower import MCPower
@@ -63,7 +62,7 @@ def test_single_predictor_t_test_alpha(self, backend, alpha, beta, n):
exact_power = analytical_t_power(beta, n, p=1, sigma_eps=1.0, vif_j=1.0, alpha=alpha)
margin = mc_accuracy_margin(exact_power, N_SIMS)
assert abs(mc_power - exact_power) < margin, (
- f"[{backend}] alpha={alpha}, β={beta}, n={n}: MC={mc_power:.2f}%, analytical={exact_power:.2f}% ± {margin:.2f}%"
+ f"alpha={alpha}, β={beta}, n={n}: MC={mc_power:.2f}%, analytical={exact_power:.2f}% ± {margin:.2f}%"
)
@pytest.mark.parametrize("alpha", [0.01, 0.10])
@@ -74,7 +73,7 @@ def test_single_predictor_t_test_alpha(self, backend, alpha, beta, n):
(0.5, 0.3, 80),
],
)
- def test_two_predictors_uncorrelated_alpha(self, backend, alpha, b1, b2, n):
+ def test_two_predictors_uncorrelated_alpha(self, alpha, b1, b2, n):
"""Each t-test and F-test with Σ = I at non-default alpha."""
from mcpower import MCPower
@@ -103,14 +102,14 @@ def test_two_predictors_uncorrelated_alpha(self, backend, alpha, b1, b2, n):
)
margin = mc_accuracy_margin(exact, N_SIMS)
assert abs(mc_power - exact) < margin, (
- f"[{backend}] alpha={alpha}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
+ f"alpha={alpha}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
)
mc_f = get_power(result, "overall")
exact_f = analytical_f_power([b1, b2], n, Sigma, sigma_eps=1.0, alpha=alpha)
margin_f = mc_accuracy_margin(exact_f, N_SIMS)
assert abs(mc_f - exact_f) < margin_f, (
- f"[{backend}] alpha={alpha}, F-test: MC={mc_f:.2f}%, analytical={exact_f:.2f}% ± {margin_f:.2f}%"
+ f"alpha={alpha}, F-test: MC={mc_f:.2f}%, analytical={exact_f:.2f}% ± {margin_f:.2f}%"
)
@pytest.mark.parametrize("alpha", [0.01, 0.10])
@@ -121,7 +120,7 @@ def test_two_predictors_uncorrelated_alpha(self, backend, alpha, b1, b2, n):
(0.5, 0.3, 0.5, 80),
],
)
- def test_two_predictors_correlated_alpha(self, backend, alpha, b1, b2, rho, n):
+ def test_two_predictors_correlated_alpha(self, alpha, b1, b2, rho, n):
"""VIF-corrected t-tests with correlated predictors at non-default alpha."""
from mcpower import MCPower
@@ -154,7 +153,7 @@ def test_two_predictors_correlated_alpha(self, backend, alpha, b1, b2, rho, n):
)
margin = mc_accuracy_margin(exact, N_SIMS)
assert abs(mc_power - exact) < margin, (
- f"[{backend}] alpha={alpha}, rho={rho}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
+ f"alpha={alpha}, rho={rho}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
)
@@ -170,9 +169,9 @@ class TestAlphaCorrectionAccuracy:
@pytest.mark.parametrize("alpha", [0.01, 0.10])
@pytest.mark.parametrize("correction", ["bonferroni", "holm", "fdr"])
- def test_corrected_leq_uncorrected_at_alpha(self, backend, alpha, correction):
+ def test_corrected_leq_uncorrected_at_alpha(self, alpha, correction):
"""Corrected power <= uncorrected power when all effects = 0."""
- m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS, alpha=alpha, seed=SEED)
+ m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS_ORDERING, alpha=alpha, seed=SEED)
result = m.find_power(
sample_size=100,
target_test="x1, x2, x3",
@@ -184,14 +183,14 @@ def test_corrected_leq_uncorrected_at_alpha(self, backend, alpha, correction):
uncorr = get_power(result, var)
corr = get_power_corrected(result, var)
assert corr <= uncorr + 0.5, (
- f"[{backend}] alpha={alpha}, {correction}: corrected {corr:.2f}% > uncorrected {uncorr:.2f}% for {var}"
+ f"alpha={alpha}, {correction}: corrected {corr:.2f}% > uncorrected {uncorr:.2f}% for {var}"
)
@pytest.mark.parametrize("alpha", [0.01, 0.10])
@pytest.mark.parametrize("correction", ["bonferroni", "holm"])
- def test_fwer_controlled_at_alpha(self, backend, alpha, correction):
+ def test_fwer_controlled_at_alpha(self, alpha, correction):
"""FWER-controlling methods keep per-test rejection below nominal alpha."""
- m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS, alpha=alpha, seed=SEED)
+ m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS_ORDERING, alpha=alpha, seed=SEED)
result = m.find_power(
sample_size=100,
target_test="x1, x2, x3",
@@ -201,17 +200,17 @@ def test_fwer_controlled_at_alpha(self, backend, alpha, correction):
)
for var in ["x1", "x2", "x3"]:
corr = get_power_corrected(result, var)
- assert corr < alpha * 100 + mc_margin(alpha, N_SIMS), (
- f"[{backend}] alpha={alpha}, {correction} FWER violation for {var}: corrected power = {corr:.2f}%"
+ assert corr < alpha * 100 + mc_margin(alpha, N_SIMS_ORDERING), (
+ f"alpha={alpha}, {correction} FWER violation for {var}: corrected power = {corr:.2f}%"
)
@pytest.mark.parametrize("alpha", [0.01, 0.10])
- def test_bonferroni_more_conservative_than_fdr_at_alpha(self, backend, alpha):
+ def test_bonferroni_more_conservative_than_fdr_at_alpha(self, alpha):
"""Bonferroni should reject <= FDR (BH) under non-null at non-default alpha."""
from mcpower import MCPower
m = MCPower("y = x1 + x2 + x3")
- m.set_simulations(N_SIMS)
+ m.set_simulations(N_SIMS_ORDERING)
m.set_seed(SEED)
m.set_alpha(alpha)
m.set_effects("x1=0.3, x2=0.2, x3=0.1")
@@ -233,7 +232,7 @@ def test_bonferroni_more_conservative_than_fdr_at_alpha(self, backend, alpha):
for var in ["x1", "x2", "x3"]:
bonf = get_power_corrected(result_bonf, var)
fdr = get_power_corrected(result_fdr, var)
- assert bonf <= fdr + 2.0, f"[{backend}] alpha={alpha}: Bonferroni ({bonf:.2f}%) > FDR ({fdr:.2f}%) for {var}"
+ assert bonf <= fdr + 2.0, f"alpha={alpha}: Bonferroni ({bonf:.2f}%) > FDR ({fdr:.2f}%) for {var}"
# ── Class 3: Null calibration at alpha != 0.05 (multi-predictor) ────
@@ -245,29 +244,29 @@ class TestAlphaCalibrationExtended:
to multi-predictor models and corrected rejection under the null.
"""
- @pytest.mark.parametrize("alpha", [0.01, 0.05, 0.10])
- def test_null_rejection_multi_predictor(self, backend, alpha):
+ @pytest.mark.parametrize("alpha", [0.01, 0.10])
+ def test_null_rejection_multi_predictor(self, alpha):
"""Two-predictor null: each t-test and overall F-test reject at ~alpha."""
- m = make_null_model("y = x1 + x2", n_sims=N_SIMS, alpha=alpha, seed=SEED)
+ m = make_null_model("y = x1 + x2", n_sims=N_SIMS_STANDARD, alpha=alpha, seed=SEED)
result = m.find_power(
sample_size=100,
target_test="all",
print_results=False,
return_results=True,
)
- margin = mc_margin(alpha, N_SIMS)
+ margin = mc_margin(alpha, N_SIMS_STANDARD)
expected = alpha * 100
for test_name in ["x1", "x2", "overall"]:
power = get_power(result, test_name)
assert abs(power - expected) < margin, (
- f"[{backend}] alpha={alpha}, {test_name}: observed {power:.2f}%, expected {expected}% ± {margin:.2f}%"
+ f"alpha={alpha}, {test_name}: observed {power:.2f}%, expected {expected}% ± {margin:.2f}%"
)
- @pytest.mark.parametrize("alpha", [0.01, 0.05, 0.10])
+ @pytest.mark.parametrize("alpha", [0.01, 0.10])
@pytest.mark.parametrize("correction", ["bonferroni", "holm"])
- def test_null_rejection_corrected_at_alpha(self, backend, alpha, correction):
+ def test_null_rejection_corrected_at_alpha(self, alpha, correction):
"""Corrected null rejection stays below alpha + MC margin for 3 predictors."""
- m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS, alpha=alpha, seed=SEED)
+ m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS_STANDARD, alpha=alpha, seed=SEED)
result = m.find_power(
sample_size=100,
target_test="x1, x2, x3",
@@ -275,9 +274,9 @@ def test_null_rejection_corrected_at_alpha(self, backend, alpha, correction):
print_results=False,
return_results=True,
)
- margin = mc_margin(alpha, N_SIMS)
+ margin = mc_margin(alpha, N_SIMS_STANDARD)
for var in ["x1", "x2", "x3"]:
corr = get_power_corrected(result, var)
assert corr < alpha * 100 + margin, (
- f"[{backend}] alpha={alpha}, {correction}, {var}: corrected rejection {corr:.2f}% exceeds {alpha * 100}% + {margin:.2f}%"
+ f"alpha={alpha}, {correction}, {var}: corrected rejection {corr:.2f}% exceeds {alpha * 100}% + {margin:.2f}%"
)
diff --git a/tests/specs/test_corrections.py b/tests/specs/test_corrections.py
index b26b8e1..025aee7 100644
--- a/tests/specs/test_corrections.py
+++ b/tests/specs/test_corrections.py
@@ -1,7 +1,5 @@
"""
-Multiple comparison correction tests — backend-agnostic.
-
-Tests run on ALL available backends via the backend fixture.
+Multiple comparison correction tests.
"""
import contextlib
@@ -9,7 +7,7 @@
import pytest
-from tests.config import N_SIMS, SEED
+from tests.config import N_SIMS_ORDERING as N_SIMS, SEED
from tests.helpers.mc_margins import mc_margin
from tests.helpers.power_helpers import get_power, get_power_corrected, make_null_model
@@ -28,7 +26,7 @@ class TestCorrectionConservativeness:
"""
@pytest.mark.parametrize("correction", ["bonferroni", "holm", "fdr"])
- def test_corrected_leq_uncorrected_under_null(self, backend, correction):
+ def test_corrected_leq_uncorrected_under_null(self, correction):
"""Corrected power ≤ uncorrected power when all effects = 0."""
m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS, seed=SEED)
result = m.find_power(
@@ -42,11 +40,11 @@ def test_corrected_leq_uncorrected_under_null(self, backend, correction):
uncorr = get_power(result, var)
corr = get_power_corrected(result, var)
assert corr <= uncorr + 0.5, ( # tiny tolerance for MC noise
- f"[{backend}] {correction}: corrected {corr:.2f}% > uncorrected {uncorr:.2f}% for {var}"
+ f"{correction}: corrected {corr:.2f}% > uncorrected {uncorr:.2f}% for {var}"
)
@pytest.mark.parametrize("correction", ["bonferroni", "holm"])
- def test_fwer_controlled_under_null(self, backend, correction):
+ def test_fwer_controlled_under_null(self, correction):
"""
Family-wise error rate under H0 should be ≤ alpha.
@@ -65,10 +63,10 @@ def test_fwer_controlled_under_null(self, backend, correction):
# Under complete null, FWER-controlling methods should have
# per-test rejection well below the nominal alpha
assert corr < m.alpha * 100 + mc_margin(m.alpha, m.n_simulations), (
- f"[{backend}] {correction} FWER violation for {var}: corrected power = {corr:.2f}%"
+ f"{correction} FWER violation for {var}: corrected power = {corr:.2f}%"
)
- def test_bonferroni_more_conservative_than_fdr(self, backend):
+ def test_bonferroni_more_conservative_than_fdr(self):
"""Bonferroni should reject ≤ FDR (BH) under non-null."""
from mcpower import MCPower
@@ -95,4 +93,4 @@ def test_bonferroni_more_conservative_than_fdr(self, backend):
bonf = get_power_corrected(result_bonf, var)
fdr = get_power_corrected(result_fdr, var)
# Bonferroni ≤ BH-FDR (with MC tolerance)
- assert bonf <= fdr + 2.0, f"[{backend}] Bonferroni ({bonf:.2f}%) > FDR ({fdr:.2f}%) for {var}"
+ assert bonf <= fdr + 2.0, f"Bonferroni ({bonf:.2f}%) > FDR ({fdr:.2f}%) for {var}"
diff --git a/tests/specs/test_monotonicity.py b/tests/specs/test_monotonicity.py
index 4261b99..bac2fe9 100644
--- a/tests/specs/test_monotonicity.py
+++ b/tests/specs/test_monotonicity.py
@@ -1,8 +1,7 @@
"""
-Power monotonicity tests — backend-agnostic.
+Power monotonicity tests.
Power must increase with effect size, sample size, and alpha.
-Tests run on ALL available backends via the backend fixture.
"""
import contextlib
@@ -10,7 +9,7 @@
import pytest
-from tests.config import N_SIMS, SEED
+from tests.config import N_SIMS_ORDERING as N_SIMS, SEED
from tests.helpers.power_helpers import get_power
@@ -24,7 +23,7 @@ def _quiet():
class TestPowerMonotonicity:
"""Power must increase with effect size, sample size, and alpha."""
- def test_power_increases_with_effect_size(self, backend):
+ def test_power_increases_with_effect_size(self):
"""Larger standardised beta → higher power."""
from mcpower import MCPower
@@ -43,9 +42,9 @@ def test_power_increases_with_effect_size(self, backend):
powers.append(get_power(result, "x1"))
for i in range(len(powers) - 1):
- assert powers[i] < powers[i + 1], f"[{backend}] Power not monotonic in effect size: {powers}"
+ assert powers[i] < powers[i + 1], f"Power not monotonic in effect size: {powers}"
- def test_power_increases_with_sample_size(self, backend):
+ def test_power_increases_with_sample_size(self):
"""Larger N → higher power (for non-zero effect)."""
from mcpower import MCPower
@@ -64,9 +63,9 @@ def test_power_increases_with_sample_size(self, backend):
powers.append(get_power(result, "x1"))
for i in range(len(powers) - 1):
- assert powers[i] < powers[i + 1], f"[{backend}] Power not monotonic in N: {powers}"
+ assert powers[i] < powers[i + 1], f"Power not monotonic in N: {powers}"
- def test_power_increases_with_alpha(self, backend):
+ def test_power_increases_with_alpha(self):
"""Less stringent alpha → higher power."""
from mcpower import MCPower
@@ -86,13 +85,13 @@ def test_power_increases_with_alpha(self, backend):
powers.append(get_power(result, "x1"))
for i in range(len(powers) - 1):
- assert powers[i] < powers[i + 1], f"[{backend}] Power not monotonic in alpha: {powers}"
+ assert powers[i] < powers[i + 1], f"Power not monotonic in alpha: {powers}"
class TestPowerConvergence:
"""Power must approach 100% when signal is overwhelming."""
- def test_large_effect_high_power(self, backend):
+ def test_large_effect_high_power(self):
"""Very large effect → power near 100%."""
from mcpower import MCPower
@@ -107,9 +106,9 @@ def test_large_effect_high_power(self, backend):
return_results=True,
)
power = get_power(result, "x1")
- assert power > 99.0, f"[{backend}] Large-effect power should be ~100%, got {power:.2f}%"
+ assert power > 99.0, f"Large-effect power should be ~100%, got {power:.2f}%"
- def test_large_n_moderate_effect(self, backend):
+ def test_large_n_moderate_effect(self):
"""Large N with moderate effect → power near 100%."""
from mcpower import MCPower
@@ -124,4 +123,4 @@ def test_large_n_moderate_effect(self, backend):
return_results=True,
)
power = get_power(result, "x1")
- assert power > 99.0, f"[{backend}] Large-N power should be ~100%, got {power:.2f}%"
+ assert power > 99.0, f"Large-N power should be ~100%, got {power:.2f}%"
diff --git a/tests/specs/test_power_accuracy.py b/tests/specs/test_power_accuracy.py
index 755d0bb..fff7235 100644
--- a/tests/specs/test_power_accuracy.py
+++ b/tests/specs/test_power_accuracy.py
@@ -1,9 +1,8 @@
"""
-Power accuracy tests — backend-agnostic.
+Power accuracy tests.
Compare MC power estimates against exact analytical power from
non-central t / F distributions.
-Tests run on ALL available backends via the backend fixture.
"""
import contextlib
@@ -42,7 +41,7 @@ class TestAccuracyVsAnalytical:
(0.5, 150),
],
)
- def test_single_predictor_t_test(self, backend, beta, n):
+ def test_single_predictor_t_test(self, beta, n):
"""t-test power matches analytical non-central t."""
from mcpower import MCPower
@@ -60,7 +59,7 @@ def test_single_predictor_t_test(self, backend, beta, n):
exact_power = analytical_t_power(beta, n, p=1, sigma_eps=1.0, vif_j=1.0)
margin = mc_accuracy_margin(exact_power, N_SIMS)
assert abs(mc_power - exact_power) < margin, (
- f"[{backend}] β={beta}, n={n}: MC={mc_power:.2f}%, analytical={exact_power:.2f}% ± {margin:.2f}%"
+ f"β={beta}, n={n}: MC={mc_power:.2f}%, analytical={exact_power:.2f}% ± {margin:.2f}%"
)
@pytest.mark.parametrize(
@@ -71,7 +70,7 @@ def test_single_predictor_t_test(self, backend, beta, n):
(0.2, 0.2, 200),
],
)
- def test_two_predictors_uncorrelated(self, backend, b1, b2, n):
+ def test_two_predictors_uncorrelated(self, b1, b2, n):
"""Each t-test and F-test with Σ = I."""
from mcpower import MCPower
@@ -91,12 +90,12 @@ def test_two_predictors_uncorrelated(self, backend, b1, b2, n):
mc_power = get_power(result, var)
exact = analytical_t_power(beta, n, p=2, sigma_eps=1.0, vif_j=1.0)
margin = mc_accuracy_margin(exact, N_SIMS)
- assert abs(mc_power - exact) < margin, f"[{backend}] {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
+ assert abs(mc_power - exact) < margin, f"{var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
mc_f = get_power(result, "overall")
exact_f = analytical_f_power([b1, b2], n, Sigma, sigma_eps=1.0)
margin_f = mc_accuracy_margin(exact_f, N_SIMS)
- assert abs(mc_f - exact_f) < margin_f, f"[{backend}] F-test: MC={mc_f:.2f}%, analytical={exact_f:.2f}% ± {margin_f:.2f}%"
+ assert abs(mc_f - exact_f) < margin_f, f"F-test: MC={mc_f:.2f}%, analytical={exact_f:.2f}% ± {margin_f:.2f}%"
@pytest.mark.parametrize(
"b1,b2,rho,n",
@@ -107,7 +106,7 @@ def test_two_predictors_uncorrelated(self, backend, b1, b2, n):
(0.5, 0.3, 0.5, 80),
],
)
- def test_two_predictors_correlated_t_tests(self, backend, b1, b2, rho, n):
+ def test_two_predictors_correlated_t_tests(self, b1, b2, rho, n):
"""Individual t-tests with correlated predictors: VIF matters."""
from mcpower import MCPower
@@ -132,5 +131,5 @@ def test_two_predictors_correlated_t_tests(self, backend, b1, b2, rho, n):
exact = analytical_t_power(beta, n, p=2, sigma_eps=1.0, vif_j=vif)
margin = mc_accuracy_margin(exact, N_SIMS)
assert abs(mc_power - exact) < margin, (
- f"[{backend}] rho={rho}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
+ f"rho={rho}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
)
diff --git a/tests/specs/test_type1_error.py b/tests/specs/test_type1_error.py
index c56b4fe..565d80e 100644
--- a/tests/specs/test_type1_error.py
+++ b/tests/specs/test_type1_error.py
@@ -1,8 +1,7 @@
"""
-Type I error control tests — backend-agnostic.
+Type I error control tests.
Under H0 (effect = 0), rejection rate must equal alpha.
-Tests run on ALL available backends via the backend fixture.
"""
import contextlib
@@ -10,7 +9,7 @@
import pytest
-from tests.config import N_SIMS, SEED
+from tests.config import N_SIMS_STANDARD as N_SIMS, SEED
from tests.helpers.mc_margins import mc_margin
from tests.helpers.power_helpers import get_power, make_null_model
@@ -25,7 +24,7 @@ def _quiet():
class TestTypeIErrorControl:
"""Under H0 (effect = 0), rejection rate must equal alpha."""
- def test_single_predictor_null_overall(self, backend):
+ def test_single_predictor_null_overall(self):
"""F-test rejection rate ≈ alpha with one predictor at zero effect."""
m = make_null_model("y = x1", n_sims=N_SIMS, seed=SEED)
result = m.find_power(
@@ -37,9 +36,9 @@ def test_single_predictor_null_overall(self, backend):
power = get_power(result, "overall")
margin = mc_margin(m.alpha, m.n_simulations)
expected = m.alpha * 100
- assert abs(power - expected) < margin, f"[{backend}] F-test power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
+ assert abs(power - expected) < margin, f"F-test power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
- def test_single_predictor_null_individual(self, backend):
+ def test_single_predictor_null_individual(self):
"""t-test rejection rate ≈ alpha for a single zero-effect predictor."""
m = make_null_model("y = x1", n_sims=N_SIMS, seed=SEED)
result = m.find_power(
@@ -51,9 +50,9 @@ def test_single_predictor_null_individual(self, backend):
power = get_power(result, "x1")
margin = mc_margin(m.alpha, m.n_simulations)
expected = m.alpha * 100
- assert abs(power - expected) < margin, f"[{backend}] t-test power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
+ assert abs(power - expected) < margin, f"t-test power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
- def test_two_predictors_null_each(self, backend):
+ def test_two_predictors_null_each(self):
"""Both predictors at zero → each t-test rejects at ~alpha."""
m = make_null_model("y = x1 + x2", n_sims=N_SIMS, seed=SEED)
result = m.find_power(
@@ -66,9 +65,9 @@ def test_two_predictors_null_each(self, backend):
expected = m.alpha * 100
for var in ["x1", "x2"]:
power = get_power(result, var)
- assert abs(power - expected) < margin, f"[{backend}] {var} power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
+ assert abs(power - expected) < margin, f"{var} power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
- def test_large_sample_null(self, backend):
+ def test_large_sample_null(self):
"""
Large N with zero effect must NOT inflate Type I error.
@@ -76,7 +75,7 @@ def test_large_sample_null(self, backend):
"""
m = make_null_model("y = x1", n_sims=N_SIMS, seed=SEED)
result = m.find_power(
- sample_size=1000,
+ sample_size=500,
target_test="x1",
print_results=False,
return_results=True,
@@ -85,7 +84,7 @@ def test_large_sample_null(self, backend):
margin = mc_margin(m.alpha, m.n_simulations)
expected = m.alpha * 100
assert abs(power - expected) < margin, (
- f"[{backend}] Large-N null power: {power:.2f}%, expected {expected}% ± {margin:.2f}% (Type I error inflated with N?)"
+ f"Large-N null power: {power:.2f}%, expected {expected}% ± {margin:.2f}% (Type I error inflated with N?)"
)
@@ -93,7 +92,7 @@ class TestAlphaCalibration:
"""Rejection rate tracks the nominal alpha across levels."""
@pytest.mark.parametrize("alpha", [0.01, 0.05, 0.10])
- def test_null_rejection_matches_alpha(self, backend, alpha):
+ def test_null_rejection_matches_alpha(self, alpha):
m = make_null_model("y = x1", n_sims=N_SIMS, alpha=alpha, seed=SEED)
result = m.find_power(
sample_size=100,
@@ -104,4 +103,4 @@ def test_null_rejection_matches_alpha(self, backend, alpha):
power = get_power(result, "x1")
margin = mc_margin(alpha, m.n_simulations)
expected = alpha * 100
- assert abs(power - expected) < margin, f"[{backend}] alpha={alpha}: observed {power:.2f}%, expected {expected}% ± {margin:.2f}%"
+ assert abs(power - expected) < margin, f"alpha={alpha}: observed {power:.2f}%, expected {expected}% ± {margin:.2f}%"
diff --git a/tests/unit/test_distributions.py b/tests/unit/test_distributions.py
index d3e77f9..693d1ff 100644
--- a/tests/unit/test_distributions.py
+++ b/tests/unit/test_distributions.py
@@ -24,7 +24,6 @@
import pytest
from mcpower.stats.distributions import (
- _BACKEND,
chi2_cdf,
chi2_ppf,
compute_critical_values_lme,
@@ -587,29 +586,6 @@ def test_studentized_range_k_too_large_returns_inf(self):
# ===========================================================================
# 15. Backend detection
-# ===========================================================================
-class TestBackendDetection:
- """Verify the distribution backend is correctly detected."""
-
- def test_backend_is_set(self):
- assert _BACKEND is not None
-
- def test_backend_is_string(self):
- assert isinstance(_BACKEND, str)
-
- def test_backend_is_known_value(self):
- assert _BACKEND in ("native", "scipy")
-
- def test_native_backend_when_compiled(self):
- """When the C++ extension is compiled, backend should be 'native'."""
- try:
- import mcpower.backends.mcpower_native # noqa: F401
-
- assert _BACKEND == "native"
- except ImportError:
- pytest.skip("C++ native backend not compiled")
-
-
# ===========================================================================
# Cross-consistency checks
# ===========================================================================
diff --git a/tests/unit/test_distributions_coverage.py b/tests/unit/test_distributions_coverage.py
new file mode 100644
index 0000000..224c8e6
--- /dev/null
+++ b/tests/unit/test_distributions_coverage.py
@@ -0,0 +1,42 @@
+"""Tests for distributions.py — optimizer functions and edge cases."""
+
+import numpy as np
+import pytest
+
+from mcpower.stats.distributions import minimize_lbfgsb, minimize_scalar_brent
+
+
+class TestOptimizerLBFGSB:
+ """L-BFGS-B optimizer via native backend."""
+
+ def test_finds_correct_minimum(self):
+ # Simple quadratic: f(x) = (x-2)^2
+ result = minimize_lbfgsb(
+ lambda x: float((x[0] - 2) ** 2),
+ x0=np.array([0.0]),
+ bounds=[(-10.0, 10.0)],
+ )
+ assert abs(result.x[0] - 2.0) < 0.01
+ assert result.fun < 0.01
+
+
+class TestOptimizerBrent:
+ """Brent scalar minimizer via native backend."""
+
+ def test_finds_correct_minimum(self):
+ # f(x) = (x - 3)^2
+ result = minimize_scalar_brent(
+ lambda x: (x - 3) ** 2,
+ bounds=(0.0, 10.0),
+ )
+ assert abs(result.x - 3.0) < 0.01
+ assert result.fun < 0.01
+
+ def test_converged_flag(self):
+ result = minimize_scalar_brent(
+ lambda x: (x - 5) ** 2,
+ bounds=(0.0, 10.0),
+ )
+ assert result.converged
+
+
diff --git a/tests/unit/test_formatters_edge.py b/tests/unit/test_formatters_edge.py
new file mode 100644
index 0000000..f52fd25
--- /dev/null
+++ b/tests/unit/test_formatters_edge.py
@@ -0,0 +1,230 @@
+"""Tests for formatter edge cases — scenario sample-size long format, cumulative recs, NaN filtering."""
+
+import math
+
+import pytest
+
+from mcpower.utils.formatters import _ResultFormatter, _is_nan
+
+
+_fmt = _ResultFormatter()
+
+
+def _make_scenario_sample_size_data(
+ target_tests=("x1", "x2"),
+ correction=None,
+ sample_sizes=(50, 100, 150),
+ optimistic_achieved=None,
+ realistic_achieved=None,
+ doomer_achieved=None,
+):
+ """Build a scenario sample_size result dict for formatting tests."""
+ if optimistic_achieved is None:
+ optimistic_achieved = {"x1": 50, "x2": 100}
+ if realistic_achieved is None:
+ realistic_achieved = {"x1": 100, "x2": 150}
+ if doomer_achieved is None:
+ doomer_achieved = {"x1": 0, "x2": 0} # Not achieved
+
+ def _make_scenario(achieved):
+ achieved_corr = {t: -1 for t in target_tests} if not correction else achieved
+ return {
+ "model": {
+ "target_tests": list(target_tests),
+ "correction": correction,
+ "sample_size_range": {"from_size": sample_sizes[0], "to_size": sample_sizes[-1]},
+ "target_power": 80.0,
+ },
+ "results": {
+ "first_achieved": achieved,
+ "first_achieved_corrected": achieved_corr,
+ "sample_sizes_tested": list(sample_sizes),
+ "powers_by_test": {
+ t: [30.0 + 25.0 * i for i in range(len(sample_sizes))]
+ for t in target_tests
+ },
+ "powers_by_test_corrected": (
+ {t: [25.0 + 25.0 * i for i in range(len(sample_sizes))] for t in target_tests}
+ if correction
+ else None
+ ),
+ },
+ }
+
+ return {
+ "analysis_type": "sample_size",
+ "scenarios": {
+ "optimistic": _make_scenario(optimistic_achieved),
+ "realistic": _make_scenario(realistic_achieved),
+ "doomer": _make_scenario(doomer_achieved),
+ },
+ "comparison": {},
+ }
+
+
+class TestScenarioSampleSizeLongFormat:
+ """Test _format_scenario_sample_size with summary='long'."""
+
+ def test_recommendations_present(self):
+ data = _make_scenario_sample_size_data()
+ output = _fmt.format("scenario_sample_size", data, "long")
+ assert "RECOMMENDATIONS" in output
+
+ def test_unachievable_tests_warning(self):
+ data = _make_scenario_sample_size_data(
+ doomer_achieved={"x1": 0, "x2": 0},
+ )
+ output = _fmt.format("scenario_sample_size", data, "long")
+ assert "Warning" in output or "may not achieve" in output
+
+ def test_realistic_recommendation_shown(self):
+ data = _make_scenario_sample_size_data(
+ realistic_achieved={"x1": 100, "x2": 150},
+ )
+ output = _fmt.format("scenario_sample_size", data, "long")
+ assert "150" in output # max N for realistic
+
+ def test_short_format_produces_table(self):
+ data = _make_scenario_sample_size_data()
+ output = _fmt.format("scenario_sample_size", data, "short")
+ assert "SCENARIO SUMMARY" in output
+
+ def test_with_correction(self):
+ data = _make_scenario_sample_size_data(correction="bonferroni")
+ output = _fmt.format("scenario_sample_size", data, "short")
+ assert "Opt(U)" in output or "Uncorrected" in output.lower() or "(U)" in output
+
+
+class TestCumulativeRecommendations:
+ """Test _format_cumulative_recommendations paths."""
+
+ def test_non_scenario_target_met(self):
+ data = {
+ "model": {
+ "target_tests": ["x1", "x2"],
+ "target_power": 80.0,
+ },
+ "results": {
+ "sample_sizes_tested": [50, 100, 150],
+ "powers_by_test": {
+ "x1": [60.0, 85.0, 95.0],
+ "x2": [70.0, 90.0, 98.0],
+ },
+ },
+ }
+ lines = _fmt._format_cumulative_recommendations(data, is_scenario=False)
+ joined = "\n".join(lines)
+ assert "N=" in joined # Found a sample size
+
+ def test_non_scenario_target_not_met(self):
+ data = {
+ "model": {
+ "target_tests": ["x1", "x2"],
+ "target_power": 80.0,
+ },
+ "results": {
+ "sample_sizes_tested": [50, 100],
+ "powers_by_test": {
+ "x1": [10.0, 20.0],
+ "x2": [15.0, 25.0],
+ },
+ },
+ }
+ lines = _fmt._format_cumulative_recommendations(data, is_scenario=False)
+ joined = "\n".join(lines)
+ assert ">100" in joined # Exceeded max tested
+
+ def test_scenario_recommendations(self):
+ data = _make_scenario_sample_size_data(
+ sample_sizes=(50, 100, 150, 200),
+ optimistic_achieved={"x1": 100, "x2": 150},
+ )
+ # Override powers so all > 80%
+ for scenario in data["scenarios"].values():
+ scenario["results"]["powers_by_test"] = {
+ "x1": [50.0, 85.0, 92.0, 98.0],
+ "x2": [40.0, 75.0, 88.0, 95.0],
+ }
+ lines = _fmt._format_cumulative_recommendations(data, is_scenario=True)
+ assert len(lines) > 0
+
+ def test_empty_scenarios(self):
+ data = {"scenarios": {}}
+ lines = _fmt._format_cumulative_recommendations(data, is_scenario=True)
+ assert lines == []
+
+ def test_no_results_key(self):
+ data = {}
+ lines = _fmt._format_cumulative_recommendations(data, is_scenario=False)
+ assert lines == []
+
+
+class TestNaNPowerFiltering:
+ """NaN power values in cumulative table should be filtered out."""
+
+ def test_nan_power_filtered_in_cumulative_sample_size_table(self):
+ lines = []
+ _fmt._add_cumulative_sample_size_table(
+ lines,
+ sample_sizes=[50, 100],
+ target_tests=["x1", "x2_nan"],
+ powers_by_test={
+ "x1": [50.0, 80.0],
+ "x2_nan": [float("nan"), float("nan")],
+ },
+ )
+ # Should still produce output for x1 (x2_nan filtered out)
+ output = "\n".join(lines)
+ assert "N=50" in output or "50" in output
+
+ def test_all_nan_produces_no_table(self):
+ lines = []
+ _fmt._add_cumulative_sample_size_table(
+ lines,
+ sample_sizes=[50],
+ target_tests=["x1"],
+ powers_by_test={"x1": [float("nan")]},
+ )
+ # All NaN → no valid tests → no table
+ assert len(lines) == 0
+
+
+class TestIsNan:
+ """Test _is_nan utility."""
+
+ def test_nan_float(self):
+ assert _is_nan(float("nan"))
+
+ def test_regular_float(self):
+ assert not _is_nan(42.0)
+
+ def test_non_float(self):
+ assert not _is_nan("nan")
+ assert not _is_nan(None)
+ assert not _is_nan(42)
+
+
+class TestExtractScenarioMeta:
+ """Test _extract_scenario_meta."""
+
+ def test_no_model_returns_none(self):
+ target_tests, correction = _fmt._extract_scenario_meta({"opt": {"results": {}}})
+ assert target_tests is None
+
+ def test_extracts_from_first_scenario(self):
+ scenarios = {
+ "optimistic": {
+ "model": {"target_tests": ["a", "b"], "correction": "holm"},
+ }
+ }
+ target_tests, correction = _fmt._extract_scenario_meta(scenarios)
+ assert target_tests == ["a", "b"]
+ assert correction == "holm"
+
+
+class TestFormatUnknownType:
+ """Unknown result type should raise."""
+
+ def test_unknown_result_type(self):
+ with pytest.raises(ValueError, match="Unknown result type"):
+ _fmt.format("nonexistent", {})
diff --git a/tests/unit/test_mixed_models_coverage.py b/tests/unit/test_mixed_models_coverage.py
new file mode 100644
index 0000000..bc99974
--- /dev/null
+++ b/tests/unit/test_mixed_models_coverage.py
@@ -0,0 +1,292 @@
+"""Tests for stats/mixed_models.py — statsmodels convergence, corrections, native wrappers.
+
+Uses pytest.mark.lme to skip when statsmodels is not installed.
+"""
+
+import warnings
+from unittest.mock import MagicMock, patch
+
+import numpy as np
+import pytest
+
+from mcpower.stats.mixed_models import (
+ _ensure_lme_crits,
+ _lme_analysis_wrapper,
+ _wrap_native_result,
+ reset_warm_start_cache,
+)
+
+pytestmark = pytest.mark.lme
+
+
+class TestWrapNativeResult:
+ """Test _wrap_native_result helper."""
+
+ def test_non_empty_non_verbose(self):
+ result = np.array([1.0, 0.0, 1.0])
+ wrapped = _wrap_native_result(result, verbose=False, solver_name="native_q1")
+ np.testing.assert_array_equal(wrapped, result)
+
+ def test_non_empty_verbose(self):
+ result = np.array([1.0, 0.0, 1.0])
+ wrapped = _wrap_native_result(result, verbose=True, solver_name="native_q1")
+ assert isinstance(wrapped, dict)
+ assert "results" in wrapped
+ assert "diagnostics" in wrapped
+ assert wrapped["diagnostics"]["solver"] == "native_q1"
+
+ def test_non_empty_verbose_with_extra_diag(self):
+ result = np.array([1.0])
+ wrapped = _wrap_native_result(
+ result, verbose=True, solver_name="native_general",
+ extra_diag={"q": 3},
+ )
+ assert wrapped["diagnostics"]["q"] == 3
+
+ def test_empty_non_verbose_returns_none(self):
+ result = np.array([])
+ assert _wrap_native_result(result, verbose=False, solver_name="native_q1") is None
+
+ def test_empty_verbose_returns_failure_dict(self):
+ result = np.array([])
+ wrapped = _wrap_native_result(result, verbose=True, solver_name="native_q1")
+ assert wrapped["results"] is None
+ assert "failure_reason" in wrapped
+ assert "empty result" in wrapped["failure_reason"]
+
+
+class TestEnsureLMECrits:
+ """Test _ensure_lme_crits computes when None."""
+
+ def test_computes_when_none(self):
+ chi2, z, crits = _ensure_lme_crits(
+ alpha=0.05, p=3, n_targets=2, correction_method=0,
+ chi2_crit=None, z_crit=None, correction_z_crits=None,
+ )
+ assert np.isfinite(chi2)
+ assert np.isfinite(z)
+ assert len(crits) == 2
+
+ def test_passthrough_when_provided(self):
+ chi2, z, crits = _ensure_lme_crits(
+ alpha=0.05, p=3, n_targets=2, correction_method=0,
+ chi2_crit=7.8, z_crit=1.96, correction_z_crits=np.array([1.96, 1.96]),
+ )
+ assert chi2 == 7.8
+ assert z == 1.96
+ assert len(crits) == 2
+
+
+class TestLMEAnalysisWrapperRouting:
+ """Test _lme_analysis_wrapper routes to correct backend."""
+
+ def test_unknown_backend_raises(self):
+ with pytest.raises(ValueError, match="Unknown backend"):
+ _lme_analysis_wrapper(
+ np.eye(10), np.ones(10), np.array([0, 1]),
+ np.zeros(10, dtype=np.int32),
+ correction_method=0, alpha=0.05, backend="nonexistent",
+ )
+
+
+class TestStatsmodelsConvergence:
+ """Test statsmodels fallback path with mocked MixedLM."""
+
+ def _make_mock_result(self, converged=True, params=None, pvalues=None, n_params=3):
+ """Create a mock MixedLM result."""
+ result = MagicMock()
+ result.converged = converged
+ result.params = params if params is not None else np.array([1.0, 0.5, 0.3])
+ result.pvalues = pvalues if pvalues is not None else np.array([0.01, 0.02, 0.04])
+ result.fe_params = result.params
+ result.bse = np.array([0.1, 0.1, 0.1])
+
+ # cov_re: random effects variance (needs .iloc[0, 0])
+ cov_re = MagicMock()
+ cov_re.iloc.__getitem__ = MagicMock(return_value=0.5)
+ result.cov_re = cov_re
+
+ result.scale = 1.0
+ result.llf = -50.0
+
+ # Make cov_params return a proper matrix
+ result.cov_params.return_value = np.eye(n_params) * 0.01
+
+ # model attribute
+ result.model = MagicMock()
+ result.model.exog = MagicMock()
+ result.model.exog.shape = (100, n_params)
+
+ return result
+
+ @patch("statsmodels.regression.mixed_linear_model.MixedLM")
+ def test_warm_start_retry_chain(self, mock_mixedlm_cls):
+ """First fit fails, cold start succeeds."""
+ from mcpower.stats.mixed_models import _lme_analysis_statsmodels, _lme_thread_local
+
+ _lme_thread_local.warm_start_params = np.array([1.0, 0.5, 0.3])
+
+ mock_model = MagicMock()
+ mock_mixedlm_cls.return_value = mock_model
+
+ good_result = self._make_mock_result()
+ mock_model.fit.side_effect = [
+ Exception("warm start diverged"),
+ good_result,
+ ]
+ mock_model.loglike.return_value = -50.0
+
+ result = _lme_analysis_statsmodels(
+ X_expanded=np.random.randn(100, 2),
+ y=np.random.randn(100),
+ target_indices=np.array([0, 1]),
+ cluster_ids=np.repeat(np.arange(10), 10),
+
+ correction_method=0,
+ alpha=0.05,
+ )
+ assert result is not None
+
+ @patch("statsmodels.regression.mixed_linear_model.MixedLM")
+ def test_all_attempts_fail_returns_none(self, mock_mixedlm_cls):
+ from mcpower.stats.mixed_models import _lme_analysis_statsmodels, _lme_thread_local
+
+ _lme_thread_local.warm_start_params = None
+
+ mock_model = MagicMock()
+ mock_mixedlm_cls.return_value = mock_model
+ mock_model.fit.side_effect = Exception("always fails")
+
+ result = _lme_analysis_statsmodels(
+ X_expanded=np.random.randn(100, 2),
+ y=np.random.randn(100),
+ target_indices=np.array([0, 1]),
+ cluster_ids=np.repeat(np.arange(10), 10),
+
+ correction_method=0,
+ alpha=0.05,
+ )
+ assert result is None
+
+ @patch("statsmodels.regression.mixed_linear_model.MixedLM")
+ def test_all_attempts_fail_verbose_returns_dict(self, mock_mixedlm_cls):
+ from mcpower.stats.mixed_models import _lme_analysis_statsmodels, _lme_thread_local
+
+ _lme_thread_local.warm_start_params = None
+
+ mock_model = MagicMock()
+ mock_mixedlm_cls.return_value = mock_model
+ mock_model.fit.side_effect = Exception("always fails")
+
+ result = _lme_analysis_statsmodels(
+ X_expanded=np.random.randn(100, 2),
+ y=np.random.randn(100),
+ target_indices=np.array([0, 1]),
+ cluster_ids=np.repeat(np.arange(10), 10),
+
+ correction_method=0,
+ alpha=0.05,
+ verbose=True,
+ )
+ assert isinstance(result, dict)
+ assert result["results"] is None
+ assert "failure_reason" in result
+
+ @patch("statsmodels.regression.mixed_linear_model.MixedLM")
+ def test_not_converged_returns_none(self, mock_mixedlm_cls):
+ """When result.converged is False for all attempts."""
+ from mcpower.stats.mixed_models import _lme_analysis_statsmodels, _lme_thread_local
+
+ _lme_thread_local.warm_start_params = None
+
+ mock_model = MagicMock()
+ mock_mixedlm_cls.return_value = mock_model
+
+ bad_result = self._make_mock_result(converged=False)
+ mock_model.fit.return_value = bad_result
+
+ result = _lme_analysis_statsmodels(
+ X_expanded=np.random.randn(100, 2),
+ y=np.random.randn(100),
+ target_indices=np.array([0, 1]),
+ cluster_ids=np.repeat(np.arange(10), 10),
+
+ correction_method=0,
+ alpha=0.05,
+ )
+ assert result is None
+
+
+class TestCorrections:
+ """Test statsmodels FDR, Holm, Bonferroni, no-correction paths."""
+
+ def _make_mock_result(self):
+ result = MagicMock()
+ result.converged = True
+ result.params = np.array([1.0, 0.5, 0.3])
+ result.pvalues = np.array([0.001, 0.02, 0.04])
+ result.fe_params = result.params
+ result.bse = np.array([0.1, 0.1, 0.1])
+ result.scale = 1.0
+ result.llf = -50.0
+ result.model = MagicMock()
+ result.model.exog = MagicMock()
+ result.model.exog.shape = (100, 3)
+
+ cov_re_mock = MagicMock()
+ cov_re_mock.iloc.__getitem__ = MagicMock(return_value=0.5)
+ result.cov_re = cov_re_mock
+ result.cov_params.return_value = np.eye(3) * 0.01
+
+ return result
+
+ def _run_with_correction(self, correction_method):
+ from mcpower.stats.mixed_models import _lme_analysis_statsmodels, _lme_thread_local
+
+ _lme_thread_local.warm_start_params = None
+
+ mock_result = self._make_mock_result()
+
+ with patch("statsmodels.regression.mixed_linear_model.MixedLM") as mock_cls:
+ mock_model = MagicMock()
+ mock_cls.return_value = mock_model
+ mock_model.fit.return_value = mock_result
+ mock_model.loglike.return_value = -50.0
+
+ out = _lme_analysis_statsmodels(
+ X_expanded=np.random.randn(100, 2),
+ y=np.random.randn(100),
+ target_indices=np.array([0, 1]),
+ cluster_ids=np.repeat(np.arange(10), 10),
+
+ correction_method=correction_method,
+ alpha=0.05,
+ )
+ return out
+
+ def test_no_correction(self):
+ result = self._run_with_correction(0)
+ assert result is not None
+
+ def test_bonferroni(self):
+ result = self._run_with_correction(1)
+ assert result is not None
+
+ def test_fdr(self):
+ result = self._run_with_correction(2)
+ assert result is not None
+
+ def test_holm(self):
+ result = self._run_with_correction(3)
+ assert result is not None
+
+
+class TestResetWarmStartCache:
+ """Test reset_warm_start_cache."""
+
+ def test_clears_params(self):
+ from mcpower.stats.mixed_models import _lme_thread_local
+
+ _lme_thread_local.warm_start_params = np.array([1.0])
+ reset_warm_start_cache()
+ assert _lme_thread_local.warm_start_params is None
diff --git a/tests/unit/test_model_coverage.py b/tests/unit/test_model_coverage.py
new file mode 100644
index 0000000..23d47c9
--- /dev/null
+++ b/tests/unit/test_model_coverage.py
@@ -0,0 +1,117 @@
+"""Tests for model.py — parallel fallback, Tukey validation, NaN under Tukey correction."""
+
+import warnings
+from unittest.mock import MagicMock, patch
+
+import numpy as np
+import pytest
+
+from mcpower import MCPower
+
+
+class TestTukeyWithoutPosthoc:
+ """Tukey correction without posthoc specs should raise ValueError."""
+
+ def test_tukey_without_posthoc_raises(self):
+ model = MCPower("y = x1 + x2")
+ model.set_effects("x1=0.5, x2=0.3")
+
+ with pytest.raises(ValueError, match="Tukey correction requires"):
+ model.find_power(
+ sample_size=100,
+ correction="tukey",
+ print_results=False,
+ )
+
+
+class TestTukeyNaNification:
+ """Non-posthoc tests should be NaN-ified under Tukey correction."""
+
+ def test_non_posthoc_tests_nan_under_tukey(self):
+ model = MCPower("y = group + x1")
+ model.set_variable_type("group=(factor,3)")
+ model.set_effects("group[2]=0.5, group[3]=0.4, x1=0.3")
+ model.n_simulations = 50
+ model.seed = 42
+
+ result = model.find_sample_size(
+ target_test="all, all-posthoc",
+ correction="tukey",
+ from_size=30,
+ to_size=60,
+ by=30,
+ print_results=False,
+ return_results=True,
+ )
+
+ assert result is not None
+ results = result["results"]
+ corrected = results.get("powers_by_test_corrected", {})
+
+ # Post-hoc comparisons should have real power values
+ # Non-posthoc tests (like "x1", "group[2]", "group[3]", "overall")
+ # should have NaN values
+ posthoc_labels = {s.label for s in model._posthoc_specs}
+ for test_name, powers in corrected.items():
+ if test_name not in posthoc_labels:
+ assert all(isinstance(v, float) and np.isnan(v) for v in powers), \
+ f"Expected NaN for non-posthoc test '{test_name}', got {powers}"
+
+ # first_achieved_corrected for non-posthoc should be -1
+ for test_name, n in results.get("first_achieved_corrected", {}).items():
+ if test_name not in posthoc_labels:
+ assert n == -1, f"Expected -1 for '{test_name}', got {n}"
+
+
+class TestParallelFallback:
+ """Parallel execution falls back to sequential on exception."""
+
+ def test_parallel_exception_falls_back(self, capsys):
+ model = MCPower("y = x1 + x2")
+ model.set_effects("x1=0.5, x2=0.3")
+ model.parallel = True
+ model.n_simulations = 50
+ model.seed = 42
+
+ # Parallel is imported inside the function via `from joblib import Parallel`,
+ # so we patch it at the joblib module level.
+ with patch("joblib.Parallel", side_effect=RuntimeError("joblib broken")):
+ # Should still complete via sequential fallback
+ result = model.find_sample_size(
+ from_size=30,
+ to_size=60,
+ by=30,
+ print_results=False,
+ return_results=True,
+ )
+ assert result is not None
+ captured = capsys.readouterr()
+ assert "Falling back to sequential" in captured.out
+
+
+class TestIsParallelEffective:
+ """Test _is_parallel_effective resolution."""
+
+ def test_true_always_parallel(self):
+ model = MCPower("y = x1 + x2")
+ model.parallel = True
+ assert model._is_parallel_effective() is True
+
+ def test_false_never_parallel(self):
+ model = MCPower("y = x1 + x2")
+ model.parallel = False
+ assert model._is_parallel_effective() is False
+
+ def test_mixedmodels_with_clusters(self):
+ model = MCPower("y ~ x1 + (1|school)")
+ model.set_cluster("school", ICC=0.2, n_clusters=20)
+ model.set_effects("x1=0.5")
+ model._apply() # cluster_specs are deferred until apply()
+ model.parallel = "mixedmodels"
+ assert model._is_parallel_effective() is True
+
+ def test_mixedmodels_without_clusters(self):
+ model = MCPower("y = x1 + x2")
+ model.set_effects("x1=0.5, x2=0.3")
+ model.parallel = "mixedmodels"
+ assert model._is_parallel_effective() is False
diff --git a/tests/unit/test_native_backend.py b/tests/unit/test_native_backend.py
new file mode 100644
index 0000000..93b4a3e
--- /dev/null
+++ b/tests/unit/test_native_backend.py
@@ -0,0 +1,60 @@
+"""Tests for mcpower.backends.native — import fallback and _prep utility."""
+
+import numpy as np
+import pytest
+from unittest.mock import patch, MagicMock
+
+from mcpower.backends.native import _prep
+
+
+class TestPrep:
+ """Test _prep array coercion for C++ interop."""
+
+ def test_contiguous_passthrough(self):
+ arr = np.array([1.0, 2.0, 3.0], dtype=np.float64)
+ result = _prep(arr)
+ assert result.flags["C_CONTIGUOUS"]
+ assert result.dtype == np.float64
+
+ def test_non_contiguous_becomes_contiguous(self):
+ arr = np.array([[1.0, 2.0], [3.0, 4.0]], dtype=np.float64)
+ col = arr[:, 1] # non-contiguous column slice
+ assert not col.flags["C_CONTIGUOUS"]
+ result = _prep(col)
+ assert result.flags["C_CONTIGUOUS"]
+ np.testing.assert_array_equal(result, [2.0, 4.0])
+
+ def test_dtype_conversion_float32_to_float64(self):
+ arr = np.array([1.0, 2.0], dtype=np.float32)
+ result = _prep(arr, np.float64)
+ assert result.dtype == np.float64
+
+ def test_dtype_conversion_int64_to_int32(self):
+ arr = np.array([0, 1, 2], dtype=np.int64)
+ result = _prep(arr, np.int32)
+ assert result.dtype == np.int32
+ np.testing.assert_array_equal(result, [0, 1, 2])
+
+ def test_2d_array(self):
+ arr = np.array([[1.0, 2.0], [3.0, 4.0]], dtype=np.float64, order="F")
+ assert not arr.flags["C_CONTIGUOUS"]
+ result = _prep(arr)
+ assert result.flags["C_CONTIGUOUS"]
+ assert result.dtype == np.float64
+
+
+class TestNativeBackendImport:
+ """Test NativeBackend init when C++ extension is unavailable."""
+
+ def test_init_raises_when_unavailable(self):
+ """NativeBackend() should raise ImportError when _NATIVE_AVAILABLE=False."""
+ with patch("mcpower.backends.native._NATIVE_AVAILABLE", False):
+ from mcpower.backends.native import NativeBackend
+ with pytest.raises(ImportError, match="Native C\\+\\+ backend not available"):
+ NativeBackend()
+
+ def test_is_native_available_reflects_module_state(self):
+ from mcpower.backends.native import is_native_available
+ # Just verify it returns a bool
+ result = is_native_available()
+ assert isinstance(result, bool)
diff --git a/tests/unit/test_ols_corrections.py b/tests/unit/test_ols_corrections.py
new file mode 100644
index 0000000..8c6728a
--- /dev/null
+++ b/tests/unit/test_ols_corrections.py
@@ -0,0 +1,251 @@
+"""Tests for OLS post-hoc contrast corrections and edge cases."""
+
+from dataclasses import dataclass
+from typing import Optional
+
+import numpy as np
+import pytest
+
+from mcpower.stats.ols import compute_posthoc_contrasts
+
+
+@dataclass
+class _PostHocSpec:
+ """Minimal PostHocSpec stub for tests."""
+ factor_name: str
+ col_idx_a: Optional[int]
+ col_idx_b: Optional[int]
+ label: str = ""
+ level_a: str = ""
+ level_b: str = ""
+ n_levels: int = 3
+
+
+def _make_ols_data(n=100, p=3, seed=42):
+ """Generate simple OLS data: X, y, and target_indices."""
+ rng = np.random.RandomState(seed)
+ X = rng.randn(n, p)
+ beta = np.array([0.5, 0.3, -0.2])[:p]
+ y = X @ beta + rng.randn(n)
+ return X, y
+
+
+class TestDegenerateDesign:
+ """When dof <= 0, posthoc should return zeros."""
+
+ def test_dof_zero_returns_zeros(self):
+ # n = p+1 → dof = 0
+ n, p = 4, 3
+ rng = np.random.RandomState(42)
+ X = rng.randn(n, p)
+ y = rng.randn(n)
+ specs = [_PostHocSpec("grp", 0, 1)]
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {}, target_indices=np.array([0, 1, 2]),
+ )
+ assert uncorr.shape == (1,)
+ assert not uncorr[0]
+ assert not corr[0]
+ assert override is None
+
+ def test_singular_contrast_variance_stays_zero(self):
+ """When both col_idx_a and col_idx_b are None, t_abs stays 0."""
+ X, y = _make_ols_data()
+ specs = [_PostHocSpec("grp", None, None)]
+
+ uncorr, corr, _ = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {},
+ )
+ assert not uncorr[0]
+ assert not corr[0]
+
+
+class TestCombinedFDR:
+ """FDR (correction_method=2) step-up across regular+posthoc t-stats."""
+
+ def test_fdr_combined_ranking(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [
+ _PostHocSpec("grp", 0, 1),
+ _PostHocSpec("grp", 0, 2),
+ ]
+ target_indices = np.array([0, 1, 2])
+ # Create combined crits of length n_regular + n_posthoc = 5
+ # Use very lenient crits so everything passes
+ combined_crits = np.full(5, 0.01)
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 0.01, {},
+ target_indices=target_indices,
+ correction_method=2,
+ correction_t_crits_combined=combined_crits,
+ )
+ assert override is not None
+ assert len(override) == 3 # n_regular
+ assert len(corr) == 2 # n_posthoc
+
+ def test_fdr_no_significant(self):
+ """With very strict crits, nothing should be significant."""
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+ # Very strict thresholds
+ combined_crits = np.full(4, 100.0)
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 100.0, {},
+ target_indices=target_indices,
+ correction_method=2,
+ correction_t_crits_combined=combined_crits,
+ )
+ assert not np.any(corr)
+ assert override is not None
+ assert not np.any(override)
+
+
+class TestCombinedHolm:
+ """Holm (correction_method=3) step-down with early termination."""
+
+ def test_holm_combined_ranking(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+ combined_crits = np.full(4, 0.01) # Very lenient
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 0.01, {},
+ target_indices=target_indices,
+ correction_method=3,
+ correction_t_crits_combined=combined_crits,
+ )
+ assert override is not None
+ assert len(override) == 3
+
+ def test_holm_early_termination(self):
+ """If the most significant test doesn't pass, none should."""
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+ combined_crits = np.full(4, 1000.0) # Impossible threshold
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 1000.0, {},
+ target_indices=target_indices,
+ correction_method=3,
+ correction_t_crits_combined=combined_crits,
+ )
+ assert not np.any(corr)
+
+
+class TestFallbackPaths:
+ """Fallback when correction_t_crits_combined is None or wrong length."""
+
+ def test_combined_crits_none_fallback(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {},
+ target_indices=target_indices,
+ correction_method=2,
+ correction_t_crits_combined=None,
+ )
+ # Fallback: corrected = uncorrected copy, no override
+ np.testing.assert_array_equal(corr, uncorr)
+ assert override is None
+
+ def test_combined_crits_wrong_length_fallback(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+ # Wrong length: should be 4 (3 regular + 1 posthoc)
+ wrong_crits = np.full(2, 2.0)
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {},
+ target_indices=target_indices,
+ correction_method=2,
+ correction_t_crits_combined=wrong_crits,
+ )
+ np.testing.assert_array_equal(corr, uncorr)
+ assert override is None
+
+
+class TestTukeyMethod:
+ """Tukey post-hoc method path."""
+
+ def test_tukey_uses_factor_crit(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1, n_levels=3)]
+ tukey_crits = {"grp": 0.01} # Very lenient
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "tukey", 2.0, tukey_crits,
+ )
+ # Tukey correction: uncorrected == corrected
+ np.testing.assert_array_equal(uncorr, corr)
+ assert override is None
+
+ def test_tukey_missing_factor_uses_inf(self):
+ """When factor not in tukey_crits, inf is used → not significant."""
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("missing_factor", 0, 1)]
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "tukey", 2.0, {},
+ )
+ assert not uncorr[0]
+ assert not corr[0]
+
+
+class TestBonferroniPosthoc:
+ """Bonferroni correction for posthoc (correction_method=1)."""
+
+ def test_bonferroni_uses_combined_first_crit(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+ combined_crits = np.full(4, 0.01) # Very lenient
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 0.01, {},
+ target_indices=target_indices,
+ correction_method=1,
+ correction_t_crits_combined=combined_crits,
+ )
+ assert override is None # Bonferroni doesn't produce override
+
+
+class TestEmptySpecs:
+ """Empty posthoc specs return empty arrays."""
+
+ def test_no_specs(self):
+ X, y = _make_ols_data()
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, [], "t-test", 2.0, {},
+ )
+ assert len(uncorr) == 0
+ assert len(corr) == 0
+ assert override is None
+
+
+class TestSingleColumnContrasts:
+ """Contrasts where one side is the reference level (None)."""
+
+ def test_col_idx_a_none(self):
+ X, y = _make_ols_data(n=200)
+ specs = [_PostHocSpec("grp", None, 1)]
+ uncorr, corr, _ = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {},
+ )
+ assert uncorr.shape == (1,)
+
+ def test_col_idx_b_none(self):
+ X, y = _make_ols_data(n=200)
+ specs = [_PostHocSpec("grp", 0, None)]
+ uncorr, corr, _ = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {},
+ )
+ assert uncorr.shape == (1,)
diff --git a/tests/unit/test_parsers_errors.py b/tests/unit/test_parsers_errors.py
new file mode 100644
index 0000000..b3c8c01
--- /dev/null
+++ b/tests/unit/test_parsers_errors.py
@@ -0,0 +1,168 @@
+"""Tests for parser error paths and edge cases."""
+
+import pytest
+
+from mcpower.utils.parsers import _AssignmentParser, _parse_equation
+
+
+_parser = _AssignmentParser()
+
+
+class TestAssignmentParserErrors:
+ """Error paths in _AssignmentParser._parse."""
+
+ def test_missing_equals_sign(self):
+ parsed, errors = _parser._parse("x1 0.5", "effect", ["x1"])
+ assert len(errors) == 1
+ assert "Invalid format" in errors[0]
+
+ def test_unknown_parse_type(self):
+ parsed, errors = _parser._parse("x1=0.5", "unknown_type", ["x1"])
+ assert len(errors) == 1
+ assert "Unknown parse type" in errors[0]
+
+ def test_unavailable_variable(self):
+ parsed, errors = _parser._parse("x_missing=0.5", "effect", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "not found" in errors[0]
+ assert "x_missing" in errors[0]
+
+ def test_invalid_effect_value(self):
+ parsed, errors = _parser._parse("x1=abc", "effect", ["x1"])
+ assert len(errors) == 1
+ assert "Invalid effect size" in errors[0]
+
+ def test_multiple_errors(self):
+ parsed, errors = _parser._parse("x_bad=abc, x_also_bad=xyz", "effect", ["x1"])
+ assert len(errors) == 2
+
+
+class TestCorrelationParserErrors:
+ """Error paths for correlation parsing."""
+
+ def test_invalid_correlation_format(self):
+ parsed, errors = _parser._parse("x1_x2=0.5", "correlation", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "Invalid format" in errors[0] or "Invalid correlation" in errors[0]
+
+ def test_correlation_var_not_found(self):
+ parsed, errors = _parser._parse("corr(x1, x_missing)=0.5", "correlation", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "not found" in errors[0]
+
+ def test_self_correlation(self):
+ parsed, errors = _parser._parse("corr(x1, x1)=0.5", "correlation", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "Cannot correlate variable with itself" in errors[0]
+
+ def test_correlation_value_out_of_range(self):
+ parsed, errors = _parser._parse("corr(x1, x2)=1.5", "correlation", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "between -1 and 1" in errors[0]
+
+ def test_invalid_correlation_value(self):
+ parsed, errors = _parser._parse("corr(x1, x2)=abc", "correlation", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "Invalid correlation value" in errors[0]
+
+
+class TestVariableTypeErrors:
+ """Error paths for variable type parsing."""
+
+ def test_unsupported_type(self):
+ parsed, errors = _parser._parse("x1=crazy_type", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "Unsupported type" in errors[0]
+
+ def test_binary_proportion_out_of_range(self):
+ parsed, errors = _parser._parse("x1=(binary,1.5)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "between 0 and 1" in errors[0]
+
+ def test_binary_non_numeric_proportion(self):
+ parsed, errors = _parser._parse("x1=(binary,abc)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "Invalid proportion" in errors[0]
+
+ def test_binary_wrong_param_count(self):
+ parsed, errors = _parser._parse("x1=(binary,0.3,0.4)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "exactly 2 values" in errors[0]
+
+ def test_factor_less_than_2_levels(self):
+ parsed, errors = _parser._parse("x1=(factor,1)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "at least 2 levels" in errors[0]
+
+ def test_factor_more_than_20_levels(self):
+ parsed, errors = _parser._parse("x1=(factor,21)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "more than 20 levels" in errors[0]
+
+ def test_factor_non_integer_levels(self):
+ parsed, errors = _parser._parse("x1=(factor,abc)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "Must be integer" in errors[0]
+
+ def test_factor_proportions_more_than_20(self):
+ props = ",".join(["0.04"] * 21)
+ parsed, errors = _parser._parse(f"x1=(factor,{props})", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "more than 20 levels" in errors[0]
+
+ def test_factor_zero_proportion(self):
+ parsed, errors = _parser._parse("x1=(factor,0.5,0.0,0.5)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "positive" in errors[0]
+
+ def test_factor_non_numeric_proportions(self):
+ parsed, errors = _parser._parse("x1=(factor,abc,def)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "numeric" in errors[0]
+
+ def test_tuple_no_comma(self):
+ parsed, errors = _parser._parse("x1=(binary)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "Invalid tuple format" in errors[0]
+
+ def test_tuple_unsupported_type_in_tuple(self):
+ parsed, errors = _parser._parse("x1=(normal,0.5)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "only supported for binary and factor" in errors[0]
+
+
+class TestEquationParsing:
+ """Edge cases in _parse_equation."""
+
+ def test_nested_random_effects(self):
+ dep, formula, ranefs = _parse_equation("y ~ x1 + (1|A/B)")
+ assert dep == "y"
+ assert len(ranefs) == 2
+ group_vars = {r["grouping_var"] for r in ranefs}
+ assert "A" in group_vars
+ assert "A:B" in group_vars
+
+ def test_duplicate_grouping_var_raises(self):
+ with pytest.raises(ValueError, match="Duplicate random effect grouping variable"):
+ _parse_equation("y ~ x1 + (1|school) + (1|school)")
+
+ def test_random_slopes(self):
+ dep, formula, ranefs = _parse_equation("y ~ x1 + (1 + x1|school)")
+ assert len(ranefs) == 1
+ assert ranefs[0]["type"] == "random_slope"
+ assert ranefs[0]["slope_vars"] == ["x1"]
+ assert ranefs[0]["grouping_var"] == "school"
+
+ def test_random_slope_duplicate_grouping_raises(self):
+ with pytest.raises(ValueError, match="Duplicate"):
+ _parse_equation("y ~ x1 + (1|school) + (1 + x1|school)")
+
+ def test_no_separator_uses_default_dep(self):
+ dep, formula, ranefs = _parse_equation("x1+x2")
+ assert dep == "explained_variable"
+ assert "x1" in formula
+ assert "x2" in formula
+
+ def test_nested_duplicate_parent_raises(self):
+ with pytest.raises(ValueError, match="Duplicate"):
+ _parse_equation("y ~ (1|A) + (1|A/B)")
diff --git a/tests/unit/test_progress.py b/tests/unit/test_progress.py
index 329acca..768420a 100644
--- a/tests/unit/test_progress.py
+++ b/tests/unit/test_progress.py
@@ -127,12 +127,6 @@ def test_completion_newline(self):
class TestTqdmReporter:
"""Test TqdmReporter with mock tqdm."""
- def test_tqdm_missing_raises(self):
- reporter = TqdmReporter()
- with patch.dict("sys.modules", {"tqdm": None}):
- with pytest.raises(ImportError, match="tqdm"):
- reporter(0, 100)
-
def test_tqdm_basic_flow(self):
mock_bar = MagicMock()
mock_bar.n = 0
@@ -154,6 +148,51 @@ def test_tqdm_basic_flow(self):
reporter(100, 100) # closes
mock_bar.close.assert_called_once()
+ def test_tqdm_successive_sessions(self):
+ """After close, a new session creates a fresh bar."""
+ mock_bar = MagicMock()
+ mock_bar.n = 0
+ mock_tqdm_cls = MagicMock(return_value=mock_bar)
+ mock_tqdm_module = MagicMock()
+ mock_tqdm_module.tqdm = mock_tqdm_cls
+
+ reporter = TqdmReporter()
+
+ with patch.dict("sys.modules", {"tqdm": mock_tqdm_module}):
+ # First session
+ reporter(0, 50)
+ mock_bar.n = 0
+ reporter(50, 50)
+ mock_bar.close.assert_called_once()
+ assert reporter._bar is None
+
+ # Second session — should create a new bar
+ mock_tqdm_cls.reset_mock()
+ mock_bar2 = MagicMock()
+ mock_bar2.n = 0
+ mock_tqdm_cls.return_value = mock_bar2
+
+ reporter(0, 200)
+ assert mock_tqdm_cls.call_count == 1
+ mock_tqdm_cls.assert_called_with(total=200, unit="sim")
+
+ def test_tqdm_no_negative_delta(self):
+ """When current <= bar.n, update should not be called with negative delta."""
+ mock_bar = MagicMock()
+ mock_bar.n = 50
+ mock_tqdm_cls = MagicMock(return_value=mock_bar)
+ mock_tqdm_module = MagicMock()
+ mock_tqdm_module.tqdm = mock_tqdm_cls
+
+ reporter = TqdmReporter()
+
+ with patch.dict("sys.modules", {"tqdm": mock_tqdm_module}):
+ reporter(0, 100) # creates bar
+ mock_bar.n = 50
+ reporter(30, 100) # current < bar.n
+ # update should NOT have been called (delta = 30 - 50 = -20, not > 0)
+ mock_bar.update.assert_not_called()
+
class TestComputeTotalSimulations:
"""Test compute_total_simulations helper."""
diff --git a/tests/unit/test_results.py b/tests/unit/test_results.py
new file mode 100644
index 0000000..31c3082
--- /dev/null
+++ b/tests/unit/test_results.py
@@ -0,0 +1,138 @@
+"""Unit tests for mcpower.core.results — ResultsProcessor and builder functions."""
+
+import numpy as np
+import pytest
+
+from mcpower.core.results import ResultsProcessor, build_power_result, build_sample_size_result
+
+
+class TestCalculatePowers:
+ """Tests for ResultsProcessor.calculate_powers."""
+
+ def test_basic_two_tests(self):
+ """Power calculation with two tests (overall + one predictor)."""
+ proc = ResultsProcessor(target_power=80.0)
+ # 10 simulations, 2 columns: [overall, x1]
+ # overall: 8/10 sig, x1: 6/10 sig
+ results = [np.array([True, True])] * 6 + [
+ np.array([True, False]),
+ np.array([True, False]),
+ np.array([False, False]),
+ np.array([False, False]),
+ ]
+ corrected = results # same for this test
+
+ out = proc.calculate_powers(results, corrected, ["overall", "x1"])
+
+ assert out["individual_powers"]["overall"] == pytest.approx(80.0)
+ assert out["individual_powers"]["x1"] == pytest.approx(60.0)
+ assert out["n_simulations_used"] == 10
+
+ def test_all_significant(self):
+ proc = ResultsProcessor()
+ results = [np.array([True, True])] * 5
+ out = proc.calculate_powers(results, results, ["overall", "x1"])
+ assert out["individual_powers"]["overall"] == pytest.approx(100.0)
+ assert out["individual_powers"]["x1"] == pytest.approx(100.0)
+
+ def test_none_significant(self):
+ proc = ResultsProcessor()
+ results = [np.array([False, False])] * 5
+ out = proc.calculate_powers(results, results, ["overall", "x1"])
+ assert out["individual_powers"]["overall"] == pytest.approx(0.0)
+ assert out["individual_powers"]["x1"] == pytest.approx(0.0)
+
+ def test_combined_probabilities(self):
+ proc = ResultsProcessor()
+ # 4 sims, 2 tests: exactly 0, 1, 2 significant
+ results = [
+ np.array([False, False]), # 0 sig
+ np.array([True, False]), # 1 sig
+ np.array([False, True]), # 1 sig
+ np.array([True, True]), # 2 sig
+ ]
+ out = proc.calculate_powers(results, results, ["overall", "x1"])
+ combined = out["combined_probabilities"]
+ assert combined["exactly_0_significant"] == pytest.approx(25.0)
+ assert combined["exactly_1_significant"] == pytest.approx(50.0)
+ assert combined["exactly_2_significant"] == pytest.approx(25.0)
+
+ def test_cumulative_probabilities(self):
+ proc = ResultsProcessor()
+ results = [
+ np.array([False, False]),
+ np.array([True, True]),
+ np.array([True, True]),
+ np.array([True, True]),
+ ]
+ out = proc.calculate_powers(results, results, ["overall", "x1"])
+ cumulative = out["cumulative_probabilities"]
+ assert cumulative["at_least_0_significant"] == pytest.approx(100.0)
+ assert cumulative["at_least_2_significant"] == pytest.approx(75.0)
+
+
+class TestBuildPowerResult:
+ """Tests for build_power_result."""
+
+ def test_basic_structure(self):
+ power_results = {
+ "individual_powers": {"overall": 80.0},
+ "n_simulations_used": 1000,
+ }
+ result = build_power_result(
+ model_type="OLS",
+ target_tests=["overall"],
+ formula_to_test=None,
+ equation="y = x1",
+ sample_size=100,
+ alpha=0.05,
+ n_simulations=1000,
+ correction=None,
+ target_power=80.0,
+ parallel=False,
+ power_results=power_results,
+ )
+ assert result["model"]["model_type"] == "OLS"
+ assert result["model"]["sample_size"] == 100
+ assert result["model"]["alpha"] == 0.05
+ assert result["results"] is power_results
+
+
+class TestBuildSampleSizeResult:
+ """Tests for build_sample_size_result."""
+
+ def test_basic_structure(self):
+ analysis_results = {"sample_sizes_tested": [50, 100]}
+ result = build_sample_size_result(
+ model_type="OLS",
+ target_tests=["overall"],
+ formula_to_test=None,
+ equation="y = x1",
+ sample_sizes=[50, 100],
+ alpha=0.05,
+ n_simulations=1000,
+ correction=None,
+ target_power=80.0,
+ parallel=False,
+ analysis_results=analysis_results,
+ )
+ assert result["model"]["sample_size_range"]["from_size"] == 50
+ assert result["model"]["sample_size_range"]["to_size"] == 100
+ assert result["model"]["sample_size_range"]["by"] == 50
+ assert result["results"] is analysis_results
+
+ def test_single_sample_size(self):
+ result = build_sample_size_result(
+ model_type="OLS",
+ target_tests=["overall"],
+ formula_to_test=None,
+ equation="y = x1",
+ sample_sizes=[100],
+ alpha=0.05,
+ n_simulations=1000,
+ correction=None,
+ target_power=80.0,
+ parallel=False,
+ analysis_results={},
+ )
+ assert result["model"]["sample_size_range"]["by"] == 1
diff --git a/tests/unit/test_scenarios_coverage.py b/tests/unit/test_scenarios_coverage.py
new file mode 100644
index 0000000..ea0d4f2
--- /dev/null
+++ b/tests/unit/test_scenarios_coverage.py
@@ -0,0 +1,218 @@
+"""Tests for scenario analysis — plot creation, correlation matrix repair, LME perturbations."""
+
+from unittest.mock import MagicMock, patch
+
+import numpy as np
+import pytest
+
+from mcpower.core.scenarios import (
+ ScenarioRunner,
+ apply_lme_perturbations,
+ apply_per_simulation_perturbations,
+)
+
+
+class TestCorrelationMatrixRepair:
+ """Spectral clipping when noise creates negative eigenvalues."""
+
+ def test_negative_eigenvalue_repaired(self):
+ """After heavy noise, result should be positive semi-definite with unit diagonal."""
+ # Create a 3x3 identity correlation matrix
+ corr = np.eye(3)
+ var_types = np.zeros(3, dtype=np.int64) # all normal
+
+ config = {
+ "correlation_noise_sd": 2.0, # Very heavy noise → guaranteed negative eigenvalues
+ "distribution_change_prob": 0.0,
+ "new_distributions": [],
+ }
+
+ perturbed_corr, _ = apply_per_simulation_perturbations(corr, var_types, config, sim_seed=42)
+
+ # Eigenvalues should all be >= 0
+ eigvals = np.linalg.eigvalsh(perturbed_corr)
+ assert np.all(eigvals >= -1e-10)
+
+ # Diagonal should be 1.0
+ np.testing.assert_allclose(np.diag(perturbed_corr), 1.0, atol=1e-10)
+
+ # Should be symmetric
+ np.testing.assert_allclose(perturbed_corr, perturbed_corr.T, atol=1e-10)
+
+ def test_no_repair_needed_when_no_noise(self):
+ corr = np.array([[1.0, 0.3], [0.3, 1.0]])
+ var_types = np.zeros(2, dtype=np.int64)
+
+ config = {
+ "correlation_noise_sd": 0.0,
+ "distribution_change_prob": 0.0,
+ "new_distributions": [],
+ }
+
+ perturbed_corr, _ = apply_per_simulation_perturbations(corr, var_types, config, sim_seed=42)
+ np.testing.assert_array_equal(perturbed_corr, corr)
+
+
+class TestDistributionPerturbation:
+ """Variable type swaps in scenario mode."""
+
+ def test_distribution_swap_occurs(self):
+ var_types = np.zeros(10, dtype=np.int64) # All normal
+ config = {
+ "correlation_noise_sd": 0.0,
+ "distribution_change_prob": 1.0, # Always swap
+ "new_distributions": ["right_skewed"],
+ }
+
+ _, perturbed_types = apply_per_simulation_perturbations(
+ np.eye(10), var_types, config, sim_seed=42,
+ )
+ # All should be swapped from 0 to 2 (right_skewed)
+ assert np.all(perturbed_types == 2)
+
+ def test_non_normal_not_swapped(self):
+ """Binary (1) and uploaded (99) vars should not be swapped."""
+ var_types = np.array([0, 1, 99], dtype=np.int64)
+ config = {
+ "correlation_noise_sd": 0.0,
+ "distribution_change_prob": 1.0,
+ "new_distributions": ["right_skewed"],
+ }
+
+ _, perturbed_types = apply_per_simulation_perturbations(
+ np.eye(3), var_types, config, sim_seed=42,
+ )
+ assert perturbed_types[0] == 2 # normal → right_skewed
+ assert perturbed_types[1] == 1 # binary unchanged
+ assert perturbed_types[2] == 99 # uploaded unchanged
+
+ def test_none_config_passthrough(self):
+ corr = np.eye(2)
+ var_types = np.zeros(2, dtype=np.int64)
+ result_corr, result_types = apply_per_simulation_perturbations(
+ corr, var_types, None, sim_seed=42,
+ )
+ np.testing.assert_array_equal(result_corr, corr)
+ np.testing.assert_array_equal(result_types, var_types)
+
+
+class TestLMEPerturbations:
+ """LME perturbation computation."""
+
+ def test_icc_noise_creates_multipliers(self):
+ cluster_specs = {"school": {"n_clusters": 20, "cluster_size": 10, "icc": 0.2}}
+ config = {
+ "icc_noise_sd": 0.3,
+ "random_effect_dist": "normal",
+ "random_effect_df": 5,
+ }
+
+ result = apply_lme_perturbations(cluster_specs, config, sim_seed=42)
+ assert result is not None
+ assert "tau_squared_multipliers" in result
+ assert "school" in result["tau_squared_multipliers"]
+ # Multiplier should be exp(N(0, 0.3)) — positive, around 1
+ mult = result["tau_squared_multipliers"]["school"]
+ assert mult > 0
+
+ def test_no_perturbation_returns_none(self):
+ cluster_specs = {"school": {"n_clusters": 20, "cluster_size": 10, "icc": 0.2}}
+ config = {
+ "icc_noise_sd": 0.0,
+ "random_effect_dist": "normal",
+ "random_effect_df": 5,
+ }
+ result = apply_lme_perturbations(cluster_specs, config, sim_seed=42)
+ assert result is None
+
+ def test_empty_cluster_specs_returns_none(self):
+ result = apply_lme_perturbations({}, {"icc_noise_sd": 0.5}, sim_seed=42)
+ assert result is None
+
+ def test_heavy_tailed_re_dist(self):
+ cluster_specs = {"school": {"n_clusters": 20, "cluster_size": 10, "icc": 0.2}}
+ config = {
+ "icc_noise_sd": 0.0,
+ "random_effect_dist": "heavy_tailed",
+ "random_effect_df": 3,
+ }
+ result = apply_lme_perturbations(cluster_specs, config, sim_seed=42)
+ assert result is not None
+ assert result["random_effect_dist"] == "heavy_tailed"
+ assert result["random_effect_df"] == 3
+
+
+class TestScenarioRunnerPlots:
+ """Test _create_scenario_plots path."""
+
+ def test_plot_creation_with_mock(self):
+ model = MagicMock()
+ model.power = 80.0
+ runner = ScenarioRunner(model)
+
+ results = {
+ "analysis_type": "sample_size",
+ "scenarios": {
+ "optimistic": {
+ "model": {
+ "target_tests": ["x1"],
+ "correction": None,
+ },
+ "results": {
+ "sample_sizes_tested": [50, 100],
+ "powers_by_test": {"x1": [50.0, 85.0]},
+ "first_achieved": {"x1": 100},
+ },
+ },
+ },
+ }
+
+ with patch("mcpower.core.scenarios._create_power_plot") as mock_plot:
+ runner._create_scenario_plots(results)
+ mock_plot.assert_called_once()
+
+ def test_plot_with_correction(self):
+ model = MagicMock()
+ model.power = 80.0
+ runner = ScenarioRunner(model)
+
+ results = {
+ "analysis_type": "sample_size",
+ "scenarios": {
+ "optimistic": {
+ "model": {
+ "target_tests": ["x1"],
+ "correction": "bonferroni",
+ },
+ "results": {
+ "sample_sizes_tested": [50, 100],
+ "powers_by_test": {"x1": [50.0, 85.0]},
+ "powers_by_test_corrected": {"x1": [40.0, 75.0]},
+ "first_achieved": {"x1": 100},
+ "first_achieved_corrected": {"x1": 150},
+ },
+ },
+ },
+ }
+
+ with patch("mcpower.core.scenarios._create_power_plot") as mock_plot:
+ runner._create_scenario_plots(results)
+ # Should be called for both uncorrected and corrected
+ assert mock_plot.call_count == 2
+
+ def test_no_plot_when_missing_sample_sizes(self):
+ model = MagicMock()
+ model.power = 80.0
+ runner = ScenarioRunner(model)
+
+ results = {
+ "scenarios": {
+ "optimistic": {
+ "results": {"powers_by_test": {"x1": [50.0]}},
+ },
+ },
+ }
+
+ with patch("mcpower.core.scenarios._create_power_plot") as mock_plot:
+ runner._create_scenario_plots(results)
+ mock_plot.assert_not_called()
diff --git a/tests/unit/test_simulation_coverage.py b/tests/unit/test_simulation_coverage.py
new file mode 100644
index 0000000..8bae7ff
--- /dev/null
+++ b/tests/unit/test_simulation_coverage.py
@@ -0,0 +1,274 @@
+"""Tests for simulation.py — failure handling, Wald fallback, verbose diagnostics, ICC mismatch."""
+
+import warnings
+from typing import Dict, List, Optional
+from unittest.mock import MagicMock, patch
+
+import numpy as np
+import pytest
+
+from mcpower.core.simulation import SimulationMetadata, SimulationRunner, _warn_icc_mismatch
+
+
+def _make_metadata(
+ n_targets=2,
+ cluster_specs=None,
+ verbose=False,
+ correction_method=0,
+):
+ """Create a minimal SimulationMetadata for testing."""
+ return SimulationMetadata(
+ target_indices=np.arange(n_targets),
+ n_non_factor_vars=n_targets,
+ correlation_matrix=np.eye(n_targets),
+ var_types=np.zeros(n_targets, dtype=np.int64),
+ var_params=np.zeros(n_targets, dtype=np.float64),
+ factor_specs=[],
+ upload_normal_values=np.zeros((2, 2), dtype=np.float64),
+ upload_data_values=np.zeros((2, 2), dtype=np.float64),
+ effect_sizes=np.array([0.5] * n_targets),
+ correction_method=correction_method,
+ cluster_specs=cluster_specs or {},
+ verbose=verbose,
+ )
+
+
+def _noop_perturbations(corr, types, config, seed):
+ return corr, types
+
+
+class TestAllSimulationsFail:
+ """When all simulations return None, RuntimeError should be raised."""
+
+ def test_all_fail_raises(self):
+ runner = SimulationRunner(n_simulations=5, seed=42)
+ metadata = _make_metadata()
+
+ def failing_sim(*args, **kwargs):
+ return None
+
+ with patch.object(runner, "_single_simulation", return_value=None):
+ with pytest.raises(RuntimeError, match="All simulations failed"):
+ runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+
+
+class TestLMEThresholdExceeded:
+ """LME failure rate exceeding threshold raises RuntimeError."""
+
+ def test_high_failure_rate_raises(self):
+ runner = SimulationRunner(n_simulations=10, seed=42, max_failed_simulations=0.05)
+ metadata = _make_metadata(cluster_specs={"school": {"n_clusters": 5, "cluster_size": 10}})
+
+ call_count = [0]
+
+ def sometimes_fail(*args, **kwargs):
+ call_count[0] += 1
+ if call_count[0] <= 5:
+ return None # 5 out of 10 fail = 50%
+ return (np.array([1, 1, 1]), np.array([1, 1, 1]), False)
+
+ with patch.object(runner, "_single_simulation", side_effect=sometimes_fail):
+ with pytest.raises(RuntimeError, match="Too many failed simulations"):
+ runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+
+
+class TestOLSHighFailureWarns:
+ """OLS high failure rate warns but doesn't raise."""
+
+ def test_ols_warns_above_10_percent(self):
+ runner = SimulationRunner(n_simulations=10, seed=42)
+ metadata = _make_metadata() # No cluster_specs = OLS
+
+ call_count = [0]
+
+ def sometimes_fail(*args, **kwargs):
+ call_count[0] += 1
+ if call_count[0] <= 2:
+ return None # 2 out of 10 fail = 20%
+ return (np.array([1, 1, 1]), np.array([1, 1, 1]))
+
+ with patch.object(runner, "_single_simulation", side_effect=sometimes_fail):
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ result = runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+ assert any("failed" in str(warning.message).lower() for warning in w)
+
+
+class TestWaldFallbackWarning:
+ """Warn if >10% iterations use Wald test."""
+
+ def test_wald_warning_above_threshold(self):
+ runner = SimulationRunner(n_simulations=10, seed=42)
+ metadata = _make_metadata()
+
+ call_count = [0]
+
+ def wald_heavy(*args, **kwargs):
+ call_count[0] += 1
+ # All return wald_flag=True
+ return (np.array([1, 1, 1]), np.array([1, 1, 1]), True)
+
+ with patch.object(runner, "_single_simulation", side_effect=wald_heavy):
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ result = runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+ assert any("Wald test fallback" in str(warning.message) for warning in w)
+ assert result["n_wald_fallbacks"] == 10
+
+
+class TestVerboseDiagnostics:
+ """Verbose mode collects diagnostics and failure reasons."""
+
+ def test_verbose_success_collects_diagnostics(self):
+ runner = SimulationRunner(n_simulations=3, seed=42)
+ metadata = _make_metadata(verbose=True)
+
+ def verbose_result(*args, **kwargs):
+ return {
+ "results": (np.array([1, 1, 1]), np.array([1, 1, 1])),
+ "diagnostics": {"icc_estimated": 0.2},
+ "wald_fallback": False,
+ }
+
+ with patch.object(runner, "_single_simulation", side_effect=verbose_result):
+ result = runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+ assert "diagnostics" in result
+ assert len(result["diagnostics"]) == 3
+
+ def test_verbose_failure_tracking(self):
+ runner = SimulationRunner(n_simulations=5, seed=42)
+ metadata = _make_metadata(verbose=True)
+
+ call_count = [0]
+
+ def mixed_results(*args, **kwargs):
+ call_count[0] += 1
+ if call_count[0] <= 2:
+ return {"failed": True, "failure_reason": "Convergence failed"}
+ return {
+ "results": (np.array([1, 1, 1]), np.array([1, 1, 1])),
+ "diagnostics": {},
+ "wald_fallback": False,
+ }
+
+ with patch.object(runner, "_single_simulation", side_effect=mixed_results):
+ with warnings.catch_warnings():
+ warnings.simplefilter("ignore", UserWarning)
+ result = runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+ assert "failure_reasons" in result
+ assert result["failure_reasons"]["Convergence failed"] == 2
+
+ def test_verbose_none_tracking(self):
+ """None results in verbose mode are tracked as unknown failures."""
+ runner = SimulationRunner(n_simulations=3, seed=42)
+ metadata = _make_metadata(verbose=True)
+
+ call_count = [0]
+
+ def mixed(*args, **kwargs):
+ call_count[0] += 1
+ if call_count[0] == 1:
+ return None
+ return {
+ "results": (np.array([1, 1, 1]), np.array([1, 1, 1])),
+ "diagnostics": {},
+ "wald_fallback": False,
+ }
+
+ with patch.object(runner, "_single_simulation", side_effect=mixed):
+ with warnings.catch_warnings():
+ warnings.simplefilter("ignore", UserWarning)
+ result = runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+ assert "Unknown (returned None)" in result["failure_reasons"]
+
+
+class TestICCMismatchWarning:
+ """ICC mismatch warning when estimated ICC differs by >50%."""
+
+ def test_large_mismatch_warns(self):
+ metadata = _make_metadata(
+ cluster_specs={"school": {"icc": 0.2, "n_clusters": 20, "cluster_size": 10}},
+ )
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ _warn_icc_mismatch(metadata, mean_estimated_icc=0.05) # 75% deviation
+ assert any("differs from specified" in str(warning.message) for warning in w)
+
+ def test_within_tolerance_no_warning(self):
+ metadata = _make_metadata(
+ cluster_specs={"school": {"icc": 0.2, "n_clusters": 20, "cluster_size": 10}},
+ )
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ _warn_icc_mismatch(metadata, mean_estimated_icc=0.18) # 10% deviation
+ icc_warnings = [x for x in w if "differs from specified" in str(x.message)]
+ assert len(icc_warnings) == 0
+
+ def test_zero_estimated_icc_no_warning(self):
+ metadata = _make_metadata(
+ cluster_specs={"school": {"icc": 0.2, "n_clusters": 20, "cluster_size": 10}},
+ )
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ _warn_icc_mismatch(metadata, mean_estimated_icc=0.0)
+ icc_warnings = [x for x in w if "differs from specified" in str(x.message)]
+ assert len(icc_warnings) == 0
+
+ def test_no_icc_in_spec_no_warning(self):
+ metadata = _make_metadata(
+ cluster_specs={"school": {"icc": None, "n_clusters": 20, "cluster_size": 10}},
+ )
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ _warn_icc_mismatch(metadata, mean_estimated_icc=0.5)
+ icc_warnings = [x for x in w if "differs from specified" in str(x.message)]
+ assert len(icc_warnings) == 0
diff --git a/tests/unit/test_test_formula_utils.py b/tests/unit/test_test_formula_utils.py
new file mode 100644
index 0000000..f3db882
--- /dev/null
+++ b/tests/unit/test_test_formula_utils.py
@@ -0,0 +1,319 @@
+"""Tests for test_formula parsing utilities."""
+
+from collections import OrderedDict
+from unittest.mock import MagicMock
+
+import numpy as np
+
+
+class TestExtractTestFormulaEffects:
+ """Test _extract_test_formula_effects helper."""
+
+ def _make_registry(
+ self,
+ effect_names,
+ factor_names=None,
+ factor_dummies=None,
+ cluster_effect_names=None,
+ ):
+ """Create a minimal mock registry for testing."""
+ reg = MagicMock()
+ reg.effect_names = effect_names
+ reg.factor_names = factor_names or []
+ reg.cluster_effect_names = cluster_effect_names or []
+
+ # Build _effects dict with correct ordering
+ effects = OrderedDict()
+ for name in effect_names:
+ eff = MagicMock()
+ eff.effect_type = "interaction" if ":" in name else "main"
+ effects[name] = eff
+ reg._effects = effects
+
+ # Factor dummies
+ reg._factor_dummies = factor_dummies or {}
+ return reg
+
+ def test_simple_subset(self):
+ """y ~ x1 + x2 from generation y ~ x1 + x2 + x3."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3"])
+ effects, random_effects = _extract_test_formula_effects("y ~ x1 + x2", registry)
+ assert effects == ["x1", "x2"]
+ assert random_effects == []
+
+ def test_single_variable(self):
+ """y ~ x1 from generation y ~ x1 + x2 + x3."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3"])
+ effects, random_effects = _extract_test_formula_effects("y ~ x1", registry)
+ assert effects == ["x1"]
+
+ def test_with_interaction(self):
+ """y ~ x1 + x2 + x1:x2 from generation y ~ x1 + x2 + x3 + x1:x2."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3", "x1:x2"])
+ effects, _ = _extract_test_formula_effects("y ~ x1 + x2 + x1:x2", registry)
+ assert effects == ["x1", "x2", "x1:x2"]
+
+ def test_interaction_omitted(self):
+ """y ~ x1 + x2 from generation y ~ x1 + x2 + x1:x2."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x1:x2"])
+ effects, _ = _extract_test_formula_effects("y ~ x1 + x2", registry)
+ assert effects == ["x1", "x2"]
+
+ def test_factor_expands_to_dummies(self):
+ """y ~ x1 + gender from generation y ~ x1 + x2 + gender."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(
+ ["x1", "x2", "gender[F]", "gender[Other]"],
+ factor_names=["gender"],
+ factor_dummies={
+ "gender[F]": {"factor_name": "gender", "level": "F"},
+ "gender[Other]": {"factor_name": "gender", "level": "Other"},
+ },
+ )
+ effects, _ = _extract_test_formula_effects("y ~ x1 + gender", registry)
+ assert effects == ["x1", "gender[F]", "gender[Other]"]
+
+ def test_factor_omitted(self):
+ """y ~ x1 from generation y ~ x1 + gender."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(
+ ["x1", "gender[F]", "gender[Other]"],
+ factor_names=["gender"],
+ factor_dummies={
+ "gender[F]": {"factor_name": "gender", "level": "F"},
+ "gender[Other]": {"factor_name": "gender", "level": "Other"},
+ },
+ )
+ effects, _ = _extract_test_formula_effects("y ~ x1", registry)
+ assert effects == ["x1"]
+
+ def test_with_random_effects(self):
+ """y ~ x1 + (1|school) extracts random effects."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2"])
+ effects, random_effects = _extract_test_formula_effects(
+ "y ~ x1 + (1|school)", registry
+ )
+ assert effects == ["x1"]
+ assert len(random_effects) == 1
+ assert random_effects[0]["grouping_var"] == "school"
+
+ def test_star_operator_expands(self):
+ """y ~ x1*x2 expands to x1 + x2 + x1:x2."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3", "x1:x2"])
+ effects, _ = _extract_test_formula_effects("y ~ x1*x2", registry)
+ assert effects == ["x1", "x2", "x1:x2"]
+
+ def test_equals_sign_formula(self):
+ """y = x1 + x2 works same as y ~ x1 + x2."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3"])
+ effects, _ = _extract_test_formula_effects("y = x1 + x2", registry)
+ assert effects == ["x1", "x2"]
+
+ def test_preserves_registry_order(self):
+ """Effects returned in registry order, not formula order."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3", "x1:x2"])
+ # Formula lists x2 before x1
+ effects, _ = _extract_test_formula_effects("y ~ x2 + x1", registry)
+ assert effects == ["x1", "x2"] # registry order preserved
+
+
+class TestComputeTestColumnIndices:
+ """Test _compute_test_column_indices helper."""
+
+ def test_subset_two_of_three(self):
+ """Selecting 2 of 3 effects gives correct indices."""
+ from mcpower.utils.test_formula_utils import _compute_test_column_indices
+
+ all_effect_names = ["x1", "x2", "x3"]
+ test_effect_names = ["x1", "x2"]
+ result = _compute_test_column_indices(all_effect_names, test_effect_names)
+ assert list(result) == [0, 1]
+
+ def test_skip_middle(self):
+ """Selecting first and last of 3 effects."""
+ from mcpower.utils.test_formula_utils import _compute_test_column_indices
+
+ all_effect_names = ["x1", "x2", "x3"]
+ test_effect_names = ["x1", "x3"]
+ result = _compute_test_column_indices(all_effect_names, test_effect_names)
+ assert list(result) == [0, 2]
+
+ def test_single_effect(self):
+ """Single effect selected."""
+ from mcpower.utils.test_formula_utils import _compute_test_column_indices
+
+ all_effect_names = ["x1", "x2", "x3"]
+ test_effect_names = ["x2"]
+ result = _compute_test_column_indices(all_effect_names, test_effect_names)
+ assert list(result) == [1]
+
+ def test_all_effects_returns_all_indices(self):
+ """Selecting all effects returns full range."""
+ from mcpower.utils.test_formula_utils import _compute_test_column_indices
+
+ all_effect_names = ["x1", "x2", "x3"]
+ test_effect_names = ["x1", "x2", "x3"]
+ result = _compute_test_column_indices(all_effect_names, test_effect_names)
+ assert list(result) == [0, 1, 2]
+
+ def test_with_interactions(self):
+ """Interaction effects have correct indices."""
+ from mcpower.utils.test_formula_utils import _compute_test_column_indices
+
+ all_effect_names = ["x1", "x2", "x3", "x1:x2"]
+ test_effect_names = ["x1", "x2", "x1:x2"]
+ result = _compute_test_column_indices(all_effect_names, test_effect_names)
+ assert list(result) == [0, 1, 3]
+
+
+class TestRemapTargetIndices:
+ """Test _remap_target_indices helper."""
+
+ def test_simple_remap(self):
+ """Target indices remapped to positions within test columns."""
+ from mcpower.utils.test_formula_utils import _remap_target_indices
+
+ # Original target_indices: [0, 1] (x1, x2 in full model)
+ # test_column_indices: [0, 1] (x1, x2 at positions 0, 1 in X_expanded)
+ # In X_test, x1 is at 0, x2 is at 1 -> remapped: [0, 1]
+ original = np.array([0, 1])
+ test_cols = np.array([0, 1])
+ result = _remap_target_indices(original, test_cols)
+ assert list(result) == [0, 1]
+
+ def test_remap_with_gap(self):
+ """Target indices remapped when test columns skip positions."""
+ from mcpower.utils.test_formula_utils import _remap_target_indices
+
+ # Full model: [x1=0, x2=1, x3=2, x1:x2=3]
+ # Test model: [x1=0, x1:x2=3] -> X_test columns at [0, 3]
+ # target_test="x1" -> original target_indices=[0]
+ # In X_test, x1 is at position 0 -> remapped: [0]
+ original = np.array([0])
+ test_cols = np.array([0, 3])
+ result = _remap_target_indices(original, test_cols)
+ assert list(result) == [0]
+
+ def test_remap_target_at_end(self):
+ """Target index that moves to different position in X_test."""
+ from mcpower.utils.test_formula_utils import _remap_target_indices
+
+ # Full model: [x1=0, x2=1, x3=2]
+ # Test model: [x2=1, x3=2] -> test_column_indices=[1, 2]
+ # target_test="x3" -> original target_indices=[2]
+ # In X_test, x3 is at position 1 (second column) -> remapped: [1]
+ original = np.array([2])
+ test_cols = np.array([1, 2])
+ result = _remap_target_indices(original, test_cols)
+ assert list(result) == [1]
+
+
+class TestPrepareMetadataWithTestFormula:
+ """Integration test: prepare_metadata with test_formula_effects."""
+
+ def test_metadata_has_test_indices_when_provided(self):
+ from mcpower import MCPower
+ from mcpower.core.simulation import prepare_metadata
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ metadata = prepare_metadata(model, ["x1", "x2"], test_formula_effects=["x1", "x2"])
+ assert metadata.test_column_indices is not None
+ assert list(metadata.test_column_indices) == [0, 1]
+ assert metadata.test_target_indices is not None
+ assert metadata.test_effect_count == 2
+
+ def test_metadata_no_test_indices_by_default(self):
+ from mcpower import MCPower
+ from mcpower.core.simulation import prepare_metadata
+
+ model = MCPower("y = x1 + x2")
+ model.set_effects("x1=0.5, x2=0.3")
+ model._apply()
+
+ metadata = prepare_metadata(model, ["x1", "x2"])
+ assert metadata.test_column_indices is None
+
+ def test_remap_skips_targets_not_in_test_formula(self):
+ from mcpower import MCPower
+ from mcpower.core.simulation import prepare_metadata
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ # target_tests = all 3, but test formula only has x1, x2
+ metadata = prepare_metadata(model, ["x1", "x2", "x3"], test_formula_effects=["x1", "x2"])
+ # test_target_indices should only have indices for x1 and x2 in X_test
+ assert len(metadata.test_target_indices) == 2
+
+
+class TestParseTargetTestsWithTestFormula:
+ """Test _parse_target_tests limits 'all' when test_formula is active."""
+
+ def test_all_expands_to_test_formula_effects_only(self):
+ from mcpower import MCPower
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ result = model._parse_target_tests("all", test_formula_effects=["x1", "x2"])
+ assert "x3" not in result
+ assert "x1" in result
+ assert "x2" in result
+ assert "overall" in result
+
+ def test_explicit_target_not_in_test_formula_raises(self):
+ from mcpower import MCPower
+
+ import pytest
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ with pytest.raises(ValueError, match="x3"):
+ model._parse_target_tests("x3", test_formula_effects=["x1", "x2"])
+
+ def test_overall_always_allowed(self):
+ from mcpower import MCPower
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ result = model._parse_target_tests("overall", test_formula_effects=["x1", "x2"])
+ assert "overall" in result
+
+ def test_no_test_formula_uses_all_effects(self):
+ from mcpower import MCPower
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ result = model._parse_target_tests("all")
+ assert "x1" in result
+ assert "x2" in result
+ assert "x3" in result
diff --git a/tests/unit/test_updates.py b/tests/unit/test_updates.py
index ee7a2a0..90b3003 100644
--- a/tests/unit/test_updates.py
+++ b/tests/unit/test_updates.py
@@ -101,14 +101,17 @@ def test_shows_warning_when_newer(self, monkeypatch):
"""Show warning when PyPI version is newer."""
monkeypatch.delenv("_MCPOWER_UPDATE_CHECKED", raising=False)
- # Write a cache file at the path the installed module actually reads from
- from datetime import datetime
-
import mcpower.utils.updates as upd_mod
+
+ # Reset the module-level dedup flag
+ upd_mod._already_checked = False
+
+ # Write a cache file at the path the module actually reads from
+ from datetime import datetime
from pathlib import Path
- cache_path = Path(upd_mod.__file__).parent.parent / ".mcpower_cache.json"
- cache_path.parent.mkdir(exist_ok=True)
+ cache_path = Path.home() / ".cache" / "mcpower" / "update_cache.json"
+ cache_path.parent.mkdir(parents=True, exist_ok=True)
cache_data = {
"last_check": datetime.now().isoformat(),
"latest_version": "99.0.0",
@@ -120,5 +123,6 @@ def test_shows_warning_when_newer(self, monkeypatch):
with pytest.warns(match="NEW MCPower VERSION"):
_check_for_updates("1.0.0")
finally:
- # Clean up the cache file
+ # Clean up the cache file and reset flag
cache_path.unlink(missing_ok=True)
+ upd_mod._already_checked = False
diff --git a/tests/unit/test_upload_data_utils.py b/tests/unit/test_upload_data_utils.py
new file mode 100644
index 0000000..c498b6a
--- /dev/null
+++ b/tests/unit/test_upload_data_utils.py
@@ -0,0 +1,62 @@
+"""Unit tests for mcpower.utils.upload_data_utils — normalize_upload_input."""
+
+import numpy as np
+import pytest
+
+from mcpower.utils.upload_data_utils import normalize_upload_input
+
+
+class TestNormalizeUploadInput:
+ """Tests for normalize_upload_input."""
+
+ def test_dict_input(self):
+ data = {"x1": [1.0, 2.0, 3.0], "x2": [4.0, 5.0, 6.0]}
+ arr, cols = normalize_upload_input(data)
+ assert cols == ["x1", "x2"]
+ assert arr.shape == (3, 2)
+ np.testing.assert_array_equal(arr[:, 0], [1.0, 2.0, 3.0])
+
+ def test_dict_with_strings(self):
+ data = {"group": ["a", "b", "a"], "x1": [1.0, 2.0, 3.0]}
+ arr, cols = normalize_upload_input(data)
+ assert arr.dtype == object
+ assert cols == ["group", "x1"]
+
+ def test_list_input(self):
+ data = [1.0, 2.0, 3.0]
+ arr, cols = normalize_upload_input(data)
+ assert arr.shape == (3, 1)
+ assert cols == ["column_1"]
+
+ def test_1d_array(self):
+ data = np.array([1.0, 2.0, 3.0])
+ arr, cols = normalize_upload_input(data)
+ assert arr.shape == (3, 1)
+ assert cols == ["column_1"]
+
+ def test_2d_array(self):
+ data = np.array([[1.0, 2.0], [3.0, 4.0]])
+ arr, cols = normalize_upload_input(data)
+ assert arr.shape == (2, 2)
+ assert cols == ["column_1", "column_2"]
+
+ def test_2d_array_with_columns(self):
+ data = np.array([[1.0, 2.0], [3.0, 4.0]])
+ arr, cols = normalize_upload_input(data, columns=["a", "b"])
+ assert cols == ["a", "b"]
+
+ def test_dataframe_input(self):
+ pd = pytest.importorskip("pandas")
+ df = pd.DataFrame({"x1": [1.0, 2.0], "x2": [3.0, 4.0]})
+ arr, cols = normalize_upload_input(df)
+ assert cols == ["x1", "x2"]
+ assert arr.shape == (2, 2)
+
+ def test_mismatched_columns_raises(self):
+ data = np.array([[1.0, 2.0], [3.0, 4.0]])
+ with pytest.raises(ValueError, match="columns length"):
+ normalize_upload_input(data, columns=["a", "b", "c"])
+
+ def test_unsupported_type_raises(self):
+ with pytest.raises(TypeError, match="data must be"):
+ normalize_upload_input("not valid data")
diff --git a/tests/unit/test_utils_mixed_models.py b/tests/unit/test_utils_mixed_models.py
new file mode 100644
index 0000000..de4d98a
--- /dev/null
+++ b/tests/unit/test_utils_mixed_models.py
@@ -0,0 +1,27 @@
+"""Tests for mcpower.utils.mixed_models backward-compat re-exports."""
+
+import threading
+
+from mcpower.utils.mixed_models import (
+ _lme_analysis_wrapper,
+ _lme_thread_local,
+ reset_warm_start_cache,
+)
+
+
+class TestReExports:
+ """Verify that the backward-compatibility re-exports resolve correctly."""
+
+ def test_lme_analysis_wrapper_is_callable(self):
+ assert callable(_lme_analysis_wrapper)
+
+ def test_lme_thread_local_is_threading_local(self):
+ assert isinstance(_lme_thread_local, threading.local)
+
+ def test_reset_warm_start_cache_is_callable(self):
+ assert callable(reset_warm_start_cache)
+
+ def test_reset_warm_start_cache_clears_params(self):
+ _lme_thread_local.warm_start_params = "dummy"
+ reset_warm_start_cache()
+ assert _lme_thread_local.warm_start_params is None
|
+
## Why MCPower?
Traditional power formulas break down with interactions, correlated predictors, categorical variables, or non-normal data. MCPower simulates instead — generates thousands of datasets like yours, fits your model, and counts how often the effects are detected.
@@ -297,19 +301,20 @@ model.set_effects("group[2]=0.4, group[3]=0.6, covariate=0.3")
# Use "vs" syntax for pairwise comparisons + correction="tukey"
model.find_power(
sample_size=150,
- target_test="group[0] vs group[1], group[0] vs group[2]",
+ target_test="group[1] vs group[2], group[1] vs group[3]",
correction="tukey"
)
```
### Test Individual Assumption Violations
```python
-# Manually add specific violations (without full scenario analysis)
-model.set_heterogeneity(0.2) # Effect sizes vary between people
-model.set_heteroskedasticity(0.15) # Violation of equal variance assumption
+# Add specific violations via custom scenario configs
+model.set_scenario_configs({
+ "my_test": {"heterogeneity": 0.2, "heteroskedasticity": 0.15}
+})
-# Run with your manual settings (no automatic scenario variations)
-model.find_sample_size(target_test="treatment")
+# Run with scenario variations
+model.find_sample_size(target_test="treatment", scenarios=True)
```
### Mixed-Effects Models
@@ -392,7 +397,7 @@ model.find_power(sample_size=200, progress_callback=False)
| **Factor effects** | **`model.set_effects("var[2]=0.5, var[3]=0.7")`** |
| Correlated predictors | `model.set_correlations("corr(var1, var2)=0.4")` |
| Multiple testing correction | Add `correction="FDR"`, `"Holm"`, `"Bonferroni"`, or `"Tukey"`|
-| Post-hoc pairwise comparison | `target_test="group[0] vs group[1]"` with `correction="tukey"` |
+| Post-hoc pairwise comparison | `target_test="group[1] vs group[2]"` with `correction="tukey"` |
| Mixed model (random intercept) | `MCPower("y ~ x + (1\|group)")` + `model.set_cluster(...)` |
| Random slopes | `MCPower("y ~ x + (1+x\|group)")` + `set_cluster(..., random_slopes=["x"], slope_variance=0.1)` |
| Nested random effects | `MCPower("y ~ x + (1\|A/B)")` + two `set_cluster()` calls |
@@ -424,7 +429,7 @@ model.find_power(sample_size=200, progress_callback=False)
- For simple models where all assumptions are clearly met.
- For large analyses with tens of thousands of observations, tiny effects, or very low alpha levels.
-## What Makes Scenarios Different? (Be careful, unvalidated, preliminary scenarios)
+## What Makes Scenarios Different? (Rule-of-thumb scenarios)
**Traditional power analysis assumes perfect conditions.** MCPower's scenarios add realistic "messiness":
@@ -478,8 +483,8 @@ model.set_variable_type("treatment=(factor,3), education=(factor,4)")
# Set effects for specific levels
model.set_effects("treatment[2]=0.5, treatment[3]=0.7, education[2]=0.3")
-# Or set same effect for all levels of a factor
-model.set_effects("treatment=0.5") # Applies to treatment[2] and treatment[3]
+# Each non-reference level needs its own effect
+model.set_effects("treatment[2]=0.5, treatment[3]=0.7")
# Important: Factors cannot be used in correlations
# This will error: model.set_correlations("corr(treatment, education)=0.3")
@@ -508,12 +513,31 @@ model.set_alpha(0.01) # Stricter significance (p < 0.01)
model.set_simulations(10000) # High precision (slower)
```
+### Model Misspecification Testing
+
+Use `test_formula` to generate data with one model but test with a simpler one -- useful for evaluating the power impact of omitting variables:
+
+```python
+# Generate with 3 predictors, test with 2 (omitting x3)
+model = MCPower("y = x1 + x2 + x3")
+model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+model.find_power(100, test_formula="y = x1 + x2")
+
+# Generate with clusters, test without (ignoring clustering)
+model = MCPower("y ~ treatment + (1|school)")
+model.set_cluster("school", ICC=0.2, n_clusters=20)
+model.set_effects("treatment=0.5")
+model.find_power(1000, test_formula="y ~ treatment")
+```
+
+See the [Test Formula Tutorial](https://github.com/pawlenartowicz/MCPower/wiki/Tutorial-Test-Formula) for details.
+
### Formula Syntax
```python
# These are equivalent:
-"y = x1 + x2 + x1*x2" # Assignment style
-"y ~ x1 + x2 + x1*x2" # R-style formula
-"x1 + x2 + x1*x2" # Predictors only
+"y = x1 + x2 + x1:x2" # Assignment style
+"y ~ x1 + x2 + x1:x2" # R-style formula
+"x1 + x2 + x1:x2" # Predictors only
# Interactions:
"x1*x2" # Main effects + interaction (x1 + x2 + x1:x2)
@@ -538,9 +562,8 @@ model.set_correlations("(x1, x2)=0.3, (x1, x3)=-0.2")
## Requirements
- Python ≥ 3.10
-- NumPy, matplotlib, joblib
+- NumPy (≥1.26.0), matplotlib, joblib, tqdm
- pandas (optional, for DataFrame input — install with `pip install mcpower[pandas]`)
-- statsmodels (optional, for mixed-effects models — install with `pip install mcpower[all]`)
## Documentation
@@ -549,11 +572,11 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
- [Quick Start](https://github.com/pawlenartowicz/MCPower/wiki/Quick-Start)
- [Model Specification](https://github.com/pawlenartowicz/MCPower/wiki/Model-Specification)
-- [Variable Types](https://github.com/pawlenartowicz/MCPower/wiki/Variable-Types)
-- [Effect Sizes](https://github.com/pawlenartowicz/MCPower/wiki/Effect-Sizes)
-- [Mixed-Effects Models](https://github.com/pawlenartowicz/MCPower/wiki/Mixed-Effects-Models) (random intercepts, slopes, nested effects)
-- [ANOVA & Post-Hoc Tests](https://github.com/pawlenartowicz/MCPower/wiki/ANOVA-and-Post-Hoc-Tests)
-- [Scenario Analysis](https://github.com/pawlenartowicz/MCPower/wiki/Scenario-Analysis)
+- [Variable Types](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Variable-Types)
+- [Effect Sizes](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Effect-Sizes)
+- [Mixed-Effects Models](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Mixed-Effects) (random intercepts, slopes, nested effects)
+- [ANOVA & Post-Hoc Tests](https://github.com/pawlenartowicz/MCPower/wiki/Tutorial-ANOVA-PostHoc)
+- [Scenario Analysis](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Scenario-Analysis)
- [API Reference](https://github.com/pawlenartowicz/MCPower/wiki/API-Reference)
## Need Help?
@@ -568,8 +591,8 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
- ✅ C++ native backend (pybind11 + Eigen, 3x speedup)
- ✅ Mixed Effects Models (random intercepts, random slopes, nested effects) — [validated against lme4](https://github.com/pawlenartowicz/MCPower/wiki/Concept-LME-Validation)
- 🚧 Logistic Regression (coming soon)
-- 🚧 ANOVA (coming soon)
-- 🚧 Guide about methods, corrections (coming soon)
+- ✅ ANOVA (factor variables as ANOVA, post-hoc pairwise comparisons)
+- ✅ Guide about methods, corrections
- 📋 2 groups comparison with alternative tests
- 📋 Robust regression methods
@@ -578,16 +601,18 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
GPL v3. If you use MCPower in research, please cite:
-Lenartowicz, P. (2025). MCPower: Monte Carlo Power Analysis for Statistical Models. Zenodo. DOI: 10.5281/zenodo.16502734
+Lenartowicz, P. (2025). MCPower: Monte Carlo Power Analysis for Complex Statistical Models (Version ) [Computer software]. Zenodo. https://doi.org/10.5281/zenodo.16502734
+
+*Replace `` with the version you used — check with `import mcpower; print(mcpower.__version__)`.*
```bibtex
@software{mcpower2025,
- author = {Pawel Lenartowicz},
- title = {MCPower: Monte Carlo Power Analysis for Statistical Models},
- year = {2025},
+ author = {Lenartowicz, Pawe{\l}},
+ title = {{MCPower}: Monte Carlo Power Analysis for Complex Statistical Models},
+ year = {2025},
publisher = {Zenodo},
- doi = {10.5281/zenodo.16502734},
- url = {https://doi.org/10.5281/zenodo.16502734}
+ doi = {10.5281/zenodo.16502734},
+ url = {https://doi.org/10.5281/zenodo.16502734}
}
```
diff --git a/cpp/src/bindings.cpp b/cpp/src/bindings.cpp
index 26fee22..8c02998 100644
--- a/cpp/src/bindings.cpp
+++ b/cpp/src/bindings.cpp
@@ -110,7 +110,9 @@ py::array_t generate_y_wrapper(
py::array_t effects,
double heterogeneity,
double heteroskedasticity,
- int seed
+ int seed,
+ int residual_dist,
+ double residual_df
) {
auto X_buf = X.request();
auto effects_buf = effects.request();
@@ -129,7 +131,8 @@ py::array_t generate_y_wrapper(
);
Eigen::VectorXd y = generate_y(
- X_map, effects_map, heterogeneity, heteroskedasticity, seed
+ X_map, effects_map, heterogeneity, heteroskedasticity, seed,
+ residual_dist, residual_df
);
py::array_t result(n);
@@ -447,7 +450,9 @@ PYBIND11_MODULE(mcpower_native, m) {
py::arg("heterogeneity") = 0.0,
py::arg("heteroskedasticity") = 0.0,
py::arg("seed") = -1,
- "Generate dependent variable with heterogeneity and heteroskedasticity"
+ py::arg("residual_dist") = 0,
+ py::arg("residual_df") = 10.0,
+ "Generate dependent variable with heterogeneity, heteroskedasticity, and non-normal residuals"
);
// LME analysis (q=1 random intercept)
diff --git a/cpp/src/ols.cpp b/cpp/src/ols.cpp
index 7d04ec2..11bd62f 100644
--- a/cpp/src/ols.cpp
+++ b/cpp/src/ols.cpp
@@ -151,19 +151,14 @@ Eigen::VectorXd generate_y(
const Eigen::Ref& effects,
double heterogeneity,
double heteroskedasticity,
- int seed
+ int seed,
+ int residual_dist,
+ double residual_df
) {
const int n = static_cast(X.rows());
const int p = static_cast(X.cols());
- // Set up random generator
std::mt19937 gen;
- if (seed >= 0) {
- gen.seed(static_cast(seed));
- } else {
- std::random_device rd;
- gen.seed(rd());
- }
std::normal_distribution normal(0.0, 1.0);
// Linear predictor with heterogeneity
@@ -176,9 +171,12 @@ Eigen::VectorXd generate_y(
// Heterogeneity: vary effect sizes per observation
linear_pred.setZero();
- // Change seed for heterogeneity noise
+ // Seed at offset +1 for heterogeneity noise
if (seed >= 0) {
gen.seed(static_cast(seed + 1));
+ } else {
+ std::random_device rd;
+ gen.seed(rd());
}
for (int j = 0; j < p; ++j) {
@@ -192,14 +190,43 @@ Eigen::VectorXd generate_y(
}
}
- // Generate errors
+ // Generate errors — seed at offset +2
if (seed >= 0) {
gen.seed(static_cast(seed + 2));
+ } else {
+ std::random_device rd;
+ gen.seed(rd());
}
Eigen::VectorXd error(n);
- for (int i = 0; i < n; ++i) {
- error(i) = normal(gen);
+
+ if (residual_dist == 1) {
+ // Heavy-tailed: Student's t distribution
+ double df = std::max(residual_df, 3.0);
+ std::student_t_distribution t_dist(df);
+ double theoretical_scale = 1.0 / std::sqrt(df / (df - 2.0));
+ for (int i = 0; i < n; ++i) {
+ error(i) = t_dist(gen) * theoretical_scale;
+ }
+ } else if (residual_dist == 2) {
+ // Skewed: chi-squared, centered and scaled
+ double df = std::max(residual_df, 3.0);
+ std::chi_squared_distribution chi2_dist(df);
+ double scale = 1.0 / std::sqrt(2.0 * df);
+ for (int i = 0; i < n; ++i) {
+ error(i) = (chi2_dist(gen) - df) * scale;
+ }
+ } else {
+ // Normal (default)
+ for (int i = 0; i < n; ++i) {
+ error(i) = normal(gen);
+ }
+ }
+
+ // Empirical re-standardization to SD = 1
+ double empirical_sd = std::sqrt(error.array().square().mean());
+ if (empirical_sd > FLOAT_NEAR_ZERO) {
+ error /= empirical_sd;
}
// Apply heteroskedasticity
diff --git a/cpp/src/ols.hpp b/cpp/src/ols.hpp
index ad1f9b6..9e046eb 100644
--- a/cpp/src/ols.hpp
+++ b/cpp/src/ols.hpp
@@ -65,6 +65,8 @@ class OLSAnalyzer {
* @param heterogeneity SD of effect size variation
* @param heteroskedasticity Correlation between predictor and error variance
* @param seed Random seed (-1 for random)
+ * @param residual_dist Error distribution: 0=normal, 1=heavy_tailed (t), 2=skewed (chi2)
+ * @param residual_df Degrees of freedom for non-normal residuals (min clamped to 3)
* @return Response vector (n_samples,)
*/
Eigen::VectorXd generate_y(
@@ -72,7 +74,9 @@ Eigen::VectorXd generate_y(
const Eigen::Ref& effects,
double heterogeneity,
double heteroskedasticity,
- int seed
+ int seed,
+ int residual_dist = 0,
+ double residual_df = 10.0
);
} // namespace mcpower
diff --git a/docs/screenshots/gui-model-setup.png b/docs/screenshots/gui-model-setup.png
new file mode 100644
index 0000000..7f87a53
Binary files /dev/null and b/docs/screenshots/gui-model-setup.png differ
diff --git a/docs/screenshots/gui-results.png b/docs/screenshots/gui-results.png
new file mode 100644
index 0000000..f84152d
Binary files /dev/null and b/docs/screenshots/gui-results.png differ
diff --git a/mcpower/__init__.py b/mcpower/__init__.py
index a52560c..675a4f3 100644
--- a/mcpower/__init__.py
+++ b/mcpower/__init__.py
@@ -16,7 +16,6 @@
from importlib.metadata import version as _get_version
-from .backends import get_backend_info, set_backend
from .model import MCPower
from .progress import PrintReporter, ProgressReporter, SimulationCancelled, TqdmReporter
@@ -27,14 +26,14 @@
__all__ = [
"MCPower",
"SimulationCancelled",
- "set_backend",
- "get_backend_info",
"ProgressReporter",
"PrintReporter",
"TqdmReporter",
]
+import threading as _threading
+
from .utils.updates import _check_for_updates
-_check_for_updates(__version__)
+_threading.Thread(target=_check_for_updates, args=(__version__,), daemon=True).start()
diff --git a/mcpower/backends/__init__.py b/mcpower/backends/__init__.py
index 7bb03f8..8b24a73 100644
--- a/mcpower/backends/__init__.py
+++ b/mcpower/backends/__init__.py
@@ -3,11 +3,9 @@
This module provides a unified interface for compute backends.
The only supported backend is native C++ (compiled via pybind11).
-
-Users can override via set_backend('c++' | 'default') or pass a ComputeBackend instance.
"""
-from typing import Optional, Protocol, Union, runtime_checkable
+from typing import Optional, Protocol, runtime_checkable
import numpy as np
@@ -24,6 +22,7 @@ def ols_analysis(
f_crit: float,
t_crit: float,
correction_t_crits: np.ndarray,
+ # correction_method encoding: 0=none, 1=Bonferroni, 2=FDR (BH), 3=Holm
correction_method: int,
) -> np.ndarray:
"""Run OLS regression and return significance flags.
@@ -40,9 +39,15 @@ def generate_y(
heterogeneity: float,
heteroskedasticity: float,
seed: int,
+ residual_dist: int = 0,
+ residual_df: float = 10.0,
) -> np.ndarray:
"""Generate the dependent variable ``y = X @ effects + error``.
+ Args:
+ residual_dist: Error distribution (0=normal, 1=heavy_tailed, 2=skewed).
+ residual_df: Degrees of freedom for non-normal residuals.
+
Returns:
1-D array of length ``n_samples``.
"""
@@ -88,12 +93,8 @@ def lme_analysis(
...
-# Valid backend names for set_backend()
-_BACKEND_NAMES = {"default", "c++"}
-
# Global backend instance
_backend_instance: Optional[ComputeBackend] = None
-_backend_forced = False
def get_backend() -> ComputeBackend:
@@ -101,7 +102,7 @@ def get_backend() -> ComputeBackend:
Get the active compute backend.
On first call, instantiates the C++ native backend.
- Subsequent calls return the cached instance unless reset_backend() is called.
+ Subsequent calls return the cached instance.
Raises:
ImportError: If the C++ extension is not compiled/installed.
@@ -117,64 +118,7 @@ def get_backend() -> ComputeBackend:
return _backend_instance
-def set_backend(backend: Union[str, ComputeBackend]) -> None:
- """
- Set the compute backend.
-
- Args:
- backend: One of:
- - 'default' -- use native C++ backend
- - 'c++' -- force native C++ backend
- - A ComputeBackend instance
-
- Raises:
- ImportError: If the C++ backend is not available.
- ValueError: If the string is not recognized.
- """
- global _backend_instance, _backend_forced
-
- if isinstance(backend, str):
- name = backend.lower().strip()
- if name not in _BACKEND_NAMES:
- raise ValueError(f"Unknown backend {backend!r}. Choose from: {', '.join(sorted(_BACKEND_NAMES))}")
-
- from .native import NativeBackend
-
- _backend_instance = NativeBackend()
- _backend_forced = name != "default"
- else:
- _backend_instance = backend
- _backend_forced = True
-
-
-def reset_backend() -> None:
- """Reset backend to automatic selection."""
- global _backend_instance, _backend_forced
- _backend_instance = None
- _backend_forced = False
-
-
-def get_backend_info() -> dict:
- """
- Get information about the current backend.
-
- Returns:
- Dictionary with backend name, type, and whether it was forced.
- """
- backend = get_backend()
- name = type(backend).__name__
- return {
- "name": name,
- "is_native": name == "NativeBackend",
- "module": type(backend).__module__,
- "forced": _backend_forced,
- }
-
-
__all__ = [
"ComputeBackend",
"get_backend",
- "set_backend",
- "reset_backend",
- "get_backend_info",
]
diff --git a/mcpower/backends/native.py b/mcpower/backends/native.py
index acd7633..2338cfe 100644
--- a/mcpower/backends/native.py
+++ b/mcpower/backends/native.py
@@ -17,6 +17,11 @@
mcpower_native = None
+def _prep(arr: np.ndarray, dtype=np.float64) -> np.ndarray:
+ """Ensure array is contiguous with the expected dtype for C++ interop."""
+ return np.ascontiguousarray(arr, dtype=dtype)
+
+
class NativeBackend:
"""
C++ compute backend using pybind11 bindings.
@@ -46,8 +51,8 @@ def _initialize_tables(self) -> None:
t3_ppf = manager.load_t3_ppf_table()
# Ensure correct dtypes
- norm_cdf = np.ascontiguousarray(norm_cdf.astype(np.float64))
- t3_ppf = np.ascontiguousarray(t3_ppf.astype(np.float64))
+ norm_cdf = _prep(norm_cdf)
+ t3_ppf = _prep(t3_ppf)
# Initialize C++ tables (generation tables only)
mcpower_native.init_tables(norm_cdf, t3_ppf)
@@ -77,10 +82,10 @@ def ols_analysis(
Returns:
Array: [f_sig, uncorrected..., corrected...]
"""
- X = np.ascontiguousarray(X, dtype=np.float64)
- y = np.ascontiguousarray(y, dtype=np.float64)
- target_indices = np.ascontiguousarray(target_indices, dtype=np.int32)
- correction_t_crits = np.ascontiguousarray(correction_t_crits, dtype=np.float64)
+ X = _prep(X)
+ y = _prep(y)
+ target_indices = _prep(target_indices, np.int32)
+ correction_t_crits = _prep(correction_t_crits)
return mcpower_native.ols_analysis(X, y, target_indices, f_crit, t_crit, correction_t_crits, correction_method) # type: ignore[no-any-return]
@@ -91,6 +96,8 @@ def generate_y(
heterogeneity: float,
heteroskedasticity: float,
seed: int,
+ residual_dist: int = 0,
+ residual_df: float = 10.0,
) -> np.ndarray:
"""
Generate dependent variable.
@@ -101,14 +108,16 @@ def generate_y(
heterogeneity: Effect size variation SD
heteroskedasticity: Error-predictor correlation
seed: Random seed (-1 for random)
+ residual_dist: Error distribution (0=normal, 1=heavy_tailed, 2=skewed)
+ residual_df: Degrees of freedom for non-normal residuals
Returns:
Response vector (n_samples,)
"""
- X = np.ascontiguousarray(X, dtype=np.float64)
- effects = np.ascontiguousarray(effects, dtype=np.float64)
+ X = _prep(X)
+ effects = _prep(effects)
- return mcpower_native.generate_y(X, effects, heterogeneity, heteroskedasticity, seed) # type: ignore[no-any-return]
+ return mcpower_native.generate_y(X, effects, heterogeneity, heteroskedasticity, seed, residual_dist, residual_df) # type: ignore[no-any-return]
def generate_X(
self,
@@ -137,11 +146,11 @@ def generate_X(
Returns:
Design matrix (n_samples, n_vars)
"""
- correlation_matrix = np.ascontiguousarray(correlation_matrix, dtype=np.float64)
- var_types = np.ascontiguousarray(var_types, dtype=np.int32)
- var_params = np.ascontiguousarray(var_params, dtype=np.float64)
- upload_normal = np.ascontiguousarray(upload_normal, dtype=np.float64)
- upload_data = np.ascontiguousarray(upload_data, dtype=np.float64)
+ correlation_matrix = _prep(correlation_matrix)
+ var_types = _prep(var_types, np.int32)
+ var_params = _prep(var_params)
+ upload_normal = _prep(upload_normal)
+ upload_data = _prep(upload_data)
return mcpower_native.generate_X( # type: ignore[no-any-return]
n_samples,
@@ -185,12 +194,15 @@ def lme_analysis(
Returns:
Array: [f_sig, uncorrected..., corrected..., wald_flag]
or empty array on failure
+
+ wald_flag: 1.0 if the Wald test was used as fallback for the overall
+ significance test (instead of the likelihood ratio test), 0.0 otherwise.
"""
- X = np.ascontiguousarray(X, dtype=np.float64)
- y = np.ascontiguousarray(y, dtype=np.float64)
- cluster_ids = np.ascontiguousarray(cluster_ids, dtype=np.int32)
- target_indices = np.ascontiguousarray(target_indices, dtype=np.int32)
- correction_z_crits = np.ascontiguousarray(correction_z_crits, dtype=np.float64)
+ X = _prep(X)
+ y = _prep(y)
+ cluster_ids = _prep(cluster_ids, np.int32)
+ target_indices = _prep(target_indices, np.int32)
+ correction_z_crits = _prep(correction_z_crits)
return mcpower_native.lme_analysis( # type: ignore[no-any-return]
X,
@@ -240,14 +252,17 @@ def lme_analysis_general(
Returns:
Array: [f_sig, uncorrected..., corrected..., wald_flag]
or empty array on failure
+
+ wald_flag: 1.0 if the Wald test was used as fallback for the overall
+ significance test (instead of the likelihood ratio test), 0.0 otherwise.
"""
- X = np.ascontiguousarray(X, dtype=np.float64)
- y = np.ascontiguousarray(y, dtype=np.float64)
- Z = np.ascontiguousarray(Z, dtype=np.float64)
- cluster_ids = np.ascontiguousarray(cluster_ids, dtype=np.int32)
- target_indices = np.ascontiguousarray(target_indices, dtype=np.int32)
- correction_z_crits = np.ascontiguousarray(correction_z_crits, dtype=np.float64)
- warm_theta = np.ascontiguousarray(warm_theta, dtype=np.float64)
+ X = _prep(X)
+ y = _prep(y)
+ Z = _prep(Z)
+ cluster_ids = _prep(cluster_ids, np.int32)
+ target_indices = _prep(target_indices, np.int32)
+ correction_z_crits = _prep(correction_z_crits)
+ warm_theta = _prep(warm_theta)
return mcpower_native.lme_analysis_general( # type: ignore[no-any-return]
X,
@@ -301,15 +316,18 @@ def lme_analysis_nested(
Returns:
Array: [f_sig, uncorrected..., corrected..., wald_flag]
or empty array on failure
+
+ wald_flag: 1.0 if the Wald test was used as fallback for the overall
+ significance test (instead of the likelihood ratio test), 0.0 otherwise.
"""
- X = np.ascontiguousarray(X, dtype=np.float64)
- y = np.ascontiguousarray(y, dtype=np.float64)
- parent_ids = np.ascontiguousarray(parent_ids, dtype=np.int32)
- child_ids = np.ascontiguousarray(child_ids, dtype=np.int32)
- child_to_parent = np.ascontiguousarray(child_to_parent, dtype=np.int32)
- target_indices = np.ascontiguousarray(target_indices, dtype=np.int32)
- correction_z_crits = np.ascontiguousarray(correction_z_crits, dtype=np.float64)
- warm_theta = np.ascontiguousarray(warm_theta, dtype=np.float64)
+ X = _prep(X)
+ y = _prep(y)
+ parent_ids = _prep(parent_ids, np.int32)
+ child_ids = _prep(child_ids, np.int32)
+ child_to_parent = _prep(child_to_parent, np.int32)
+ target_indices = _prep(target_indices, np.int32)
+ correction_z_crits = _prep(correction_z_crits)
+ warm_theta = _prep(warm_theta)
return mcpower_native.lme_analysis_nested( # type: ignore[no-any-return]
X,
diff --git a/mcpower/core/results.py b/mcpower/core/results.py
index 45b178a..dfe6b77 100644
--- a/mcpower/core/results.py
+++ b/mcpower/core/results.py
@@ -54,6 +54,7 @@ def calculate_powers(
# Individual powers
individual_powers = {}
individual_powers_corrected = {}
+ non_overall_tests = [t for t in target_tests if t != "overall"]
for test in target_tests:
if test == "overall":
@@ -62,7 +63,6 @@ def calculate_powers(
individual_powers_corrected[test] = np.mean(results_corrected_array[:, 0]) * 100
else:
# Find position among non-'overall' tests and add 1 for F-test offset
- non_overall_tests = [t for t in target_tests if t != "overall"]
pos = non_overall_tests.index(test)
col_idx = pos + 1 # +1 because column 0 is F-test
individual_powers[test] = np.mean(results_array[:, col_idx]) * 100
diff --git a/mcpower/core/scenarios.py b/mcpower/core/scenarios.py
index 454f8e3..2d2dd01 100644
--- a/mcpower/core/scenarios.py
+++ b/mcpower/core/scenarios.py
@@ -13,35 +13,51 @@
from ..utils.visualization import _create_power_plot
# Default scenario configurations.
+# "optimistic" is the zero-perturbation baseline — also used as the default
+# scenario_config when scenarios=False and as a template for custom scenarios
+# (ensures all required keys exist).
# "realistic" introduces moderate assumption violations; "doomer" introduces
# severe violations. Each simulation iteration draws random perturbations
# from these parameters (correlation noise, distribution swaps, etc.).
DEFAULT_SCENARIO_CONFIG = {
+ "optimistic": {
+ "heterogeneity": 0.0,
+ "heteroskedasticity": 0.0,
+ "correlation_noise_sd": 0.0,
+ "distribution_change_prob": 0.0,
+ "new_distributions": ["right_skewed", "left_skewed", "uniform"],
+ # Mixed model perturbations (only consumed when cluster_specs present)
+ "random_effect_dist": "normal",
+ "random_effect_df": 5,
+ "icc_noise_sd": 0.0,
+ # Residual distribution perturbations (all model types)
+ "residual_dists": ["heavy_tailed", "skewed"],
+ "residual_change_prob": 0.0,
+ "residual_df": 10,
+ },
"realistic": {
"heterogeneity": 0.2,
- "heteroskedasticity": 0.1,
- "correlation_noise_sd": 0.2,
- "distribution_change_prob": 0.3,
+ "heteroskedasticity": 0.15,
+ "correlation_noise_sd": 0.15,
+ "distribution_change_prob": 0.5,
"new_distributions": ["right_skewed", "left_skewed", "uniform"],
- # LME-specific keys (only consumed when cluster_specs present)
"random_effect_dist": "heavy_tailed",
- "random_effect_df": 5,
+ "random_effect_df": 10,
"icc_noise_sd": 0.15,
- "residual_dist": "heavy_tailed",
- "residual_change_prob": 0.3,
- "residual_df": 10,
+ "residual_dists": ["heavy_tailed", "skewed"],
+ "residual_change_prob": 0.5,
+ "residual_df": 8,
},
"doomer": {
"heterogeneity": 0.4,
- "heteroskedasticity": 0.2,
- "correlation_noise_sd": 0.4,
- "distribution_change_prob": 0.6,
+ "heteroskedasticity": 0.35,
+ "correlation_noise_sd": 0.30,
+ "distribution_change_prob": 0.8,
"new_distributions": ["right_skewed", "left_skewed", "uniform"],
- # LME-specific keys (only consumed when cluster_specs present)
"random_effect_dist": "heavy_tailed",
- "random_effect_df": 3,
+ "random_effect_df": 5,
"icc_noise_sd": 0.30,
- "residual_dist": "heavy_tailed",
+ "residual_dists": ["heavy_tailed", "skewed"],
"residual_change_prob": 0.8,
"residual_df": 5,
},
@@ -111,15 +127,7 @@ def run_power_analysis(
if progress is not None:
progress.start()
- # Optimistic (user's original settings)
- results["optimistic"] = run_find_power_func(
- sample_size=sample_size,
- target_tests=target_tests,
- correction=correction,
- scenario_config=None,
- )
-
- # Realistic & Doomer scenarios
+ # Run all scenarios (optimistic is always present as zero-perturbation baseline)
for scenario_name, config in self.configs.items():
results[scenario_name] = run_find_power_func(
sample_size=sample_size,
@@ -175,15 +183,7 @@ def run_sample_size_analysis(
if progress is not None:
progress.start()
- # Optimistic
- results["optimistic"] = run_sample_size_func(
- sample_sizes=sample_sizes,
- target_tests=target_tests,
- correction=correction,
- scenario_config=None,
- )
-
- # Other scenarios
+ # Run all scenarios (optimistic is always present as zero-perturbation baseline)
for scenario_name, config in self.configs.items():
results[scenario_name] = run_sample_size_func(
sample_sizes=sample_sizes,
@@ -209,8 +209,9 @@ def run_sample_size_analysis(
def _create_scenario_plots(self, results: Dict) -> None:
"""Create visualizations for scenario analysis."""
scenarios = results["scenarios"]
- scenario_names = ["optimistic", "realistic", "doomer"]
- scenario_labels = ["Optimistic", "Realistic", "Doomer"]
+ # Derive scenario order from results: optimistic first, then config keys
+ scenario_names = ["optimistic"] + [k for k in scenarios if k != "optimistic"]
+ scenario_labels = [name.title() for name in scenario_names]
first_scenario = scenarios.get("optimistic", {})
if "results" not in first_scenario or "sample_sizes_tested" not in first_scenario["results"]:
@@ -286,7 +287,7 @@ def apply_lme_perturbations(
if icc_noise_sd == 0.0 and re_dist == "normal":
return None
- rng = np.random.RandomState(sim_seed + 5000 if sim_seed is not None else None)
+ rng = np.random.RandomState(sim_seed + 6 if sim_seed is not None else None)
# ICC jitter: multiplicative noise on tau_squared per grouping variable
tau_squared_multipliers: Dict[str, float] = {}
@@ -304,70 +305,6 @@ def apply_lme_perturbations(
}
-def apply_lme_residual_perturbations(
- y: np.ndarray,
- scenario_config: Dict,
- sim_seed: Optional[int],
-) -> np.ndarray:
- """Replace normal residuals with non-normal if coin flip succeeds.
-
- For each simulation, independently flips a coin (probability
- ``residual_change_prob``) to decide whether residuals are replaced.
- If activated, reproduces the original N(0,1) errors via the known
- seed, generates replacements from t(df) or shifted χ², and applies
- the correction ``y += (new_error - original_error)``.
-
- Args:
- y: Dependent variable array (modified in-place).
- scenario_config: Scenario parameters with residual keys.
- sim_seed: Random seed for reproducibility.
-
- Returns:
- The (possibly modified) dependent variable array.
- """
- residual_dist = scenario_config.get("residual_dist", "normal")
- residual_change_prob = scenario_config.get("residual_change_prob", 0.0)
- residual_df = scenario_config.get("residual_df", 10)
-
- if residual_dist == "normal" or residual_change_prob <= 0.0:
- return y
-
- rng = np.random.RandomState(sim_seed + 6000 if sim_seed is not None else None)
-
- # Coin flip: should this simulation have non-normal residuals?
- if rng.random() > residual_change_prob:
- return y
-
- n = len(y)
-
- # Reproduce the original N(0,1) errors using the same seed as generate_y
- # generate_y uses sim_seed + 2 for error generation
- original_rng = np.random.RandomState(sim_seed + 2 if sim_seed is not None else None)
- original_errors = original_rng.standard_normal(n)
-
- # Generate replacement errors
- replacement_rng = np.random.RandomState(sim_seed + 6001 if sim_seed is not None else None)
-
- if residual_dist == "heavy_tailed":
- # t(df) scaled to have variance 1
- df = max(residual_df, 3)
- raw = replacement_rng.standard_t(df, size=n)
- # t(df) has variance df/(df-2), scale to unit variance
- scale = 1.0 / np.sqrt(df / (df - 2))
- new_errors = raw * scale
- elif residual_dist == "skewed":
- # Shifted chi-squared: mean=0, variance=1
- df = max(residual_df, 3)
- raw = replacement_rng.chisquare(df, size=n)
- new_errors = (raw - df) / np.sqrt(2 * df)
- else:
- return y
-
- # Apply correction: swap out original errors for new ones
- y = y + (new_errors - original_errors)
- return y
-
-
def apply_per_simulation_perturbations(
correlation_matrix: np.ndarray,
var_types: np.ndarray,
@@ -393,19 +330,22 @@ def apply_per_simulation_perturbations(
if scenario_config is None:
return correlation_matrix, var_types
- rng = np.random.RandomState(sim_seed)
+ rng = np.random.RandomState(sim_seed + 5 if sim_seed is not None else None)
# Perturb correlation matrix
perturbed_corr = correlation_matrix
- if correlation_matrix is not None and scenario_config["correlation_noise_sd"] > 0:
+ if correlation_matrix is not None and scenario_config.get("correlation_noise_sd", 0) > 0:
perturbed_corr = correlation_matrix.copy()
noise = rng.normal(0, scenario_config["correlation_noise_sd"], correlation_matrix.shape)
noise = (noise + noise.T) / 2 # Keep symmetric
perturbed_corr += noise
+ # Clip off-diagonal correlations to [-0.8, 0.8] to prevent near-singular
+ # matrices that cause Cholesky decomposition failures in data generation.
perturbed_corr = np.clip(perturbed_corr, -0.8, 0.8)
np.fill_diagonal(perturbed_corr, 1.0)
- # Ensure positive semi-definiteness via eigenvalue clipping
+ # Nearest correlation matrix repair via spectral clipping: set negative
+ # eigenvalues to zero and reconstruct, then re-normalize to unit diagonal.
eigvals, eigvecs = np.linalg.eigh(perturbed_corr)
if np.any(eigvals < 0):
eigvals = np.maximum(eigvals, 0.0)
@@ -417,7 +357,7 @@ def apply_per_simulation_perturbations(
# Perturb variable types
perturbed_var_types = var_types.copy()
- if scenario_config["distribution_change_prob"] > 0:
+ if scenario_config.get("distribution_change_prob", 0) > 0:
type_mapping = {"right_skewed": 2, "left_skewed": 3, "uniform": 5}
new_type_codes = [type_mapping[distribution] for distribution in scenario_config["new_distributions"]]
diff --git a/mcpower/core/simulation.py b/mcpower/core/simulation.py
index 266a39e..2223324 100644
--- a/mcpower/core/simulation.py
+++ b/mcpower/core/simulation.py
@@ -61,7 +61,7 @@ def __init__(
Args:
n_simulations: Number of Monte Carlo iterations.
seed: Base random seed. Each iteration uses
- ``seed + 4 * sim_id``.
+ ``seed + 12 * sim_id``.
alpha: Significance level for hypothesis tests.
parallel: Parallel processing mode (unused inside the
runner itself; parallelism is handled at the
@@ -143,12 +143,19 @@ def run_power_simulations(
if metadata.cluster_specs:
from ..stats.lme_solver import compute_lme_critical_values
- n_fixed = len(metadata.target_indices)
- # n_fixed_effects = number of columns in X_expanded (excluding intercept)
- # This equals the total effect count minus cluster effects
- n_fixed_total = len(metadata.effect_sizes)
- if metadata.cluster_effect_indices:
- n_fixed_total -= len(metadata.cluster_effect_indices)
+ # Use test formula dimensions when subsetting with random effects
+ if metadata.test_column_indices is not None and metadata.test_has_random_effects:
+ if metadata.test_target_indices is None:
+ raise RuntimeError("test_target_indices must be set when test_column_indices is present")
+ n_fixed = len(metadata.test_target_indices)
+ n_fixed_total = metadata.test_effect_count
+ else:
+ n_fixed = len(metadata.target_indices)
+ # n_fixed_effects = number of columns in X_expanded (excluding intercept)
+ # This equals the total effect count minus cluster effects
+ n_fixed_total = len(metadata.effect_sizes)
+ if metadata.cluster_effect_indices:
+ n_fixed_total -= len(metadata.cluster_effect_indices)
chi2_crit, z_crit, correction_z_crits = compute_lme_critical_values(
self.alpha, n_fixed_total, n_fixed, metadata.correction_method
)
@@ -162,19 +169,18 @@ def run_power_simulations(
raise SimulationCancelled("Simulation cancelled by user")
- sim_seed = self.seed + 4 * sim_id if self.seed is not None else None
-
- # Apply perturbations if in scenario mode
- if scenario_config is not None and apply_perturbations_func is not None:
- perturbed_corr, perturbed_types = apply_perturbations_func(
- metadata.correlation_matrix,
- metadata.var_types,
- scenario_config,
- sim_seed,
- )
- else:
- perturbed_corr = metadata.correlation_matrix
- perturbed_types = metadata.var_types
+ sim_seed = self.seed + 12 * sim_id if self.seed is not None else None
+
+ # Apply per-simulation perturbations (correlation noise, distribution swaps)
+ # Zero-valued params in optimistic scenario are no-ops
+ if apply_perturbations_func is None:
+ raise RuntimeError("apply_perturbations_func must be provided")
+ perturbed_corr, perturbed_types = apply_perturbations_func(
+ metadata.correlation_matrix,
+ metadata.var_types,
+ scenario_config,
+ sim_seed,
+ )
result = self._single_simulation(
sim_id=sim_id,
@@ -326,7 +332,9 @@ def _single_simulation(
first_spec = next(iter(metadata.cluster_specs.values()))
sample_size = first_spec["n_clusters"] * first_spec["cluster_size"]
- # Check if strict mode with uploaded data
+ # Strict-mode bootstrap: resample whole rows from uploaded data to
+ # preserve exact inter-variable relationships, then generate y from
+ # the bootstrapped X. This bypasses the normal X-generation pipeline.
if metadata.preserve_correlation == "strict" and metadata.uploaded_raw_data is not None:
# Strict mode: bootstrap uploaded data + generate created variables separately
from ..stats.data_generation import bootstrap_uploaded_data
@@ -336,7 +344,7 @@ def _single_simulation(
sample_size,
metadata.uploaded_raw_data,
metadata.uploaded_var_metadata,
- sim_seed,
+ sim_seed + 3 if sim_seed is not None else None,
)
# Merge uploaded and created non-factor variables
@@ -367,7 +375,7 @@ def _single_simulation(
X_factors = X_uploaded_factors
else:
# Mixed: generate all factors, replace uploaded factor columns
- X_factors = _generate_factors(sample_size, metadata.factor_specs, sim_seed)
+ X_factors = _generate_factors(sample_size, metadata.factor_specs, sim_seed + 3 if sim_seed is not None else None)
# Overwrite uploaded factor dummy columns with bootstrapped data
if X_uploaded_factors.shape[1] > 0:
col_offset = 0
@@ -400,14 +408,14 @@ def _single_simulation(
X_non_factors = np.empty((sample_size, 0), dtype=float)
# Generate factor variables (as dummy variables)
- X_factors = _generate_factors(sample_size, metadata.factor_specs, sim_seed)
+ X_factors = _generate_factors(sample_size, metadata.factor_specs, sim_seed + 3 if sim_seed is not None else None)
# Compute LME perturbations (ICC jitter, non-normal RE dist)
lme_perturbations = None
- if metadata.cluster_specs and scenario_config is not None:
+ if metadata.cluster_specs:
from ..core.scenarios import apply_lme_perturbations
- lme_perturbations = apply_lme_perturbations(metadata.cluster_specs, scenario_config, sim_seed)
+ lme_perturbations = apply_lme_perturbations(metadata.cluster_specs, scenario_config or {}, sim_seed)
# Generate cluster random effects (independent of upload mode)
re_result = None # Phase 2: random effects result for slopes/nesting
@@ -448,6 +456,16 @@ def _single_simulation(
# Create extended design matrix with interactions (excludes cluster effects)
X_expanded = create_X_extended_func(X)
+ # Test formula column subsetting: use reduced design matrix for analysis
+ if metadata.test_column_indices is not None:
+ X_test = X_expanded[:, metadata.test_column_indices]
+ if metadata.test_target_indices is None:
+ raise RuntimeError("test_target_indices must be set when test_column_indices is present")
+ test_target_indices = metadata.test_target_indices
+ else:
+ X_test = X_expanded
+ test_target_indices = metadata.target_indices
+
# Split effect sizes: fixed effects vs cluster effects
# Use precomputed values (Phase 2 optimization)
if metadata.cluster_effect_indices:
@@ -457,6 +475,21 @@ def _single_simulation(
fixed_effect_sizes = metadata.fixed_effect_sizes_cached
cluster_effect_sizes = None
+ # Residual coin flip: decide whether this simulation uses non-normal errors
+ residual_dist = 0 # normal
+ residual_df = 10.0
+ residual_change_prob = scenario_config.get("residual_change_prob", 0.0) if scenario_config else 0.0
+ if residual_change_prob > 0:
+ if scenario_config is None:
+ raise RuntimeError("scenario_config must be provided when residual_change_prob > 0")
+ coin_rng = np.random.RandomState(sim_seed + 7 if sim_seed is not None else None)
+ if coin_rng.random() < residual_change_prob:
+ residual_dists = scenario_config.get("residual_dists", ["heavy_tailed", "skewed"])
+ picked = coin_rng.choice(residual_dists)
+ dist_map = {"heavy_tailed": 1, "skewed": 2}
+ residual_dist = dist_map.get(picked, 0)
+ residual_df = float(scenario_config.get("residual_df", 10))
+
# Generate dependent variable with fixed effects only
y = generate_y_func(
X_expanded=X_expanded,
@@ -464,6 +497,8 @@ def _single_simulation(
heterogeneity=metadata.heterogeneity,
heteroskedasticity=metadata.heteroskedasticity,
sim_seed=sim_seed,
+ residual_dist=residual_dist,
+ residual_df=residual_df,
)
# Add cluster random effects contribution
@@ -478,12 +513,6 @@ def _single_simulation(
if re_result is not None and not np.allclose(re_result.slope_contribution, 0):
y = y + re_result.slope_contribution
- # Apply LME residual perturbations (non-normal residuals)
- if metadata.cluster_specs and scenario_config is not None:
- from ..core.scenarios import apply_lme_residual_perturbations
-
- y = apply_lme_residual_perturbations(y, scenario_config, sim_seed)
-
# Determine cluster IDs for the solver
cluster_ids: Optional[np.ndarray]
if re_result is not None:
@@ -496,23 +525,25 @@ def _single_simulation(
cluster_ids = metadata.cluster_ids_template
# Route to correct analysis method
- if cluster_ids is not None:
+ # When test_formula specifies no random effects, use OLS even if generation has clusters
+ use_lme = cluster_ids is not None and not (metadata.test_column_indices is not None and not metadata.test_has_random_effects)
+ if use_lme:
# Mixed model path (LME)
from ..stats.mixed_models import _lme_analysis_wrapper
+ assert cluster_ids is not None # narrowed by use_lme guard above
lme_result = _lme_analysis_wrapper(
- X_expanded,
+ X_test,
y,
- metadata.target_indices,
+ test_target_indices,
cluster_ids,
- metadata.cluster_column_indices,
metadata.correction_method,
self.alpha,
backend="custom",
verbose=metadata.verbose,
- chi2_crit=getattr(metadata, "lme_chi2_crit", None),
- z_crit=getattr(metadata, "lme_z_crit", None),
- correction_z_crits=getattr(metadata, "lme_correction_z_crits", None),
+ chi2_crit=metadata.lme_chi2_crit,
+ z_crit=metadata.lme_z_crit,
+ correction_z_crits=metadata.lme_correction_z_crits,
re_result=re_result,
)
@@ -539,16 +570,20 @@ def _single_simulation(
else:
# Standard OLS path
results = analyze_func(
- X_expanded,
+ X_test,
y,
- metadata.target_indices,
+ test_target_indices,
self.alpha,
metadata.correction_method,
)
diagnostics = None
- # Extract results: [f_sig, uncorr..., corr..., (wald_flag)]
- n_targets = len(metadata.target_indices)
+ # Result array layout: [F_sig, uncorrected[n_targets], corrected[n_targets], wald_flag?]
+ # - F_sig (index 0): overall model F-test significance (1.0 or 0.0)
+ # - uncorrected[1..n]: per-target t-test significance without correction
+ # - corrected[n+1..2n]: per-target significance with multiple-comparison correction
+ # - wald_flag (optional, LME only): 1.0 if Wald test was used instead of LRT
+ n_targets = len(test_target_indices)
f_significant = bool(results[0])
uncorrected = results[1 : 1 + n_targets].astype(bool)
corrected = results[1 + n_targets : 1 + 2 * n_targets].astype(bool)
@@ -560,19 +595,19 @@ def _single_simulation(
wald_flag = bool(results[expected_len])
# Post-hoc pairwise contrasts (OLS path only)
- if metadata.posthoc_specs and cluster_ids is None:
+ if metadata.posthoc_specs and not use_lme:
from ..stats.ols import compute_posthoc_contrasts
ph_uncorr, ph_corr, regular_override = compute_posthoc_contrasts(
- X_expanded,
+ X_test,
y,
metadata.posthoc_specs,
metadata.posthoc_method,
metadata.posthoc_t_crit,
metadata.posthoc_tukey_crits,
- target_indices=metadata.target_indices,
+ target_indices=test_target_indices,
correction_method=metadata.correction_method,
- correction_t_crits_combined=getattr(metadata, "posthoc_correction_t_crits_combined", None),
+ correction_t_crits_combined=metadata.posthoc_correction_t_crits_combined,
)
# If FDR/Holm combined correction was applied, override regular corrected
@@ -645,7 +680,7 @@ class SimulationMetadata:
correction_method: Encoded multiple-comparison correction
(0=none, 1=Bonferroni, 2=BH, 3=Holm).
heterogeneity: SD of random effect-size multiplier.
- heteroskedasticity: Correlation between first predictor and error SD.
+ heteroskedasticity: Correlation between predicted values and error SD.
preserve_correlation: Upload correlation mode
(``"no"``/``"partial"``/``"strict"``).
uploaded_raw_data: Normalised raw data for strict-mode bootstrap.
@@ -728,6 +763,13 @@ def __init__(
self.posthoc_method: str = "t-test"
self.posthoc_tukey_crits: Dict[str, float] = {}
self.posthoc_t_crit: float = 0.0
+ self.posthoc_correction_t_crits_combined: Optional[np.ndarray] = None
+
+ # Test formula fields (for model misspecification testing)
+ self.test_column_indices: Optional[np.ndarray] = None
+ self.test_target_indices: Optional[np.ndarray] = None
+ self.test_effect_count: Optional[int] = None # p for critical value computation
+ self.test_has_random_effects: bool = False # Whether test formula has (1|group) etc.
def _compute_fixed_effect_variance(registry) -> float:
@@ -779,8 +821,13 @@ def _compute_fixed_effect_variance(registry) -> float:
factor_info = registry._factors[factor_name]
proportions = factor_info.get("proportions")
if proportions is not None:
- # level is 1-indexed; proportions list is 0-indexed
- p_k = proportions[level - 1]
+ level_labels = factor_info.get("level_labels")
+ if level_labels is not None:
+ # String level labels — look up position by label
+ p_k = proportions[level_labels.index(str(level))]
+ else:
+ # Integer levels are 1-indexed; proportions list is 0-indexed
+ p_k = proportions[level - 1]
else:
# Equal proportions (default)
n_levels = factor_info["n_levels"]
@@ -814,6 +861,7 @@ def prepare_metadata(
model,
target_tests: List[str],
correction: Optional[str] = None,
+ test_formula_effects: Optional[List[str]] = None,
) -> SimulationMetadata:
"""
Prepare simulation metadata from model state.
@@ -825,6 +873,9 @@ def prepare_metadata(
model: MCPowerModel instance
target_tests: List of effects to test
correction: Multiple comparison correction method
+ test_formula_effects: Optional list of effect names from a test
+ formula. When provided, the metadata will include column
+ indices for subsetting X_expanded to the test model.
Returns:
SimulationMetadata instance
@@ -960,8 +1011,6 @@ def prepare_metadata(
upload_data_values=model.upload_data_values if model.upload_data_values is not None else np.zeros((2, 2), dtype=np.float64),
effect_sizes=effect_sizes,
correction_method=correction_method,
- heterogeneity=model.heterogeneity,
- heteroskedasticity=model.heteroskedasticity,
preserve_correlation=model._preserve_correlation,
uploaded_raw_data=model._uploaded_raw_data,
uploaded_var_metadata=model._uploaded_var_metadata,
@@ -982,4 +1031,20 @@ def prepare_metadata(
metadata.posthoc_specs = model._posthoc_specs
metadata.posthoc_method = "tukey" if is_tukey_correction else "t-test"
+ # Test formula column subsetting
+ if test_formula_effects is not None:
+ from ..utils.test_formula_utils import _compute_test_column_indices, _remap_target_indices
+
+ # Get all non-cluster effect names in registry order
+ all_effect_names = [name for name in registry._effects if name not in registry.cluster_effect_names]
+
+ test_col_indices = _compute_test_column_indices(all_effect_names, test_formula_effects)
+ metadata.test_column_indices = test_col_indices
+ metadata.test_effect_count = len(test_col_indices)
+
+ # Remap target indices to X_test space
+ # Only remap targets that exist in the test formula
+ valid_targets = np.array([idx for idx in target_indices if idx in test_col_indices], dtype=np.int64)
+ metadata.test_target_indices = _remap_target_indices(valid_targets, test_col_indices)
+
return metadata
diff --git a/mcpower/core/variables.py b/mcpower/core/variables.py
index e311606..338d40b 100644
--- a/mcpower/core/variables.py
+++ b/mcpower/core/variables.py
@@ -367,76 +367,42 @@ def expand_factors(self) -> None:
level_labels = factor_info.get("level_labels")
reference_level = factor_info.get("reference_level", 1)
+ # Compute non-reference levels once
if level_labels is not None:
- # Named levels: skip the reference, create dummies for the rest
- non_ref_labels = [lb for lb in level_labels if lb != str(reference_level)]
- for label in non_ref_labels:
- dummy_name = f"{factor_name}[{label}]"
-
- # Create dummy predictor
- dummy_pred = PredictorVar(
- name=dummy_name,
- var_type="factor_dummy",
- is_dummy=True,
- factor_source=factor_name,
- factor_level=label,
- column_index=col_idx,
- level_labels=level_labels,
- )
- new_predictors[dummy_name] = dummy_pred
-
- # Create main effect for dummy
- dummy_eff = Effect(
- name=dummy_name,
- effect_type="main",
- var_names=[dummy_name],
- column_index=col_idx,
- factor_source=factor_name,
- factor_level=label,
- )
- new_effects[dummy_name] = dummy_eff
-
- # Store dummy mapping
- self._factor_dummies[dummy_name] = {
- "factor_name": factor_name,
- "level": label,
- }
-
- col_idx += 1
+ non_ref = [lb for lb in level_labels if lb != str(reference_level)]
else:
- # Original integer-indexed behavior
- for level in range(2, n_levels + 1):
- dummy_name = f"{factor_name}[{level}]"
-
- # Create dummy predictor
- dummy_pred = PredictorVar(
- name=dummy_name,
- var_type="factor_dummy",
- is_dummy=True,
- factor_source=factor_name,
- factor_level=level,
- column_index=col_idx,
- )
- new_predictors[dummy_name] = dummy_pred
-
- # Create main effect for dummy
- dummy_eff = Effect(
- name=dummy_name,
- effect_type="main",
- var_names=[dummy_name],
- column_index=col_idx,
- factor_source=factor_name,
- factor_level=level,
- )
- new_effects[dummy_name] = dummy_eff
+ non_ref = list(range(2, n_levels + 1))
+
+ for level in non_ref:
+ dummy_name = f"{factor_name}[{level}]"
+
+ dummy_pred = PredictorVar(
+ name=dummy_name,
+ var_type="factor_dummy",
+ is_dummy=True,
+ factor_source=factor_name,
+ factor_level=level,
+ column_index=col_idx,
+ level_labels=level_labels if level_labels is not None else None,
+ )
+ new_predictors[dummy_name] = dummy_pred
+
+ dummy_eff = Effect(
+ name=dummy_name,
+ effect_type="main",
+ var_names=[dummy_name],
+ column_index=col_idx,
+ factor_source=factor_name,
+ factor_level=level,
+ )
+ new_effects[dummy_name] = dummy_eff
- # Store dummy mapping
- self._factor_dummies[dummy_name] = {
- "factor_name": factor_name,
- "level": level,
- }
+ self._factor_dummies[dummy_name] = {
+ "factor_name": factor_name,
+ "level": level,
+ }
- col_idx += 1
+ col_idx += 1
# Handle interactions involving factors — Cartesian product of
# non-reference dummy levels across all factor components.
@@ -503,6 +469,13 @@ def get_effect_sizes(self) -> np.ndarray:
def get_var_types(self) -> np.ndarray:
"""Get variable types as numpy array (for data generation)."""
+ # Type codes: 0-5 are parametric distributions generated from scratch.
+ # 97/98/99 are sentinel codes for uploaded-data variables whose values
+ # come from bootstrapped/quantile-matched empirical data rather than
+ # parametric generation:
+ # 97 = uploaded_factor (factor from uploaded data)
+ # 98 = uploaded_binary (binary from uploaded data)
+ # 99 = uploaded_data (continuous from uploaded data)
type_mapping = {
"normal": 0,
"binary": 1,
@@ -717,28 +690,20 @@ def register_cluster(
def _reindex_predictors(self) -> None:
"""Reindex all predictors to maintain order: non_factor | cluster_effect | dummies."""
- col_idx = 0
+ non_factor = []
+ cluster = []
+ dummies = []
- # Non-factor predictors first
- for name in sorted(self._predictors.keys(), key=lambda x: self._predictors[x].column_index or 0):
- pred = self._predictors[name]
- if not pred.is_factor and not pred.is_dummy and pred.var_type != "cluster_effect":
- pred.column_index = col_idx
- col_idx += 1
-
- # Cluster effect predictors second
- for name in sorted(self._predictors.keys(), key=lambda x: self._predictors[x].column_index or 0):
- pred = self._predictors[name]
- if pred.var_type == "cluster_effect":
- pred.column_index = col_idx
- col_idx += 1
-
- # Factor dummies last
for name in sorted(self._predictors.keys(), key=lambda x: self._predictors[x].column_index or 0):
pred = self._predictors[name]
if pred.is_dummy:
- pred.column_index = col_idx
- col_idx += 1
+ dummies.append(pred)
+ elif pred.var_type == "cluster_effect":
+ cluster.append(pred)
+ elif not pred.is_factor:
+ non_factor.append(pred)
+
+ for col_idx, pred in enumerate(non_factor + cluster + dummies):
+ pred.column_index = col_idx
- # Update effect indices
self._update_effect_indices()
diff --git a/mcpower/model.py b/mcpower/model.py
index 4fa2c4b..9a5f813 100644
--- a/mcpower/model.py
+++ b/mcpower/model.py
@@ -123,10 +123,9 @@ def __init__(self, data_generation_formula: str):
self._pending_factor_levels: Optional[str] = None
self._pending_effects: Optional[str] = None
self._pending_correlations: Optional[Union[str, np.ndarray]] = None
- self._pending_heterogeneity: Optional[float] = None
- self._pending_heteroskedasticity: Optional[float] = None
self._pending_data: Optional[Dict[str, Any]] = None
self._pending_clusters: Dict[str, Dict] = {} # {grouping_var: {n_clusters, cluster_size, icc}}
+ self._effects_set: bool = False # True after set_effects() has been called
# Detect mixed model formula
if self._registry._random_effects_parsed:
@@ -134,8 +133,6 @@ def __init__(self, data_generation_formula: str):
# Applied state
self._applied = False
- self.heterogeneity = 0.0
- self.heteroskedasticity = 0.0
# Data storage
self.upload_normal_values: Optional[np.ndarray] = None
@@ -385,6 +382,7 @@ def set_effects(self, effects_string: str):
raise ValueError("effects_string cannot be empty")
self._pending_effects = effects_string
+ self._effects_set = True
self._applied = False
return self
@@ -432,13 +430,16 @@ def set_variable_type(self, variable_types_string: str):
- ``"normal"`` — standard normal (default).
- ``"binary"`` or ``"binary(p)"`` — Bernoulli with proportion *p*
(default 0.5).
- - ``"skewed"`` — heavy-tailed (t-distribution, df=3).
+ - ``"right_skewed"`` — positively skewed distribution.
+ - ``"left_skewed"`` — negatively skewed distribution.
+ - ``"high_kurtosis"`` — heavy-tailed (t-distribution, df=3).
+ - ``"uniform"`` — uniform distribution.
- ``"factor(k)"`` — categorical with *k* levels (creates *k-1*
dummy variables).
- ``"factor(k, p1, p2, ...)"`` — factor with custom level
proportions.
- Example: ``"x1=binary, x2=skewed, x3=factor(3)"``.
+ Example: ``"x1=binary, x2=right_skewed, x3=factor(3)"``.
Returns:
self: For method chaining.
@@ -479,62 +480,6 @@ def set_factor_levels(self, spec: str):
self._applied = False
return self
- def set_heterogeneity(self, heterogeneity: float):
- """Set heterogeneity (random variation) in effect sizes.
-
- When non-zero, each simulation draws a per-simulation effect-size
- multiplier from a normal distribution with mean 1 and the given
- standard deviation. This models uncertainty about the true effect
- size — for example, ``heterogeneity=0.1`` means effect sizes vary
- by roughly +/- 10% across simulations.
-
- This setting is deferred until ``apply()`` is called.
-
- Args:
- heterogeneity: Standard deviation of the random effect-size
- multiplier. Must be non-negative. Default is 0 (no variation).
-
- Returns:
- self: For method chaining.
-
- Raises:
- TypeError: If *heterogeneity* is not numeric.
- """
- if not isinstance(heterogeneity, (int, float)):
- raise TypeError("heterogeneity must be a number")
-
- self._pending_heterogeneity = float(heterogeneity)
- self._applied = False
- return self
-
- def set_heteroskedasticity(self, heteroskedasticity_correlation: float):
- """Set heteroskedasticity (non-constant error variance).
-
- Introduces a correlation between the first predictor's values and
- the error standard deviation, producing variance that increases (or
- decreases) with the predictor. This violates the homoskedasticity
- assumption and typically reduces power.
-
- This setting is deferred until ``apply()`` is called.
-
- Args:
- heteroskedasticity_correlation: Correlation between the first
- predictor and the error standard deviation, in the range
- [-1, 1]. Default is 0 (homoskedastic errors).
-
- Returns:
- self: For method chaining.
-
- Raises:
- TypeError: If the value is not numeric.
- """
- if not isinstance(heteroskedasticity_correlation, (int, float)):
- raise TypeError("heteroskedasticity_correlation must be a number")
-
- self._pending_heteroskedasticity = float(heteroskedasticity_correlation)
- self._applied = False
- return self
-
def set_cluster(
self,
grouping_var: str,
@@ -769,6 +714,7 @@ def upload_data(
"preserve_factor_level_names": preserve_factor_level_names,
}
self._applied = False
+ return self
def set_scenario_configs(self, configs_dict: Dict):
"""Set custom scenario configurations for robustness analysis.
@@ -786,7 +732,9 @@ def set_scenario_configs(self, configs_dict: Dict):
configs_dict: Mapping of scenario names to configuration dicts.
Each configuration may include keys such as
``"heterogeneity"``, ``"heteroskedasticity"``,
- ``"effect_size_jitter"``, and ``"distribution_jitter"``.
+ ``"correlation_noise_sd"``, and ``"distribution_change_prob"``.
+ See ``DEFAULT_SCENARIO_CONFIG`` in ``mcpower.core.scenarios``
+ for the full list of keys.
Returns:
self: For method chaining.
@@ -802,7 +750,8 @@ def set_scenario_configs(self, configs_dict: Dict):
if scenario in merged:
merged[scenario].update(config)
else:
- merged[scenario] = config
+ # New custom scenarios inherit all keys from optimistic baseline
+ merged[scenario] = {**DEFAULT_SCENARIO_CONFIG["optimistic"], **config}
self._scenario_configs = merged
print(f"Custom scenario configs set: {', '.join(configs_dict.keys())}")
@@ -812,7 +761,7 @@ def set_scenario_configs(self, configs_dict: Dict):
# Apply method (processes all pending settings)
# =========================================================================
- def apply(self):
+ def _apply(self):
"""
Apply all pending settings to the model.
@@ -857,16 +806,22 @@ def apply(self):
# 7. Apply correlations
self._apply_correlations(_parser)
- # 8. Apply heterogeneity/heteroskedasticity
- self._apply_heterogeneity()
-
- # 9. Validate model is ready
+ # 8. Validate model is ready
model_result = _validate_model_ready(self)
model_result.raise_if_invalid()
- # Invalidate effect plan cache when settings change (Phase 2 optimization)
+ # Invalidate the effect plan cache — apply() rebuilds the variable
+ # registry state, so any cached column mappings are now stale.
self._effect_plan_cache = None
+ # Clear pending state to prevent double-application
+ self._pending_variable_types = None
+ self._pending_factor_levels = None
+ self._pending_effects = None
+ self._pending_correlations = None
+ self._pending_data = None
+ self._pending_clusters = {}
+
self._applied = True
print("Model settings applied successfully")
return self
@@ -1024,6 +979,21 @@ def _apply_data(self):
# Extract matched data
matched_data = data[:, matched_indices]
+ # Reject NaN values early
+ try:
+ if np.isnan(matched_data.astype(np.float64)).any():
+ nan_cols = [
+ matched_columns[i] for i in range(matched_data.shape[1]) if np.isnan(matched_data[:, i].astype(np.float64)).any()
+ ]
+ raise ValueError(
+ f"Uploaded data contains NaN values in columns: {', '.join(nan_cols)}. "
+ f"Remove or impute missing values before uploading."
+ )
+ except (ValueError, TypeError):
+ # Object dtype columns (strings) can't be converted to float for NaN check.
+ # NaN check for numeric columns will happen after string encoding below.
+ pass
+
# Convert to float64 if object dtype (common with mixed-type DataFrames)
# String columns are encoded to integer indices; mapping is stored in string_col_indices
string_col_indices = {}
@@ -1178,11 +1148,7 @@ def _apply_data_normal_mode(self, data, columns, type_info, mode, data_types_ove
level_labels = info.get("level_labels")
# Determine reference from data_types tuple override
- reference_level = None
- if col in data_types_override:
- dt = data_types_override[col]
- if isinstance(dt, tuple) and len(dt) == 2:
- reference_level = str(dt[1])
+ reference_level = self._extract_reference_level(data_types_override, col)
# Calculate proportions for each level
proportions = []
@@ -1200,7 +1166,10 @@ def _apply_data_normal_mode(self, data, columns, type_info, mode, data_types_ove
else: # continuous
# Normalize: mean=0, sd=1
- normalized = (col_data - np.mean(col_data)) / np.std(col_data, ddof=1)
+ std = np.std(col_data, ddof=1)
+ if std < 1e-15:
+ raise ValueError(f"Column '{col}' has zero variance (constant value). Remove it from the model or check your data.")
+ normalized = (col_data - np.mean(col_data)) / std
# Create lookup tables (type 99)
normal_vals, uploaded_vals = create_uploaded_lookup_tables(normalized.reshape(-1, 1))
@@ -1324,11 +1293,7 @@ def _apply_data_strict_mode(self, data, columns, type_info, data_types_override=
level_labels = info.get("level_labels")
# Determine reference from data_types tuple override
- reference_level = None
- if col in data_types_override:
- dt = data_types_override[col]
- if isinstance(dt, tuple) and len(dt) == 2:
- reference_level = str(dt[1])
+ reference_level = self._extract_reference_level(data_types_override, col)
self._uploaded_var_metadata[col] = {
"type": "factor",
@@ -1355,7 +1320,10 @@ def _apply_data_strict_mode(self, data, columns, type_info, data_types_override=
continuous_cols.append(idx)
# Normalize
col_data = data[:, idx]
- normalized_data[:, idx] = (col_data - np.mean(col_data)) / np.std(col_data, ddof=1)
+ std = np.std(col_data, ddof=1)
+ if std < 1e-15:
+ raise ValueError(f"Column '{col}' has zero variance (constant value). Remove it from the model or check your data.")
+ normalized_data[:, idx] = (col_data - np.mean(col_data)) / std
self._uploaded_var_metadata[col] = {
"type": "continuous",
@@ -1481,22 +1449,6 @@ def _apply_correlations(self, _parser):
self._registry.set_correlation_matrix(correlations_input)
print("Correlation matrix set")
- def _apply_heterogeneity(self):
- """Validate and apply pending heterogeneity and heteroskedasticity settings."""
- if self._pending_heterogeneity is not None:
- if self._pending_heterogeneity < 0:
- raise ValueError("heterogeneity must be non-negative")
- self.heterogeneity = self._pending_heterogeneity
- if self.heterogeneity > 0:
- print(f"Heterogeneity: SD = {self.heterogeneity}")
-
- if self._pending_heteroskedasticity is not None:
- if not -1 <= self._pending_heteroskedasticity <= 1:
- raise ValueError("heteroskedasticity_correlation must be between -1 and 1")
- self.heteroskedasticity = self._pending_heteroskedasticity
- if abs(self.heteroskedasticity) > 1e-8:
- print(f"Heteroskedasticity: correlation = {self.heteroskedasticity}")
-
# =========================================================================
# Analysis methods
# =========================================================================
@@ -1507,7 +1459,7 @@ def find_power(
target_test: str = "all",
correction: Optional[str] = None,
print_results: bool = True,
- scenarios: bool = False,
+ scenarios: Union[bool, List[str]] = False,
summary: str = "short",
return_results: bool = False,
test_formula: str = "",
@@ -1529,12 +1481,16 @@ def find_power(
Duplicate tests raise ``ValueError``.
correction: Multiple comparison correction (None, "bonferroni", "benjamini-hochberg", "holm")
print_results: Whether to print results
- scenarios: Run scenario analysis
+ scenarios: Scenario analysis control:
+ - ``False`` (default): no scenario analysis.
+ - ``True``: run all configured scenarios.
+ - List of scenario names: run only the specified scenarios
+ (e.g. ``["optimistic", "extreme"]``). Case-insensitive.
summary: Output detail level ("short" or "long")
return_results: Return results dict
test_formula: Formula for statistical testing (default: use data generation formula).
If the formula contains random effects like (1|school), analysis switches to
- mixed model testing (not yet implemented).
+ mixed model testing.
progress_callback: Progress reporting control:
- ``None`` (default): auto-use ``PrintReporter`` when
*print_results* is ``True``.
@@ -1549,7 +1505,10 @@ def find_power(
"""
# Auto-apply if settings have changed
if not self._applied:
- self.apply()
+ self._apply()
+
+ # Resolve scenarios parameter
+ scenario_filter = self._resolve_scenarios(scenarios)
# Validate sample size (basic: >= 20, type check)
_validate_sample_size(sample_size).raise_if_invalid()
@@ -1558,9 +1517,6 @@ def find_power(
n_variables = len(self._registry.effect_names)
_validate_sample_size_for_model(sample_size, n_variables).raise_if_invalid()
- # Validate and adjust cluster sample sizes
- self._validate_cluster_sample_size(sample_size)
-
# Warn if sample size is much larger than uploaded data
if self._uploaded_data_n > 0 and sample_size > 3 * self._uploaded_data_n:
print(
@@ -1570,33 +1526,13 @@ def find_power(
)
self._validate_analysis_inputs(correction)
- resolved_test_formula = self._resolve_test_formula(test_formula)
- target_tests = self._parse_target_tests(target_test)
-
- if correction and correction.lower() == "tukey" and not self._posthoc_specs:
- raise ValueError(
- "Tukey correction requires at least one post-hoc comparison "
- "(e.g., target_test='group[0] vs group[1]'). "
- "Tukey HSD only applies to pairwise contrasts between factor levels."
- )
-
- # Resolve progress callback
- from .progress import PrintReporter, ProgressReporter, compute_total_simulations
-
- if progress_callback is None:
- effective_cb = PrintReporter() if print_results else None
- elif progress_callback is False:
- effective_cb = None
- else:
- effective_cb = progress_callback
+ resolved_test_formula, test_formula_effects, test_random_effects = self._resolve_test_formula(test_formula)
+ target_tests = self._parse_target_tests(target_test, test_formula_effects=test_formula_effects)
+ self._validate_tukey_posthoc(correction)
- reporter = None
- if effective_cb is not None:
- n_scenarios = (len(self._scenario_configs or DEFAULT_SCENARIO_CONFIG) + 1) if scenarios else 1
- total = compute_total_simulations(self._effective_n_simulations, 1, n_scenarios)
- reporter = ProgressReporter(total, effective_cb)
+ reporter = self._resolve_progress(progress_callback, print_results, scenario_filter)
- if scenarios:
+ if scenario_filter is not None:
result = self._run_scenario_analysis(
"power",
sample_size=sample_size,
@@ -1605,8 +1541,11 @@ def find_power(
summary=summary,
print_results=print_results,
test_formula=resolved_test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=reporter,
cancel_check=cancel_check,
+ scenario_filter=scenario_filter,
)
else:
if reporter is not None:
@@ -1615,7 +1554,10 @@ def find_power(
sample_size,
target_tests,
correction,
+ scenario_config=DEFAULT_SCENARIO_CONFIG["optimistic"],
test_formula=resolved_test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=reporter,
cancel_check=cancel_check,
)
@@ -1623,7 +1565,7 @@ def find_power(
if reporter is not None:
reporter.finish()
- if not scenarios and print_results:
+ if scenario_filter is None and print_results:
print(f"\n{'=' * 80}")
print("MONTE CARLO POWER ANALYSIS RESULTS")
print(f"{'=' * 80}")
@@ -1641,7 +1583,7 @@ def find_sample_size(
by: int = 5,
correction: Optional[str] = None,
print_results: bool = True,
- scenarios: bool = False,
+ scenarios: Union[bool, List[str]] = False,
summary: str = "short",
return_results: bool = False,
test_formula: str = "",
@@ -1659,12 +1601,16 @@ def find_sample_size(
by: Step size between sample sizes
correction: Multiple comparison correction
print_results: Whether to print results
- scenarios: Run scenario analysis
+ scenarios: Scenario analysis control:
+ - ``False`` (default): no scenario analysis.
+ - ``True``: run all configured scenarios.
+ - List of scenario names: run only the specified scenarios
+ (e.g. ``["optimistic", "extreme"]``). Case-insensitive.
summary: Output detail level
return_results: Return results dict
test_formula: Formula for statistical testing (default: use data generation formula).
If the formula contains random effects like (1|school), analysis switches to
- mixed model testing (not yet implemented).
+ mixed model testing.
progress_callback: Progress reporting control:
- ``None`` (default): auto-use ``PrintReporter`` when
*print_results* is ``True``.
@@ -1680,7 +1626,10 @@ def find_sample_size(
"""
# Auto-apply if settings have changed
if not self._applied:
- self.apply()
+ self._apply()
+
+ # Resolve scenarios parameter
+ scenario_filter = self._resolve_scenarios(scenarios)
# Validate from_size meets minimum requirements
_validate_sample_size(from_size).raise_if_invalid()
@@ -1696,40 +1645,20 @@ def find_sample_size(
)
self._validate_analysis_inputs(correction)
- resolved_test_formula = self._resolve_test_formula(test_formula)
+ resolved_test_formula, test_formula_effects, test_random_effects = self._resolve_test_formula(test_formula)
validation_result = _validate_sample_size_range(from_size, to_size, by)
for warning in validation_result.warnings:
print(f"Warning: {warning}")
validation_result.raise_if_invalid()
- target_tests = self._parse_target_tests(target_test)
-
- if correction and correction.lower() == "tukey" and not self._posthoc_specs:
- raise ValueError(
- "Tukey correction requires at least one post-hoc comparison "
- "(e.g., target_test='group[0] vs group[1]'). "
- "Tukey HSD only applies to pairwise contrasts between factor levels."
- )
+ target_tests = self._parse_target_tests(target_test, test_formula_effects=test_formula_effects)
+ self._validate_tukey_posthoc(correction)
sample_sizes = list(range(from_size, to_size + 1, by))
- # Resolve progress callback
- from .progress import PrintReporter, ProgressReporter, compute_total_simulations
-
- if progress_callback is None:
- effective_cb = PrintReporter() if print_results else None
- elif progress_callback is False:
- effective_cb = None
- else:
- effective_cb = progress_callback
-
- reporter = None
- if effective_cb is not None:
- n_scenarios = (len(self._scenario_configs or DEFAULT_SCENARIO_CONFIG) + 1) if scenarios else 1
- total = compute_total_simulations(self._effective_n_simulations, len(sample_sizes), n_scenarios)
- reporter = ProgressReporter(total, effective_cb)
+ reporter = self._resolve_progress(progress_callback, print_results, scenario_filter, n_sample_sizes=len(sample_sizes))
- if scenarios:
+ if scenario_filter is not None:
result = self._run_scenario_analysis(
"sample_size",
target_tests=target_tests,
@@ -1738,8 +1667,11 @@ def find_sample_size(
summary=summary,
print_results=print_results,
test_formula=resolved_test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=reporter,
cancel_check=cancel_check,
+ scenario_filter=scenario_filter,
)
else:
if reporter is not None:
@@ -1748,7 +1680,10 @@ def find_sample_size(
sample_sizes,
target_tests,
correction,
+ scenario_config=DEFAULT_SCENARIO_CONFIG["optimistic"],
test_formula=resolved_test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=reporter,
cancel_check=cancel_check,
)
@@ -1756,7 +1691,7 @@ def find_sample_size(
if reporter is not None:
reporter.finish()
- if not scenarios and print_results:
+ if scenario_filter is None and print_results:
print(f"\n{'=' * 80}")
print("SAMPLE SIZE ANALYSIS RESULTS")
print(f"{'=' * 80}")
@@ -1780,6 +1715,8 @@ def _generate_dependent_variable(
heterogeneity: float = 0.0,
heteroskedasticity: float = 0.0,
sim_seed: Optional[int] = None,
+ residual_dist: int = 0,
+ residual_df: float = 10.0,
) -> np.ndarray:
"""Generate the dependent variable as y = X @ beta + error via the active backend."""
return get_backend().generate_y(
@@ -1788,19 +1725,103 @@ def _generate_dependent_variable(
heterogeneity,
heteroskedasticity,
sim_seed if sim_seed is not None else -1,
+ residual_dist,
+ residual_df,
)
# =========================================================================
# Internal methods
# =========================================================================
+ @staticmethod
+ def _extract_reference_level(data_types_override, col):
+ """Extract reference level from data_types_override tuple for a column."""
+ dt = data_types_override.get(col)
+ if isinstance(dt, tuple) and len(dt) == 2:
+ return str(dt[1])
+ return None
+
+ def _resolve_scenarios(self, scenarios: Union[bool, List[str]]) -> Optional[List[str]]:
+ """Resolve the scenarios parameter into a list of scenario names or None.
+
+ Args:
+ scenarios: ``False`` for no scenarios, ``True`` for all configured
+ scenarios, or a list of scenario names (case-insensitive).
+
+ Returns:
+ List of validated, lowercase scenario names, or ``None`` if
+ scenarios are disabled.
+
+ Raises:
+ ValueError: If any requested scenario name is not configured.
+ TypeError: If *scenarios* is not ``bool`` or a list of strings.
+ """
+ if scenarios is False:
+ return None
+
+ all_configs = self._scenario_configs or DEFAULT_SCENARIO_CONFIG
+ available = set(all_configs.keys())
+
+ if scenarios is True:
+ return list(all_configs.keys())
+
+ if not isinstance(scenarios, list):
+ raise TypeError(f"scenarios must be True, False, or a list of scenario names, got {type(scenarios).__name__}")
+
+ # Case-insensitive matching
+ available_lower = {k.lower(): k for k in available}
+ resolved = []
+ invalid = []
+ for name in scenarios:
+ if not isinstance(name, str):
+ raise TypeError(f"Scenario names must be strings, got {type(name).__name__}")
+ key = available_lower.get(name.lower())
+ if key is None:
+ invalid.append(name)
+ else:
+ resolved.append(key)
+
+ if invalid:
+ raise ValueError(f"Unknown scenario(s): {', '.join(repr(n) for n in invalid)}. Available: {', '.join(sorted(available))}")
+
+ return resolved
+
+ def _resolve_progress(self, progress_callback, print_results, scenario_filter, n_sample_sizes=1):
+ """Resolve progress_callback into a ProgressReporter or None."""
+ from .progress import PrintReporter, ProgressReporter, compute_total_simulations
+
+ if progress_callback is None:
+ effective_cb = PrintReporter() if print_results else None
+ elif progress_callback is False:
+ effective_cb = None
+ else:
+ effective_cb = progress_callback
+
+ if effective_cb is None:
+ return None
+
+ n_scenarios = len(scenario_filter) if scenario_filter is not None else 1
+ total = compute_total_simulations(self._effective_n_simulations, n_sample_sizes, n_scenarios)
+ return ProgressReporter(total, effective_cb)
+
def _validate_analysis_inputs(self, correction):
"""Validate the multiple-comparison correction method before analysis."""
result = _validate_correction_method(correction)
result.raise_if_invalid()
+ def _validate_tukey_posthoc(self, correction):
+ """Raise if Tukey correction is requested without posthoc specs."""
+ if correction and correction.lower() == "tukey" and not self._posthoc_specs:
+ raise ValueError(
+ "Tukey correction requires at least one post-hoc comparison "
+ "(e.g., target_test='group[0] vs group[1]'). "
+ "Tukey HSD only applies to pairwise contrasts between factor levels."
+ )
+
def _validate_cluster_sample_size(self, sample_size: int):
"""Derive missing cluster dimensions from sample_size and validate minimums."""
+ # NOTE: This method both validates AND mutates — it derives missing
+ # cluster_size/n_clusters from sample_size before checking minimums.
if not self._registry.cluster_names:
return # No clusters, nothing to do
@@ -1811,10 +1832,12 @@ def _validate_cluster_sample_size(self, sample_size: int):
if spec.n_clusters is not None:
spec.cluster_size = sample_size // spec.n_clusters
else:
- assert spec.cluster_size is not None
+ if spec.cluster_size is None:
+ raise RuntimeError(f"Cluster '{gv}': either n_clusters or cluster_size must be set")
spec.n_clusters = sample_size // spec.cluster_size
- assert spec.n_clusters is not None and spec.cluster_size is not None
+ if spec.n_clusters is None or spec.cluster_size is None:
+ raise RuntimeError(f"Cluster '{gv}': failed to derive n_clusters and cluster_size from sample_size={sample_size}")
actual_n = spec.n_clusters * spec.cluster_size
if actual_n != sample_size:
print(
@@ -1825,7 +1848,7 @@ def _validate_cluster_sample_size(self, sample_size: int):
_validate_cluster_sample_size(sample_size, spec.n_clusters, spec.cluster_size).raise_if_invalid()
- def _parse_target_tests(self, target_test: Union[str, List[str]]) -> List[str]:
+ def _parse_target_tests(self, target_test: Union[str, List[str]], test_formula_effects: Optional[List[str]] = None) -> List[str]:
"""Parse a target_test argument into a list of effect names to test.
Supports regular effect names (e.g. ``"x1"``, ``"overall"``),
@@ -1875,7 +1898,10 @@ def _parse_target_tests(self, target_test: Union[str, List[str]]) -> List[str]:
cluster_effects = self._registry.cluster_effect_names
if "all" in keywords:
- fixed_effects = [e for e in self._registry.effect_names if e not in cluster_effects]
+ if test_formula_effects is not None:
+ fixed_effects = [e for e in test_formula_effects if e not in cluster_effects]
+ else:
+ fixed_effects = [e for e in self._registry.effect_names if e not in cluster_effects]
keyword_expansion += ["overall"] + fixed_effects
if "all-posthoc" in keywords:
@@ -1929,6 +1955,17 @@ def _parse_target_tests(self, target_test: Union[str, List[str]]) -> List[str]:
"(e.g. 'all'), do not also list tests that are already included."
)
+ # -- Phase 7b: Validate explicit tests against test formula ----------------
+ if test_formula_effects is not None:
+ test_formula_set = set(test_formula_effects)
+ for test in expanded:
+ if " vs " in test or test == "overall":
+ continue
+ if test not in test_formula_set:
+ raise ValueError(
+ f"Target test '{test}' is not in the test formula. Available effects: {', '.join(test_formula_effects)}"
+ )
+
# -- Phase 8: Parse posthoc specs + validate ------------------------------
regular_tests: list[str] = []
posthoc_specs: list[PostHocSpec] = []
@@ -1982,6 +2019,8 @@ def _parse_target_tests(self, target_test: Union[str, List[str]]) -> List[str]:
# User level k (k≥2) = dummy factor[k]
effect_order = list(self._registry._effects.keys())
+ # Returns None for the reference level, which is absorbed into the
+ # intercept in dummy coding and has no dedicated design matrix column.
def _level_to_col(factor_name, user_level, _effect_order=effect_order):
factor_info = self._registry._factors[factor_name]
reference = factor_info.get("reference_level", 1)
@@ -2069,30 +2108,60 @@ def _create_X_extended(self, X):
return np.column_stack(columns) if columns else np.empty((X.shape[0], 0))
- def _prepare_metadata(self, target_tests, correction=None):
+ def _prepare_metadata(self, target_tests, correction=None, test_formula_effects=None):
"""Pre-compute all static simulation metadata from the current model state."""
- return prepare_metadata(self, target_tests, correction)
+ return prepare_metadata(self, target_tests, correction, test_formula_effects=test_formula_effects)
- def _resolve_test_formula(self, test_formula: str) -> str:
- """Resolve test formula and update _test_method accordingly.
+ def _resolve_test_formula(self, test_formula: str):
+ """Resolve test formula, validate, parse, and update _test_method.
- Returns the resolved formula string.
+ Returns:
+ Tuple of (formula_string, test_effect_names, random_effects).
+ test_effect_names is None when test_formula is empty (use generation formula).
"""
+ from .utils.parsers import _parse_equation
+
if not test_formula:
resolved = self._registry.equation
- else:
- resolved = test_formula
+ _, _, random_effects = _parse_equation(resolved)
+ if random_effects:
+ self._test_method = "mixed_model"
+ else:
+ self._test_method = "linear_regression"
+ return resolved, None, []
- from .utils.parsers import _parse_equation
+ # Validate test formula variables exist in the model
+ from .utils.validators import _validate_test_formula
- _, _, random_effects = _parse_equation(resolved)
+ available_vars = (
+ [self._registry.dependent] + self._registry.non_factor_names + self._registry.factor_names + self._registry.cluster_names
+ )
+ validation = _validate_test_formula(test_formula, available_vars)
+ validation.raise_if_invalid()
+
+ # Parse test formula to get effects and random effects
+ from .utils.test_formula_utils import _extract_test_formula_effects
+
+ test_effects, random_effects = _extract_test_formula_effects(test_formula, self._registry)
+
+ if not test_effects:
+ raise ValueError(f"test_formula '{test_formula}' contains no testable effects from the data generation model.")
+
+ # Check for OLS -> LME cross (invalid: no cluster data to fit)
+ if random_effects and not self._registry._cluster_specs:
+ grouping_vars = [re["grouping_var"] for re in random_effects]
+ raise ValueError(
+ f"test_formula contains random effects ({grouping_vars}) but the "
+ f"data generation model has no cluster structure. Cannot fit a "
+ f"mixed model to data without clusters."
+ )
if random_effects:
self._test_method = "mixed_model"
else:
self._test_method = "linear_regression"
- return resolved
+ return test_formula, test_effects, random_effects
def _run_find_power(
self,
@@ -2101,6 +2170,8 @@ def _run_find_power(
correction,
scenario_config=None,
test_formula=None,
+ test_formula_effects=None,
+ test_random_effects=None,
progress=None,
cancel_check=None,
):
@@ -2109,13 +2180,15 @@ def _run_find_power(
self._validate_cluster_sample_size(sample_size)
# Route based on test method (routing logic handled in simulation.py)
- metadata = self._prepare_metadata(target_tests, correction)
+ metadata = self._prepare_metadata(target_tests, correction, test_formula_effects)
- if scenario_config:
- metadata.heterogeneity = scenario_config["heterogeneity"]
- metadata.heteroskedasticity = scenario_config["heteroskedasticity"]
- if metadata.cluster_specs:
- metadata.lme_scenario_config = scenario_config
+ # Set the random effects flag for test formula
+ if test_random_effects:
+ metadata.test_has_random_effects = True
+
+ # scenario_config is always a dict (SCENARIO_ZERO or user-provided)
+ metadata.heterogeneity = scenario_config["heterogeneity"]
+ metadata.heteroskedasticity = scenario_config["heteroskedasticity"]
runner = SimulationRunner(
n_simulations=self._effective_n_simulations,
@@ -2127,9 +2200,15 @@ def _run_find_power(
)
# Compute critical values once before the simulation loop
- p = len(metadata.effect_sizes)
+ # Use test formula's effect count for critical values when subsetting
+ if metadata.test_column_indices is not None:
+ p = metadata.test_effect_count
+ n_targets = len(metadata.test_target_indices)
+ else:
+ p = len(metadata.effect_sizes)
+ n_targets = len(metadata.target_indices)
+
dof = sample_size - p - 1
- n_targets = len(metadata.target_indices)
n_posthoc = len(metadata.posthoc_specs)
if n_posthoc > 0 and metadata.posthoc_method == "t-test":
@@ -2185,7 +2264,7 @@ def analyze_func(X, y, indices, alpha, correction):
analyze_func=analyze_func,
create_X_extended_func=self._create_X_extended,
scenario_config=scenario_config,
- apply_perturbations_func=(apply_per_simulation_perturbations if scenario_config else None),
+ apply_perturbations_func=apply_per_simulation_perturbations,
progress=progress,
cancel_check=cancel_check,
)
@@ -2193,11 +2272,17 @@ def analyze_func(X, y, indices, alpha, correction):
if not sim_results:
return {}
+ # When test formula is active, filter target_tests to only effects in the test model
+ effective_target_tests = target_tests
+ if test_formula_effects is not None:
+ test_effect_set = set(test_formula_effects)
+ effective_target_tests = [t for t in target_tests if t == "overall" or t in test_effect_set]
+
processor = ResultsProcessor(target_power=self.power)
power_results = processor.calculate_powers(
sim_results["all_results"],
sim_results["all_results_corrected"],
- target_tests,
+ effective_target_tests,
)
# Add n_simulations_failed to power_results
@@ -2207,13 +2292,13 @@ def analyze_func(X, y, indices, alpha, correction):
# Tukey correction only applies to pairwise contrasts; NaN-ify others
if correction and correction.lower() == "tukey" and power_results.get("individual_powers_corrected"):
posthoc_labels = {s.label for s in self._posthoc_specs}
- for test in target_tests:
+ for test in effective_target_tests:
if test not in posthoc_labels:
power_results["individual_powers_corrected"][test] = float("nan")
return build_power_result(
model_type=self.model_type,
- target_tests=target_tests,
+ target_tests=effective_target_tests,
formula_to_test=test_formula,
equation=self.equation,
sample_size=sample_size,
@@ -2243,12 +2328,15 @@ def _run_sample_size_analysis(
correction,
scenario_config=None,
test_formula=None,
+ test_formula_effects=None,
+ test_random_effects=None,
progress=None,
cancel_check=None,
):
"""Iterate over sample sizes, running power analysis for each."""
from .progress import SimulationCancelled
+ use_sequential = True
if self._is_parallel_effective():
from joblib import Parallel, delayed
@@ -2258,7 +2346,18 @@ def _run_sample_size_analysis(
backend="loky",
verbose=0,
return_as="generator",
- )(delayed(self._run_find_power)(ss, target_tests, correction, scenario_config, test_formula) for ss in sample_sizes)
+ )(
+ delayed(self._run_find_power)(
+ ss,
+ target_tests,
+ correction,
+ scenario_config,
+ test_formula,
+ test_formula_effects,
+ test_random_effects,
+ )
+ for ss in sample_sizes
+ )
results = []
for ss, result in zip(sample_sizes, power_results, strict=False):
if cancel_check is not None and cancel_check():
@@ -2266,25 +2365,13 @@ def _run_sample_size_analysis(
results.append((ss, result))
if progress is not None:
progress.advance(self._effective_n_simulations)
+ use_sequential = False
except Exception as e:
if isinstance(e, SimulationCancelled):
raise
print(f"Warning: Parallel execution failed ({e}). Falling back to sequential.")
- results = []
- for ss in sample_sizes:
- if cancel_check is not None and cancel_check():
- raise SimulationCancelled("Simulation cancelled by user") from None
- result = self._run_find_power(
- ss,
- target_tests,
- correction,
- scenario_config,
- test_formula,
- progress=progress,
- cancel_check=cancel_check,
- )
- results.append((ss, result))
- else:
+
+ if use_sequential:
results = []
for sample_size in sample_sizes:
if cancel_check is not None and cancel_check():
@@ -2295,19 +2382,28 @@ def _run_sample_size_analysis(
correction,
scenario_config,
test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=progress,
cancel_check=cancel_check,
)
results.append((sample_size, power_result))
processor = ResultsProcessor(target_power=self.power)
- analysis_results = processor.process_sample_size_results(results, target_tests, correction)
+ # Filter target_tests to match test formula effects
+ if test_formula_effects is not None:
+ test_set = set(test_formula_effects)
+ effective_target_tests = [t for t in target_tests if t in test_set or t == "overall"]
+ else:
+ effective_target_tests = target_tests
+
+ analysis_results = processor.process_sample_size_results(results, effective_target_tests, correction)
# Tukey correction only applies to pairwise contrasts; NaN-ify others
if correction and correction.lower() == "tukey":
posthoc_labels = {s.label for s in self._posthoc_specs}
if analysis_results.get("powers_by_test_corrected"):
- for test in target_tests:
+ for test in effective_target_tests:
if test not in posthoc_labels:
n_points = len(analysis_results["powers_by_test_corrected"][test])
analysis_results["powers_by_test_corrected"][test] = [float("nan")] * n_points
@@ -2315,7 +2411,7 @@ def _run_sample_size_analysis(
return build_sample_size_result(
model_type=self.model_type,
- target_tests=target_tests,
+ target_tests=effective_target_tests,
formula_to_test=test_formula,
equation=self.equation,
sample_sizes=sample_sizes,
@@ -2331,9 +2427,16 @@ def _run_scenario_analysis(self, analysis_type, **kwargs):
"""Delegate to ScenarioRunner for multi-scenario power or sample-size analysis."""
from functools import partial
- configs = self._scenario_configs or DEFAULT_SCENARIO_CONFIG
+ all_configs = self._scenario_configs or DEFAULT_SCENARIO_CONFIG
+ scenario_filter = kwargs.pop("scenario_filter", None)
+ if scenario_filter is not None:
+ configs = {k: all_configs[k] for k in scenario_filter}
+ else:
+ configs = all_configs
scenario_runner = ScenarioRunner(self, configs)
test_formula = kwargs.get("test_formula")
+ test_formula_effects = kwargs.get("test_formula_effects")
+ test_random_effects = kwargs.get("test_random_effects")
progress = kwargs.get("progress")
cancel_check = kwargs.get("cancel_check")
@@ -2341,6 +2444,8 @@ def _run_scenario_analysis(self, analysis_type, **kwargs):
run_power_func = partial(
self._run_find_power,
test_formula=test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=progress,
cancel_check=cancel_check,
)
@@ -2357,6 +2462,8 @@ def _run_scenario_analysis(self, analysis_type, **kwargs):
run_ss_func = partial(
self._run_sample_size_analysis,
test_formula=test_formula,
+ test_formula_effects=test_formula_effects,
+ test_random_effects=test_random_effects,
progress=progress,
cancel_check=cancel_check,
)
diff --git a/mcpower/progress.py b/mcpower/progress.py
index e733148..dca3c25 100644
--- a/mcpower/progress.py
+++ b/mcpower/progress.py
@@ -87,10 +87,7 @@ def __init__(self, **tqdm_kwargs):
self._bar = None
def __call__(self, current: int, total: int):
- try:
- from tqdm import tqdm
- except ImportError:
- raise ImportError("tqdm is required for TqdmReporter. Install with: pip install tqdm") from None
+ from tqdm import tqdm
if self._bar is None:
self._bar = tqdm(total=total, unit="sim", **self._tqdm_kwargs)
diff --git a/mcpower/stats/data_generation.py b/mcpower/stats/data_generation.py
index 0d8800c..3c46d89 100644
--- a/mcpower/stats/data_generation.py
+++ b/mcpower/stats/data_generation.py
@@ -23,7 +23,6 @@
SKEW_STD = np.sqrt(np.exp(2) - np.exp(1))
NORM_SCALE = (DIST_RESOLUTION - 1) / (NORM_RANGE[1] - NORM_RANGE[0])
PERC_SCALE = (DIST_RESOLUTION - 1) / (PERCENTILE_RANGE[1] - PERCENTILE_RANGE[0])
-FLOAT_NEAR_ZERO = 1e-15
# Global lookup tables
NORM_CDF_TABLE = None
@@ -58,13 +57,12 @@ def _compute_t3_sd():
Replicates the vectorised norm-CDF -> t(3)-PPF lookup chain on a large
fixed-seed sample to get a stable SD estimate.
"""
- assert NORM_CDF_TABLE is not None
- assert T3_PPF_TABLE is not None
+ if NORM_CDF_TABLE is None or T3_PPF_TABLE is None:
+ raise RuntimeError("Distribution tables not initialized — _init_tables() must be called first")
- rng_state = np.random.get_state()
- np.random.seed(999999)
- z = np.random.standard_normal(200000)
- np.random.set_state(rng_state)
+ # Use a local RNG to avoid affecting the global state and to be thread-safe.
+ rng = np.random.RandomState(999999)
+ z = rng.standard_normal(200000)
# Step 1: Normal CDF lookup (z -> percentile)
z_clipped = np.clip(z, NORM_RANGE[0], NORM_RANGE[1])
@@ -107,9 +105,16 @@ def create_uploaded_lookup_tables(
for var_idx in range(n_vars):
data = data_matrix[:, var_idx]
- normalized = (data - np.mean(data)) / np.std(data)
+ std = np.std(data)
+ if std < 1e-15:
+ raise ValueError(
+ f"Variable at index {var_idx} has zero variance (constant value). Remove it from the model or check your data."
+ )
+ normalized = (data - np.mean(data)) / std
sorted_uploaded = np.sort(normalized)
+ # Weibull plotting positions: i/(n+1) avoids 0 and 1, which would map
+ # to -inf/+inf under the normal PPF, giving well-behaved quantiles.
percentiles = np.linspace(1 / (n_samples + 1), n_samples / (n_samples + 1), n_samples)
normal_quantiles = norm_ppf_array(percentiles)
@@ -126,13 +131,12 @@ def _generate_factors(sample_size, factor_specs, seed):
Args:
sample_size: Number of observations
factor_specs: List of {'n_levels': int, 'proportions': [float, ...]}
- seed: Random seed
+ seed: Random seed (callers pass sim_seed + 3)
Returns:
X_factors: (sample_size, total_dummies) array
"""
- if seed is not None:
- np.random.seed(seed)
+ rng = np.random.RandomState(seed)
if not factor_specs:
return np.empty((sample_size, 0), dtype=float)
@@ -141,7 +145,7 @@ def _generate_factors(sample_size, factor_specs, seed):
for spec in factor_specs:
n_levels = spec["n_levels"]
proportions = spec["proportions"]
- factor_data = np.random.choice(n_levels, size=sample_size, p=proportions)
+ factor_data = rng.choice(n_levels, size=sample_size, p=proportions)
dummies = np.eye(n_levels, dtype=float)[factor_data]
factor_columns.append(dummies[:, 1:])
@@ -170,12 +174,11 @@ def bootstrap_uploaded_data(
X_non_factors: Non-factor variables (continuous + binary mapped to 0-1)
X_factors: Factor dummy variables
"""
- if seed is not None:
- np.random.seed(seed)
+ rng = np.random.RandomState(seed)
# Bootstrap whole rows
n_samples = raw_data.shape[0]
- row_indices = np.random.choice(n_samples, size=sample_size, replace=True)
+ row_indices = rng.choice(n_samples, size=sample_size, replace=True)
bootstrapped_data = raw_data[row_indices, :]
# Separate by type
@@ -286,12 +289,17 @@ def _generate_cluster_effects(
Returns:
X_cluster: (sample_size, n_cluster_vars) array of random effect columns
"""
- if sim_seed is not None:
- # Use a derived seed to avoid collision with X generation seed
- np.random.seed(sim_seed + 3)
+ rng = np.random.RandomState(sim_seed + 4 if sim_seed is not None else None)
columns = []
+ # Extract perturbation defaults once
+ perturb = lme_perturbations or {}
+ tau_mults = perturb.get("tau_squared_multipliers", {})
+ re_dist_val = perturb.get("random_effect_dist", "normal")
+ re_df_val = perturb.get("random_effect_df", 5)
+ has_perturb = lme_perturbations is not None
+
for gv, spec in cluster_specs.items():
n_clusters = spec["n_clusters"]
cluster_size = spec["cluster_size"]
@@ -302,19 +310,16 @@ def _generate_cluster_effects(
cluster_size = sample_size // n_clusters
# Apply LME perturbations if present
- if lme_perturbations is not None:
- multiplier = lme_perturbations["tau_squared_multipliers"].get(gv, 1.0)
- tau_sq = tau_sq * multiplier
+ if has_perturb:
+ tau_sq = tau_sq * tau_mults.get(gv, 1.0)
tau = np.sqrt(tau_sq)
# Generate random intercepts (possibly non-normal)
- if lme_perturbations is not None:
- re_dist = lme_perturbations.get("random_effect_dist", "normal")
- re_df = lme_perturbations.get("random_effect_df", 5)
- random_intercepts = _generate_non_normal_intercepts(n_clusters, tau, re_dist, re_df)
+ if has_perturb:
+ random_intercepts = _generate_non_normal_intercepts(n_clusters, tau, re_dist_val, re_df_val, rng_state=rng)
else:
- random_intercepts = np.random.normal(0, tau, size=n_clusters)
+ random_intercepts = rng.normal(0, tau, size=n_clusters)
# Create id_effect column: repeat each cluster's intercept
# cluster_id assignment: [0,0,...,0, 1,1,...,1, ..., K-1,K-1,...,K-1]
@@ -413,14 +418,20 @@ def _generate_random_effects(
A :class:`RandomEffectsResult` with intercept columns, slope
contributions, cluster IDs, Z matrices, and nesting metadata.
"""
- if sim_seed is not None:
- np.random.seed(sim_seed + 3)
+ rng = np.random.RandomState(sim_seed + 4 if sim_seed is not None else None)
intercept_cols: List[np.ndarray] = []
slope_contribution = np.zeros(sample_size)
cluster_ids_dict: Dict[str, np.ndarray] = {}
Z_matrices: Dict[str, np.ndarray] = {}
+ # Extract perturbation defaults once (avoids repeated dict lookups)
+ perturb = lme_perturbations or {}
+ tau_multipliers = perturb.get("tau_squared_multipliers", {})
+ re_dist = perturb.get("random_effect_dist", "normal")
+ re_df = perturb.get("random_effect_df", 5)
+ has_perturbations = lme_perturbations is not None
+
# Nested model bookkeeping
child_to_parent: Optional[np.ndarray] = None
K_parent = 0
@@ -460,19 +471,16 @@ def _generate_random_effects(
# Apply LME perturbations: ICC jitter on tau_squared
tau_sq = spec["tau_squared"]
- if lme_perturbations is not None:
- multiplier = lme_perturbations["tau_squared_multipliers"].get(gv, 1.0)
- tau_sq = tau_sq * multiplier
+ if has_perturbations:
+ tau_sq = tau_sq * tau_multipliers.get(gv, 1.0)
if q == 1:
# --- Random intercept only ---
tau = np.sqrt(tau_sq)
- if lme_perturbations is not None:
- re_dist = lme_perturbations.get("random_effect_dist", "normal")
- re_df = lme_perturbations.get("random_effect_df", 5)
- random_intercepts = _generate_non_normal_intercepts(n_clusters, tau, re_dist, re_df)
+ if has_perturbations:
+ random_intercepts = _generate_non_normal_intercepts(n_clusters, tau, re_dist, re_df, rng_state=rng)
else:
- random_intercepts = np.random.normal(0, tau, size=n_clusters)
+ random_intercepts = rng.normal(0, tau, size=n_clusters)
id_effect = _trim_or_pad(np.repeat(random_intercepts, cluster_size), sample_size)
intercept_cols.append(id_effect)
@@ -482,33 +490,29 @@ def _generate_random_effects(
slope_vars = spec.get("random_slope_vars", [])
# Apply ICC jitter to G_matrix intercept variance
- if lme_perturbations is not None:
+ if has_perturbations:
ratio = tau_sq / spec["tau_squared"] if spec["tau_squared"] > 0 else 1.0
# Scale intercept row/column of G by sqrt(ratio)
sqrt_ratio = np.sqrt(ratio)
G_matrix[0, :] *= sqrt_ratio
G_matrix[:, 0] *= sqrt_ratio
- # Draw correlated [b_int, b_slope1, ...] per cluster
- re_dist = lme_perturbations.get("random_effect_dist", "normal") if lme_perturbations else "normal"
- re_df = lme_perturbations.get("random_effect_df", 5) if lme_perturbations else 5
-
- if re_dist == "heavy_tailed" and lme_perturbations is not None:
+ if re_dist == "heavy_tailed" and has_perturbations:
# Multivariate t: MVN(0, G * (df-2)/df) × sqrt(df / chi2(df))
df = max(re_df, 3)
G_scaled = G_matrix * ((df - 2.0) / df)
- b_normal = np.random.multivariate_normal(np.zeros(q), G_scaled, size=n_clusters)
- chi2_samples = np.random.chisquare(df, size=n_clusters)
+ b_normal = rng.multivariate_normal(np.zeros(q), G_scaled, size=n_clusters)
+ chi2_samples = rng.chisquare(df, size=n_clusters)
mixing = np.sqrt(df / chi2_samples)
b = b_normal * mixing[:, np.newaxis]
- elif re_dist == "skewed" and lme_perturbations is not None:
+ elif re_dist == "skewed" and has_perturbations:
# Independent skewed marginals via shifted chi-squared, scaled by Cholesky
df = max(re_df, 3)
L = np.linalg.cholesky(G_matrix)
- raw = (np.random.chisquare(df, size=(n_clusters, q)) - df) / np.sqrt(2 * df)
+ raw = (rng.chisquare(df, size=(n_clusters, q)) - df) / np.sqrt(2 * df)
b = raw @ L.T
else:
- b = np.random.multivariate_normal(np.zeros(q), G_matrix, size=n_clusters)
+ b = rng.multivariate_normal(np.zeros(q), G_matrix, size=n_clusters)
# Intercept component
intercept_effect = _trim_or_pad(np.repeat(b[:, 0], cluster_size), sample_size)
@@ -549,21 +553,19 @@ def _generate_random_effects(
tau_sq_parent = p_spec["tau_squared"]
tau_sq_child = c_spec["tau_squared"]
- if lme_perturbations is not None:
- tau_sq_parent *= lme_perturbations["tau_squared_multipliers"].get(p_gv, 1.0)
- tau_sq_child *= lme_perturbations["tau_squared_multipliers"].get(c_gv, 1.0)
+ if has_perturbations:
+ tau_sq_parent *= tau_multipliers.get(p_gv, 1.0)
+ tau_sq_child *= tau_multipliers.get(c_gv, 1.0)
tau_parent = np.sqrt(tau_sq_parent)
tau_child = np.sqrt(tau_sq_child)
- if lme_perturbations is not None:
- re_dist = lme_perturbations.get("random_effect_dist", "normal")
- re_df = lme_perturbations.get("random_effect_df", 5)
- b_parent = _generate_non_normal_intercepts(K_parent, tau_parent, re_dist, re_df)
- b_child = _generate_non_normal_intercepts(K_child, tau_child, re_dist, re_df)
+ if has_perturbations:
+ b_parent = _generate_non_normal_intercepts(K_parent, tau_parent, re_dist, re_df, rng_state=rng)
+ b_child = _generate_non_normal_intercepts(K_child, tau_child, re_dist, re_df, rng_state=rng)
else:
- b_parent = np.random.normal(0, tau_parent, size=K_parent)
- b_child = np.random.normal(0, tau_child, size=K_child)
+ b_parent = rng.normal(0, tau_parent, size=K_parent)
+ b_child = rng.normal(0, tau_child, size=K_child)
# IDs: parent_ids assigns each observation to a parent cluster,
# child_ids assigns each observation to a child cluster.
diff --git a/mcpower/stats/distributions.py b/mcpower/stats/distributions.py
index cf13bca..6ee5265 100644
--- a/mcpower/stats/distributions.py
+++ b/mcpower/stats/distributions.py
@@ -3,9 +3,7 @@
Provides F, t, chi2, normal, and studentized range distribution
functions plus batch critical-value computation and table generation.
-Backend priority:
- 1. C++ native (Boost.Math + R Tukey port) via mcpower_native
- 2. scipy (optional shim, for when C++ is not compiled)
+All functions are provided by the C++ native backend (Boost.Math + R Tukey port).
Usage:
from mcpower.stats.distributions import norm_ppf, compute_critical_values_ols
@@ -14,13 +12,11 @@
import numpy as np
# ============================================================================
-# Backend selection
+# Backend — native C++ only
# ============================================================================
-_BACKEND = None
-
try:
- from mcpower.backends.mcpower_native import ( # type: ignore[import]
+ from mcpower.backends.mcpower_native import ( # type: ignore[import] # noqa: F401
chi2_cdf,
chi2_ppf,
compute_critical_values_lme,
@@ -36,171 +32,17 @@
t_ppf,
)
- _BACKEND = "native"
-
-except ImportError:
- # -------------------------------------------------------------------
- # scipy shim -- temporary fallback for when C++ is not compiled.
- # Will be removed when Python fallback backends are fully dropped.
- # -------------------------------------------------------------------
- try:
- from scipy.stats import ( # isort: skip
- chi2 as _chi2_dist,
- f as _f_dist,
- norm as _norm_dist,
- studentized_range as _sr_dist,
- t as _t_dist,
- )
-
- def norm_ppf(p): # noqa: F811
- """Standard normal quantile function (inverse CDF)."""
- return float(_norm_dist.ppf(p))
-
- def norm_cdf(x): # noqa: F811
- """Standard normal CDF."""
- return float(_norm_dist.cdf(x))
-
- def t_ppf(p, df): # noqa: F811
- """Student's t quantile function."""
- return float(_t_dist.ppf(p, df))
-
- def f_ppf(p, dfn, dfd): # noqa: F811
- """Fisher F quantile function."""
- return float(_f_dist.ppf(p, dfn, dfd))
-
- def chi2_ppf(p, df): # noqa: F811
- """Chi-squared quantile function."""
- return float(_chi2_dist.ppf(p, df))
-
- def chi2_cdf(x, df): # noqa: F811
- """Chi-squared CDF."""
- return float(_chi2_dist.cdf(x, df))
-
- def studentized_range_ppf(p, k, df): # noqa: F811
- """Studentized range quantile (Tukey). k=groups, df=denom df."""
- if df < 2 or k < 2 or k > 200 or p <= 0.0 or p >= 1.0:
- return float("inf")
- return float(_sr_dist.ppf(p, k, df))
-
- def compute_critical_values_ols(alpha, dfn, dfd, n_targets, correction_method): # noqa: F811
- """Compute OLS critical values using scipy (fallback).
-
- Args:
- alpha: Significance level.
- dfn: Numerator degrees of freedom (number of predictors).
- dfd: Denominator degrees of freedom (n - p - 1).
- n_targets: Number of individual effects being tested.
- correction_method: 0=none, 1=Bonferroni, 2=FDR (BH), 3=Holm.
-
- Returns:
- Tuple of (f_crit, t_crit, correction_t_crits) where
- correction_t_crits is an ndarray of length n_targets.
- """
- if dfd <= 0:
- return np.inf, np.inf, np.full(max(n_targets, 1), np.inf)
-
- f_crit = _f_dist.ppf(1 - alpha, dfn, dfd) if dfn > 0 else np.inf
- t_crit = _t_dist.ppf(1 - alpha / 2, dfd)
-
- m = n_targets
- if m == 0:
- return f_crit, t_crit, np.empty(0)
-
- if correction_method == 0: # None
- correction_t_crits = np.full(m, t_crit)
- elif correction_method == 1: # Bonferroni
- bonf_crit = _t_dist.ppf(1 - alpha / (2 * m), dfd)
- correction_t_crits = np.full(m, bonf_crit)
- elif correction_method == 2: # FDR (Benjamini-Hochberg)
- correction_t_crits = np.array(
- [_t_dist.ppf(1 - (k + 1) / m * alpha / 2, dfd) if (k + 1) / m * alpha / 2 >= 1e-12 else np.inf for k in range(m)]
- )
- elif correction_method == 3: # Holm
- correction_t_crits = np.array(
- [_t_dist.ppf(1 - alpha / (2 * (m - k)), dfd) if alpha / (2 * (m - k)) >= 1e-12 else np.inf for k in range(m)]
- )
- else:
- correction_t_crits = np.full(m, t_crit)
-
- return f_crit, t_crit, correction_t_crits
-
- def compute_tukey_critical_value(alpha, n_levels, dfd): # noqa: F811
- """Compute Tukey HSD critical value (q / sqrt(2))."""
- if dfd <= 0:
- return np.inf
- q_crit = _sr_dist.ppf(1 - alpha, n_levels, dfd)
- return q_crit / np.sqrt(2)
-
- def compute_critical_values_lme(alpha, n_fixed, n_targets, correction_method): # noqa: F811
- """Compute LME critical values using scipy (fallback).
-
- Args:
- alpha: Significance level.
- n_fixed: Number of fixed effects (excluding intercept).
- n_targets: Number of individual effects being tested.
- correction_method: 0=none, 1=Bonferroni, 2=FDR (BH), 3=Holm.
-
- Returns:
- Tuple of (chi2_crit, z_crit, correction_z_crits) where
- correction_z_crits is an ndarray of length n_targets.
- """
- chi2_crit = _chi2_dist.ppf(1 - alpha, n_fixed) if n_fixed > 0 else np.inf
- z_crit = _norm_dist.ppf(1 - alpha / 2)
-
- m = n_targets
- if m == 0:
- return chi2_crit, z_crit, np.empty(0)
-
- if correction_method == 0: # None
- correction_z_crits = np.full(m, z_crit)
- elif correction_method == 1: # Bonferroni
- bonf = _norm_dist.ppf(1 - alpha / (2 * m))
- correction_z_crits = np.full(m, bonf)
- elif correction_method == 2: # FDR (Benjamini-Hochberg)
- correction_z_crits = np.array(
- [_norm_dist.ppf(1 - (k + 1) / m * alpha / 2) if (k + 1) / m * alpha / 2 >= 1e-12 else np.inf for k in range(m)]
- )
- elif correction_method == 3: # Holm
- correction_z_crits = np.array(
- [_norm_dist.ppf(1 - alpha / (2 * (m - k))) if alpha / (2 * (m - k)) >= 1e-12 else np.inf for k in range(m)]
- )
- else:
- correction_z_crits = np.full(m, z_crit)
-
- return chi2_crit, z_crit, correction_z_crits
-
- def generate_norm_cdf_table(x_min, x_max, resolution): # noqa: F811
- """Generate normal CDF lookup table."""
- x = np.linspace(x_min, x_max, resolution)
- return _norm_dist.cdf(x).astype(np.float64)
-
- def generate_t3_ppf_table(perc_min, perc_max, resolution): # noqa: F811
- """Generate t(3) PPF lookup table (divided by sqrt(3))."""
- p = np.linspace(perc_min, perc_max, resolution)
- return (_t_dist.ppf(p, 3) / np.sqrt(3)).astype(np.float64)
-
- def norm_ppf_array(percentiles): # noqa: F811
- """Vectorized normal PPF for percentile array."""
- return _norm_dist.ppf(np.asarray(percentiles)).astype(np.float64)
-
- _BACKEND = "scipy"
-
- except ImportError as exc:
- raise ImportError(
- "No distribution backend available. "
- "Install from PyPI for prebuilt C++ wheels: pip install MCPower\n"
- "Or install scipy as fallback: pip install scipy"
- ) from exc
+except ImportError as exc:
+ raise ImportError("Native C++ backend not available. Install from PyPI for prebuilt wheels: pip install MCPower") from exc
# ============================================================================
-# Also re-export scipy optimizer shims for lme_solver.py
-# These replace scipy.optimize.minimize and minimize_scalar
+# Optimizer wrappers for lme_solver.py
# ============================================================================
def minimize_lbfgsb(objective, x0, bounds, maxiter=200, ftol=1e-10, gtol=1e-6):
- """L-BFGS-B minimization -- C++ native or scipy fallback.
+ """L-BFGS-B minimization via native C++ backend.
Args:
objective: Callable f(x) -> float
@@ -213,53 +55,15 @@ def minimize_lbfgsb(objective, x0, bounds, maxiter=200, ftol=1e-10, gtol=1e-6):
Returns:
Object with .x (optimal point), .fun (optimal value), .converged (bool)
"""
- if _BACKEND == "native":
- try:
- from mcpower.backends.mcpower_native import lbfgsb_minimize_fd # type: ignore[import]
-
- lb = np.array([b[0] for b in bounds])
- ub = np.array([b[1] for b in bounds])
- return lbfgsb_minimize_fd(objective, np.asarray(x0, dtype=np.float64), lb, ub, maxiter, ftol, gtol)
- except ImportError:
- import warnings
-
- warnings.warn(
- "Native L-BFGS-B optimizer not available despite native backend being loaded. Falling back to scipy.",
- RuntimeWarning,
- stacklevel=2,
- )
- except Exception as e:
- import warnings
+ from mcpower.backends.mcpower_native import lbfgsb_minimize_fd # type: ignore[import]
- warnings.warn(
- f"Native L-BFGS-B optimizer failed ({type(e).__name__}: {e}), falling back to scipy.",
- RuntimeWarning,
- stacklevel=2,
- )
-
- # scipy fallback
- from scipy.optimize import minimize
-
- result = minimize(
- objective,
- x0,
- method="L-BFGS-B",
- bounds=bounds,
- options={"maxiter": maxiter, "ftol": ftol, "gtol": gtol},
- )
-
- class _Result:
- __slots__ = ("x", "fun", "converged")
-
- r = _Result()
- r.x = result.x
- r.fun = result.fun
- r.converged = result.success
- return r
+ lb = np.array([b[0] for b in bounds])
+ ub = np.array([b[1] for b in bounds])
+ return lbfgsb_minimize_fd(objective, np.asarray(x0, dtype=np.float64), lb, ub, maxiter, ftol, gtol)
def minimize_scalar_brent(objective, bounds, tol=1e-8, maxiter=150):
- """Brent 1D minimization -- C++ native or scipy fallback.
+ """Brent 1D minimization via native C++ backend.
Args:
objective: Callable f(x) -> float
@@ -270,43 +74,6 @@ def minimize_scalar_brent(objective, bounds, tol=1e-8, maxiter=150):
Returns:
Object with .x (optimal point), .fun (optimal value), .converged (bool)
"""
- if _BACKEND == "native":
- try:
- from mcpower.backends.mcpower_native import brent_minimize_scalar # type: ignore[import]
-
- return brent_minimize_scalar(objective, bounds[0], bounds[1], tol, maxiter)
- except ImportError:
- import warnings
-
- warnings.warn(
- "Native Brent optimizer not available despite native backend being loaded. Falling back to scipy.",
- RuntimeWarning,
- stacklevel=2,
- )
- except Exception as e:
- import warnings
-
- warnings.warn(
- f"Native Brent optimizer failed ({type(e).__name__}: {e}), falling back to scipy.",
- RuntimeWarning,
- stacklevel=2,
- )
-
- # scipy fallback
- from scipy.optimize import minimize_scalar
-
- result = minimize_scalar(
- objective,
- bounds=bounds,
- method="bounded",
- options={"xatol": tol, "maxiter": maxiter},
- )
-
- class _Result:
- __slots__ = ("x", "fun", "converged")
+ from mcpower.backends.mcpower_native import brent_minimize_scalar # type: ignore[import]
- r = _Result()
- r.x = result.x
- r.fun = result.fun
- r.converged = bool(getattr(result, "success", True))
- return r
+ return brent_minimize_scalar(objective, bounds[0], bounds[1], tol, maxiter)
diff --git a/mcpower/stats/mixed_models.py b/mcpower/stats/mixed_models.py
index 414e911..07de5e8 100644
--- a/mcpower/stats/mixed_models.py
+++ b/mcpower/stats/mixed_models.py
@@ -11,10 +11,12 @@
import threading
import warnings
-from typing import Any, Dict, List, Optional, Union
+from typing import Any, Dict, Optional, Union
import numpy as np
+from ..backends.native import _prep
+
# Suppress statsmodels convergence warnings (expected with small samples/low ICC).
# Module-level filterwarnings with module= is unreliable for statsmodels internals,
# so we also use catch_warnings() context managers around .fit() calls below.
@@ -30,7 +32,6 @@ def _lme_analysis_wrapper(
y: np.ndarray,
target_indices: np.ndarray,
cluster_ids: np.ndarray,
- cluster_column_indices: List[int],
correction_method: int,
alpha: float,
backend: str = "custom",
@@ -51,7 +52,6 @@ def _lme_analysis_wrapper(
y: (n,) response vector
target_indices: Coefficient indices to test (fixed effects only)
cluster_ids: (n,) cluster membership array [0,0,0, 1,1,1, ...]
- cluster_column_indices: Indices of cluster effect columns (unused)
correction_method: 0=none, 1=Bonferroni, 2=FDR, 3=Holm
alpha: Significance level
backend: "custom" (default) or "statsmodels" (fallback)
@@ -118,9 +118,7 @@ def _lme_analysis_wrapper(
verbose=verbose,
)
elif backend == "statsmodels":
- return _lme_analysis_statsmodels(
- X_expanded, y, target_indices, cluster_ids, cluster_column_indices, correction_method, alpha, verbose
- )
+ return _lme_analysis_statsmodels(X_expanded, y, target_indices, cluster_ids, correction_method, alpha, verbose)
else:
raise ValueError(f"Unknown backend: {backend}")
@@ -130,7 +128,6 @@ def _lme_analysis_statsmodels(
y: np.ndarray,
target_indices: np.ndarray,
cluster_ids: np.ndarray,
- cluster_column_indices: List[int],
correction_method: int,
alpha: float,
verbose: bool = False,
@@ -145,11 +142,10 @@ def _lme_analysis_statsmodels(
- Convergence retry strategy (allows ≤3% failures)
Args:
- X_expanded: (n, p) design matrix (includes cluster effect columns)
+ X_expanded: (n, p) design matrix (excludes cluster effect columns)
y: (n,) response vector
target_indices: Coefficient indices to test (fixed effects only)
cluster_ids: (n,) cluster membership array
- cluster_column_indices: Indices of cluster effect columns to remove
correction_method: 0=none, 1=Bonferroni, 2=FDR, 3=Holm
alpha: Significance level
verbose: Return detailed diagnostics
@@ -172,8 +168,6 @@ def _lme_analysis_statsmodels(
n, p = X_expanded.shape
n_targets = len(target_indices)
- # Note: X_expanded already excludes cluster effects (they're not in the design matrix)
- # cluster_column_indices is now unused in this function but kept for API compatibility
X_fixed = X_expanded
# Step 1: Add intercept to fixed effects
@@ -530,6 +524,29 @@ def _compute_wald_test(result, alpha):
return results_array
+def _ensure_lme_crits(alpha, p, n_targets, correction_method, chi2_crit, z_crit, correction_z_crits):
+ """Compute LME critical values on-the-fly if not precomputed."""
+ if z_crit is None or chi2_crit is None or correction_z_crits is None:
+ from .lme_solver import compute_lme_critical_values
+
+ return compute_lme_critical_values(alpha, p, n_targets, correction_method)
+ return chi2_crit, z_crit, correction_z_crits
+
+
+def _wrap_native_result(result, verbose, solver_name, extra_diag=None) -> Optional[Union[np.ndarray, Dict]]:
+ """Wrap C++ solver result with optional verbose diagnostics."""
+ if len(result) > 0:
+ if verbose:
+ diag = {"solver": solver_name}
+ if extra_diag:
+ diag.update(extra_diag)
+ return {"results": result, "diagnostics": diag}
+ return np.asarray(result)
+ if verbose:
+ return {"results": None, "failure_reason": f"C++ {solver_name} returned empty result"}
+ return None
+
+
def _lme_analysis_custom(
X_expanded: np.ndarray,
y: np.ndarray,
@@ -545,39 +562,29 @@ def _lme_analysis_custom(
"""LME analysis for random-intercept models via C++ backend.
Uses precomputed critical values (chi2_crit, z_crit) to avoid
- per-simulation scipy calls. Falls back to computing them if not provided.
+ per-simulation distribution calls. Falls back to computing them if not provided.
"""
n, p = X_expanded.shape
n_targets = len(target_indices)
K = int(cluster_ids.max()) + 1
- if z_crit is None or chi2_crit is None or correction_z_crits is None:
- from .lme_solver import compute_lme_critical_values
-
- chi2_crit, z_crit, correction_z_crits = compute_lme_critical_values(alpha, p, n_targets, correction_method)
+ chi2_crit, z_crit, correction_z_crits = _ensure_lme_crits(alpha, p, n_targets, correction_method, chi2_crit, z_crit, correction_z_crits)
from mcpower.backends import mcpower_native as _native # type: ignore[attr-defined]
result = _native.lme_analysis(
- np.ascontiguousarray(X_expanded, dtype=np.float64),
- np.ascontiguousarray(y, dtype=np.float64),
- np.ascontiguousarray(cluster_ids, dtype=np.int32),
+ _prep(X_expanded),
+ _prep(y),
+ _prep(cluster_ids, np.int32),
K,
- np.ascontiguousarray(target_indices, dtype=np.int32),
+ _prep(target_indices, np.int32),
float(chi2_crit),
float(z_crit),
- np.ascontiguousarray(correction_z_crits, dtype=np.float64),
+ _prep(correction_z_crits),
int(correction_method),
float(-1.0),
)
- if len(result) > 0:
- if verbose:
- return {"results": result, "diagnostics": {"solver": "native_q1"}}
- return result # type: ignore[no-any-return]
-
- if verbose:
- return {"results": None, "failure_reason": "C++ solver returned empty result"}
- return None
+ return _wrap_native_result(result, verbose, "native_q1")
def _lme_analysis_custom_general(
@@ -594,8 +601,6 @@ def _lme_analysis_custom_general(
verbose: bool = False,
) -> Optional[Union[np.ndarray, Dict]]:
"""LME analysis for random slopes (q > 1) via C++ backend."""
- from .lme_solver import compute_lme_critical_values
-
n, p = X_expanded.shape
n_targets = len(target_indices)
@@ -604,34 +609,26 @@ def _lme_analysis_custom_general(
q = Z.shape[1]
K = int(cluster_ids.max()) + 1
- if z_crit is None or chi2_crit is None or correction_z_crits is None:
- chi2_crit, z_crit, correction_z_crits = compute_lme_critical_values(alpha, p, n_targets, correction_method)
+ chi2_crit, z_crit, correction_z_crits = _ensure_lme_crits(alpha, p, n_targets, correction_method, chi2_crit, z_crit, correction_z_crits)
from mcpower.backends import mcpower_native as _native # type: ignore[attr-defined]
warm_theta_arr = np.empty(0, dtype=np.float64)
result = _native.lme_analysis_general(
- np.ascontiguousarray(X_expanded, dtype=np.float64),
- np.ascontiguousarray(y, dtype=np.float64),
- np.ascontiguousarray(Z, dtype=np.float64),
- np.ascontiguousarray(cluster_ids, dtype=np.int32),
+ _prep(X_expanded),
+ _prep(y),
+ _prep(Z),
+ _prep(cluster_ids, np.int32),
K,
q,
- np.ascontiguousarray(target_indices, dtype=np.int32),
+ _prep(target_indices, np.int32),
float(chi2_crit),
float(z_crit),
- np.ascontiguousarray(correction_z_crits, dtype=np.float64),
+ _prep(correction_z_crits),
int(correction_method),
warm_theta_arr,
)
- if len(result) > 0:
- if verbose:
- return {"results": result, "diagnostics": {"solver": "native_general", "q": q}}
- return result # type: ignore[no-any-return]
-
- if verbose:
- return {"results": None, "failure_reason": "C++ general solver returned empty result"}
- return None
+ return _wrap_native_result(result, verbose, "native_general", extra_diag={"q": q})
def _lme_analysis_custom_nested(
@@ -647,8 +644,6 @@ def _lme_analysis_custom_nested(
verbose: bool = False,
) -> Optional[Union[np.ndarray, Dict]]:
"""LME analysis for nested random intercepts via C++ backend."""
- from .lme_solver import compute_lme_critical_values
-
n, p = X_expanded.shape
n_targets = len(target_indices)
@@ -658,35 +653,27 @@ def _lme_analysis_custom_nested(
K_child = re_result.K_child
child_to_parent = re_result.child_to_parent
- if z_crit is None or chi2_crit is None or correction_z_crits is None:
- chi2_crit, z_crit, correction_z_crits = compute_lme_critical_values(alpha, p, n_targets, correction_method)
+ chi2_crit, z_crit, correction_z_crits = _ensure_lme_crits(alpha, p, n_targets, correction_method, chi2_crit, z_crit, correction_z_crits)
from mcpower.backends import mcpower_native as _native # type: ignore[attr-defined]
warm_theta_arr = np.empty(0, dtype=np.float64)
result = _native.lme_analysis_nested(
- np.ascontiguousarray(X_expanded, dtype=np.float64),
- np.ascontiguousarray(y, dtype=np.float64),
- np.ascontiguousarray(parent_ids, dtype=np.int32),
- np.ascontiguousarray(child_ids, dtype=np.int32),
+ _prep(X_expanded),
+ _prep(y),
+ _prep(parent_ids, np.int32),
+ _prep(child_ids, np.int32),
K_parent,
K_child,
- np.ascontiguousarray(child_to_parent, dtype=np.int32),
- np.ascontiguousarray(target_indices, dtype=np.int32),
+ _prep(child_to_parent, np.int32),
+ _prep(target_indices, np.int32),
float(chi2_crit),
float(z_crit),
- np.ascontiguousarray(correction_z_crits, dtype=np.float64),
+ _prep(correction_z_crits),
int(correction_method),
warm_theta_arr,
)
- if len(result) > 0:
- if verbose:
- return {"results": result, "diagnostics": {"solver": "native_nested", "K_parent": K_parent, "K_child": K_child}}
- return result # type: ignore[no-any-return]
-
- if verbose:
- return {"results": None, "failure_reason": "C++ nested solver returned empty result"}
- return None
+ return _wrap_native_result(result, verbose, "native_nested", extra_diag={"K_parent": K_parent, "K_child": K_child})
def reset_warm_start_cache():
diff --git a/mcpower/tables/lookup.py b/mcpower/tables/lookup.py
index b66fefa..5f4d691 100644
--- a/mcpower/tables/lookup.py
+++ b/mcpower/tables/lookup.py
@@ -15,7 +15,7 @@ class LookupTableManager:
"""Manages pre-computed lookup tables for data-generation transforms.
Tables are lazily loaded from disk (``tables/data/*.npz``) on first
- access and generated from scipy if the cache files are missing.
+ access and generated via the C++ native backend if the cache files are missing.
The C++ native backend consumes these tables for distribution
transforms.
@@ -47,47 +47,37 @@ def ensure_data_dir(self) -> None:
"""Ensure data directory exists."""
self.data_dir.mkdir(parents=True, exist_ok=True)
- def load_norm_cdf_table(self) -> np.ndarray:
- """Load (or generate and cache) the normal CDF lookup table.
+ def _load_table(self, key: str, generate_fn) -> np.ndarray:
+ """Load a table from cache, disk, or generate it on the fly.
+
+ Args:
+ key: Cache key and npz array name (e.g. ``"norm_cdf"``).
+ generate_fn: Bound method to generate and cache the table.
Returns:
1-D float64 array of length ``DIST_RESOLUTION``.
"""
- if "norm_cdf" in self._tables:
- return self._tables["norm_cdf"]
-
- cache_file = self.data_dir / "norm_cdf.npz"
+ if key in self._tables:
+ return self._tables[key]
+ cache_file = self.data_dir / f"{key}.npz"
try:
data = np.load(cache_file)
- self._tables["norm_cdf"] = data["norm_cdf"]
- return self._tables["norm_cdf"]
+ self._tables[key] = data[key]
+ return self._tables[key]
except (FileNotFoundError, KeyError):
pass
- self._generate_norm_cdf_table()
- return self._tables["norm_cdf"]
-
- def load_t3_ppf_table(self) -> np.ndarray:
- """Load (or generate and cache) the t(df=3) PPF lookup table.
+ generate_fn()
+ return self._tables[key]
- Returns:
- 1-D float64 array of length ``DIST_RESOLUTION``.
- """
- if "t3_ppf" in self._tables:
- return self._tables["t3_ppf"]
-
- cache_file = self.data_dir / "t3_ppf.npz"
-
- try:
- data = np.load(cache_file)
- self._tables["t3_ppf"] = data["t3_ppf"]
- return self._tables["t3_ppf"]
- except (FileNotFoundError, KeyError):
- pass
+ def load_norm_cdf_table(self) -> np.ndarray:
+ """Load (or generate and cache) the normal CDF lookup table."""
+ return self._load_table("norm_cdf", self._generate_norm_cdf_table)
- self._generate_t3_ppf_table()
- return self._tables["t3_ppf"]
+ def load_t3_ppf_table(self) -> np.ndarray:
+ """Load (or generate and cache) the t(df=3) PPF lookup table."""
+ return self._load_table("t3_ppf", self._generate_t3_ppf_table)
def load_all_generation_tables(self) -> Tuple[np.ndarray, np.ndarray]:
"""
@@ -110,6 +100,8 @@ def _generate_norm_cdf_table(self) -> None:
self._tables["norm_cdf"] = norm_cdf
self.ensure_data_dir()
+ # Silently ignore cache write failures (e.g. read-only filesystem,
+ # permission denied). Tables are still usable from memory.
try:
np.savez_compressed(self.data_dir / "norm_cdf.npz", norm_cdf=norm_cdf, x_range=x_norm)
except Exception:
@@ -125,6 +117,8 @@ def _generate_t3_ppf_table(self) -> None:
self._tables["t3_ppf"] = t3_ppf
self.ensure_data_dir()
+ # Silently ignore cache write failures (e.g. read-only filesystem,
+ # permission denied). Tables are still usable from memory.
try:
np.savez_compressed(
self.data_dir / "t3_ppf.npz",
diff --git a/mcpower/utils/formatters.py b/mcpower/utils/formatters.py
index b8bd0b0..8bc91b3 100644
--- a/mcpower/utils/formatters.py
+++ b/mcpower/utils/formatters.py
@@ -6,6 +6,7 @@
"""
import math
+from itertools import combinations
from typing import Any, Dict, List, Optional
import numpy as np
@@ -13,6 +14,11 @@
__all__ = []
+def _is_nan(value) -> bool:
+ """Check if a value is NaN (float type check + math.isnan)."""
+ return isinstance(value, float) and math.isnan(value)
+
+
class _TableFormatter:
"""Static helpers for building fixed-width text tables."""
@@ -25,7 +31,10 @@ def _create_table(
"""Create formatted table with headers and rows."""
if not col_widths:
- col_widths = [max(len(str(h)), max(len(str(row[i])) + 2 for row in rows)) for i, h in enumerate(headers)]
+ if rows:
+ col_widths = [max(len(str(h)), max(len(str(row[i])) + 2 for row in rows)) for i, h in enumerate(headers)]
+ else:
+ col_widths = [len(str(h)) for h in headers]
lines = []
@@ -131,7 +140,7 @@ def _format_short_power(self, data: Dict) -> str:
for test in model["target_tests"]:
power_corr = results["individual_powers_corrected"][test]
- if isinstance(power_corr, float) and math.isnan(power_corr):
+ if _is_nan(power_corr):
rows_corrected.append([test, "-", f"{target:.0f}", "-"])
else:
status = "✓" if power_corr >= target else "✗"
@@ -162,7 +171,7 @@ def _format_long_power(self, data: Dict) -> str:
power = results["individual_powers"][test]
power_corr = results.get("individual_powers_corrected", {}).get(test, power)
target = model.get("target_power", 80.0)
- if isinstance(power_corr, float) and math.isnan(power_corr):
+ if _is_nan(power_corr):
rows.append([test, f"{power:.2f}", "-", f"{target:.1f}", "-"])
else:
achieved = "✓" if power_corr >= target else "✗"
@@ -331,13 +340,15 @@ def _format_scenario_power_short(self, scenarios: Dict, target_tests: List[str],
lines = [f"\n{'=' * 80}", "SCENARIO SUMMARY", f"{'=' * 80}"]
+ scenario_names = list(scenarios.keys())
+ headers = ["Test"] + [name.title() for name in scenario_names]
+ col_widths = [40] + [12] * len(scenario_names)
+
# Uncorrected table
- headers = ["Test", "Optimistic", "Realistic", "Doomer"]
rows = []
-
for test in target_tests:
row = [test]
- for scenario in ["optimistic", "realistic", "doomer"]:
+ for scenario in scenario_names:
if scenario in scenarios and "results" in scenarios[scenario]:
power = scenarios[scenario]["results"]["individual_powers"][test]
row.append(f"{power:.1f}")
@@ -346,17 +357,17 @@ def _format_scenario_power_short(self, scenarios: Dict, target_tests: List[str],
rows.append(row)
lines.append("\nUncorrected Power:")
- lines.append(self._table._create_table(headers, rows, [40, 12, 12, 12]))
+ lines.append(self._table._create_table(headers, rows, col_widths))
# Corrected table if applicable
if correction:
rows_corr = []
for test in target_tests:
row = [test]
- for scenario in ["optimistic", "realistic", "doomer"]:
+ for scenario in scenario_names:
if scenario in scenarios and "results" in scenarios[scenario]:
power_corr = scenarios[scenario]["results"]["individual_powers_corrected"][test]
- if isinstance(power_corr, float) and math.isnan(power_corr):
+ if _is_nan(power_corr):
row.append("-")
else:
row.append(f"{power_corr:.1f}")
@@ -365,7 +376,7 @@ def _format_scenario_power_short(self, scenarios: Dict, target_tests: List[str],
rows_corr.append(row)
lines.append(f"\nCorrected Power ({correction}):")
- lines.append(self._table._create_table(headers, rows_corr, [40, 12, 12, 12]))
+ lines.append(self._table._create_table(headers, rows_corr, col_widths))
lines.append(f"{'=' * 80}")
@@ -395,74 +406,74 @@ def _format_scenario_power_long(
lines.append("DETAILED SCENARIO RESULTS")
lines.append(f"{'=' * 80}")
- for scenario_name in ["optimistic", "realistic", "doomer"]:
- if scenario_name in scenarios:
- lines.append(f"\n{'-' * 80}")
- lines.append(f"{scenario_name.upper()} SCENARIO")
- lines.append(f"{'-' * 80}")
-
- # Use regular power formatter for each scenario
- scenario_data = {
- "model": scenarios[scenario_name]["model"],
- "results": scenarios[scenario_name]["results"],
- }
- lines.append(self._format_long_power(scenario_data))
-
- # 3. Comparison analysis
- lines.append(f"\n{'=' * 80}")
- lines.append("ROBUSTNESS ANALYSIS")
- lines.append(f"{'=' * 80}")
-
- # Power reduction table
- headers = ["Test", "Opt→Real Drop", "Opt→Doom Drop", "Vulnerability"]
- rows = []
- vulnerable_tests = []
- inflated_tests = []
+ for scenario_name in scenarios:
+ lines.append(f"\n{'-' * 80}")
+ lines.append(f"{scenario_name.upper()} SCENARIO")
+ lines.append(f"{'-' * 80}")
+
+ scenario_data = {
+ "model": scenarios[scenario_name]["model"],
+ "results": scenarios[scenario_name]["results"],
+ }
+ lines.append(self._format_long_power(scenario_data))
+
+ # 3. Comparison analysis — compare each non-optimistic scenario to optimistic
+ if "optimistic" in scenarios and len(scenarios) > 1:
+ lines.append(f"\n{'=' * 80}")
+ lines.append("ROBUSTNESS ANALYSIS")
+ lines.append(f"{'=' * 80}")
+
+ other_scenarios = [s for s in scenarios if s != "optimistic"]
+ headers = ["Test"] + [f"Opt→{s.title()} Drop" for s in other_scenarios] + ["Vulnerability"]
+ rows = []
+ vulnerable_tests = []
+ inflated_tests = []
- for test in target_tests:
- opt_power = scenarios["optimistic"]["results"]["individual_powers"][test]
- real_power = scenarios.get("realistic", {}).get("results", {}).get("individual_powers", {}).get(test, opt_power)
- doom_power = scenarios.get("doomer", {}).get("results", {}).get("individual_powers", {}).get(test, opt_power)
-
- real_drop = opt_power - real_power
- doom_drop = opt_power - doom_power
-
- # Format drops with proper signs
- real_drop_str = f"+{abs(real_drop):.1f}%" if real_drop < 0 else f"-{real_drop:.1f}%"
- doom_drop_str = f"+{abs(doom_drop):.1f}%" if doom_drop < 0 else f"-{doom_drop:.1f}%"
-
- # Vulnerability assessment and categorization
- if doom_drop > HIGH_VULNERABILITY_THRESHOLD:
- vulnerability = "HIGH"
- vulnerable_tests.append(test)
- elif doom_drop > MEDIUM_VULNERABILITY_THRESHOLD:
- vulnerability = "MEDIUM"
- elif doom_drop < INFLATED_ERROR_THRESHOLD:
- vulnerability = "INFLATED FALSE POSITIVES"
- inflated_tests.append(test)
- else:
- vulnerability = "LOW"
+ for test in target_tests:
+ opt_power = scenarios["optimistic"]["results"]["individual_powers"][test]
+ row = [test]
+ max_drop = 0.0
+
+ for scenario in other_scenarios:
+ other_power = scenarios.get(scenario, {}).get("results", {}).get("individual_powers", {}).get(test, opt_power)
+ drop = opt_power - other_power
+ max_drop = max(max_drop, drop)
+ drop_str = f"+{abs(drop):.1f}%" if drop < 0 else f"-{drop:.1f}%"
+ row.append(drop_str)
+
+ if max_drop > HIGH_VULNERABILITY_THRESHOLD:
+ vulnerability = "HIGH"
+ vulnerable_tests.append(test)
+ elif max_drop > MEDIUM_VULNERABILITY_THRESHOLD:
+ vulnerability = "MEDIUM"
+ elif max_drop < INFLATED_ERROR_THRESHOLD:
+ vulnerability = "INFLATED FALSE POSITIVES"
+ inflated_tests.append(test)
+ else:
+ vulnerability = "LOW"
- rows.append([test, real_drop_str, doom_drop_str, vulnerability])
+ row.append(vulnerability)
+ rows.append(row)
- lines.append(self._table._create_table(headers, rows))
+ lines.append(self._table._create_table(headers, rows))
# 4. Recommendations
- lines.append(f"\n{'=' * 80}")
- lines.append("RECOMMENDATIONS")
- lines.append(f"{'=' * 80}")
+ if "optimistic" in scenarios and len(scenarios) > 1:
+ lines.append(f"\n{'=' * 80}")
+ lines.append("RECOMMENDATIONS")
+ lines.append(f"{'=' * 80}")
- if vulnerable_tests:
- lines.append(f"• High vulnerability tests: {', '.join(vulnerable_tests)}")
- lines.append("• Consider increasing sample size to maintain power under adverse conditions")
+ if vulnerable_tests:
+ lines.append(f"• High vulnerability tests: {', '.join(vulnerable_tests)}")
+ lines.append("• Consider increasing sample size to maintain power under adverse conditions")
- if inflated_tests:
- lines.append(f"• Inflated false positive risk: {', '.join(inflated_tests)}")
- lines.append("• Be careful about interpretation")
+ if inflated_tests:
+ lines.append(f"• Inflated false positive risk: {', '.join(inflated_tests)}")
+ lines.append("• Be careful about interpretation")
- if not vulnerable_tests and not inflated_tests:
- lines.append("• Power analysis appears robust to assumption violations")
- lines.append("• Original sample size should be sufficient")
+ if not vulnerable_tests and not inflated_tests:
+ lines.append("• Power analysis appears robust to assumption violations")
+ lines.append("• Original sample size should be sufficient")
return "\n".join(lines)
@@ -484,25 +495,22 @@ def _format_scenario_sample_size_short(self, scenarios: Dict, target_tests: List
"""Short scenario sample size summary."""
lines = [f"\n{'=' * 80}", "SCENARIO SUMMARY", f"{'=' * 80}"]
+ scenario_names = list(scenarios.keys())
if correction:
# Combined table with uncorrected and corrected
lines.append("\nSample Size Requirements:")
- headers = [
- "Test",
- "Opt(U)",
- "Opt(C)",
- "Real(U)",
- "Real(C)",
- "Doom(U)",
- "Doom(C)",
- ]
+ headers = ["Test"]
+ for name in scenario_names:
+ abbrev = name[:4].title()
+ headers.extend([f"{abbrev}(U)", f"{abbrev}(C)"])
+ col_widths = [40] + [8] * (len(scenario_names) * 2)
rows = []
for test in target_tests:
- row = [test[:40]] # Truncate to 40 chars
+ row = [test[:40]]
- for scenario in ["optimistic", "realistic", "doomer"]:
+ for scenario in scenario_names:
if scenario in scenarios and "results" in scenarios[scenario]:
n_uncorr = scenarios[scenario]["results"]["first_achieved"][test]
n_corr = scenarios[scenario]["results"]["first_achieved_corrected"][test]
@@ -520,16 +528,17 @@ def _format_scenario_sample_size_short(self, scenarios: Dict, target_tests: List
row.extend(["N/A", "N/A"])
rows.append(row)
- lines.append(self._table._create_table(headers, rows, [40, 8, 8, 8, 8, 8, 8]))
+ lines.append(self._table._create_table(headers, rows, col_widths))
lines.append("Note: (U) = Uncorrected, (C) = Corrected")
else:
# Uncorrected only
- headers = ["Test", "Optimistic", "Realistic", "Doomer"]
+ headers = ["Test"] + [name.title() for name in scenario_names]
+ col_widths = [40] + [12] * len(scenario_names)
rows = []
for test in target_tests:
- row = [test[:40]] # Truncate to 40 chars
- for scenario in ["optimistic", "realistic", "doomer"]:
+ row = [test[:40]]
+ for scenario in scenario_names:
if scenario in scenarios and "results" in scenarios[scenario]:
n_required = scenarios[scenario]["results"]["first_achieved"][test]
if n_required > 0:
@@ -542,7 +551,7 @@ def _format_scenario_sample_size_short(self, scenarios: Dict, target_tests: List
rows.append(row)
lines.append("\nUncorrected Sample Sizes:")
- lines.append(self._table._create_table(headers, rows, [40, 12, 12, 12]))
+ lines.append(self._table._create_table(headers, rows, col_widths))
lines.append(f"{'=' * 80}")
@@ -562,39 +571,33 @@ def _format_scenario_sample_size_long(
# 1. Overall summary
lines.append(self._format_scenario_sample_size_short(scenarios, target_tests, correction))
- # 2. Recommendations
+ # 2. Recommendations — summarize max N per non-optimistic scenario
lines.append(f"\n{'=' * 80}")
lines.append("RECOMMENDATIONS")
lines.append(f"{'=' * 80}")
- # Calculate max required N across scenarios
- max_n_realistic = max(
- (scenarios.get("realistic", {}).get("results", {}).get("first_achieved", {}).get(test, 0) for test in target_tests),
- default=0,
- )
- max_n_doomer = max(
- (scenarios.get("doomer", {}).get("results", {}).get("first_achieved", {}).get(test, 0) for test in target_tests),
- default=0,
- )
-
- max_tested = scenarios.get("realistic", {}).get("model", {}).get("sample_size_range", {}).get("to_size", 200)
-
- if max_n_realistic > 0 and max_n_realistic <= max_tested:
- lines.append(f"• For robust power under realistic conditions: N = {max_n_realistic}")
- elif max_n_realistic <= 0:
- lines.append(f"• For robust power under realistic conditions: N > {max_tested}")
-
- if max_n_doomer > 0 and max_n_doomer <= max_tested:
- lines.append(f"• For power under worst-case conditions: N = {max_n_doomer}")
- elif max_n_doomer <= 0:
- lines.append(f"• For power under worst-case conditions: N > {max_tested}")
-
- # Check if any tests couldn't achieve power
- unachievable = [
- test for test in target_tests if scenarios.get("doomer", {}).get("results", {}).get("first_achieved", {}).get(test, -1) <= 0
- ]
- if unachievable:
- lines.append(f"• Warning: These tests may not achieve target power under adverse conditions: {', '.join(unachievable)}")
+ other_scenarios = [s for s in scenarios if s != "optimistic"]
+ for scenario in other_scenarios:
+ max_n = max(
+ (scenarios.get(scenario, {}).get("results", {}).get("first_achieved", {}).get(test, 0) for test in target_tests),
+ default=0,
+ )
+ max_tested = scenarios.get(scenario, {}).get("model", {}).get("sample_size_range", {}).get("to_size", 200)
+ label = scenario.title()
+
+ if max_n > 0 and max_n <= max_tested:
+ lines.append(f"• For power under {label} conditions: N = {max_n}")
+ elif max_n <= 0:
+ lines.append(f"• For power under {label} conditions: N > {max_tested}")
+
+ # Check unachievable across worst scenario (last non-optimistic)
+ if other_scenarios:
+ worst = other_scenarios[-1]
+ unachievable = [
+ test for test in target_tests if scenarios.get(worst, {}).get("results", {}).get("first_achieved", {}).get(test, -1) <= 0
+ ]
+ if unachievable:
+ lines.append(f"• Warning: These tests may not achieve target power under {worst} conditions: {', '.join(unachievable)}")
# Add cumulative probability analysis
cumulative_lines = self._format_cumulative_recommendations(data, is_scenario=True)
@@ -706,7 +709,7 @@ def _add_cumulative_sample_size_table(
# Filter out tests with NaN power (e.g. non-contrast tests under Tukey correction)
def _has_nan_power(t: str) -> bool:
vals = powers_by_test[t]
- return bool(vals and isinstance(vals[0], float) and math.isnan(vals[0]))
+ return bool(vals and _is_nan(vals[0]))
valid_tests = [t for t in target_tests if not _has_nan_power(t)]
if not valid_tests:
@@ -741,8 +744,6 @@ def _has_nan_power(t: str) -> bool:
else: # ≥k cases
# Approximate using independence assumption
prob_at_least_k = 0.0
- from itertools import combinations
-
# Sum over all ways to choose at least k tests
for num_sig in range(k, n_tests + 1):
for combo in combinations(range(n_tests), num_sig):
@@ -859,6 +860,12 @@ def _format_cumulative_recommendations(self, results: Dict, is_scenario: bool =
if prob >= target_power:
min_n_target = sample_sizes[i]
break
+
+ if min_n_target:
+ lines.append(f"• N={min_n_target} for {target_power:.0f}% chance all tests significant")
+ else:
+ max_tested = sample_sizes[-1]
+ lines.append(f"• >{max_tested} needed for {target_power:.0f}% chance all tests significant")
return lines
diff --git a/mcpower/utils/parsers.py b/mcpower/utils/parsers.py
index e89d140..c14533f 100644
--- a/mcpower/utils/parsers.py
+++ b/mcpower/utils/parsers.py
@@ -105,6 +105,8 @@ def _split_assignments(self, input_string: str) -> List[str]:
paren_count += 1
elif char == ")":
paren_count -= 1
+ if paren_count < 0:
+ raise ValueError("Unbalanced parentheses: unexpected ')'")
current.append(char)
if current:
@@ -424,7 +426,11 @@ def _parse_independent_variables(formula: str) -> Tuple[Dict, Dict]:
"""
from itertools import combinations
- terms = re.split(r"[+\-]", formula)
+ # Check for minus sign (term removal) which is not supported
+ if re.search(r"(? Tuple[List[str], List[Dict]]:
+ """Extract effect names from a test formula, matched against the registry.
+
+ Parses the test formula, expands factor variables to their dummies,
+ and returns the list of effect names (in registry order) that belong
+ to the test formula.
+
+ Args:
+ test_formula: Formula string (e.g. ``"y ~ x1 + x2"``).
+ registry: ``VariableRegistry`` instance.
+
+ Returns:
+ Tuple of ``(effect_names, random_effects)`` where *effect_names*
+ are the registry effect names present in the test formula (in
+ registry order), and *random_effects* is the list of parsed
+ random-effect dicts from the test formula.
+ """
+ _dep_var, fixed_formula, random_effects = _parse_equation(test_formula)
+
+ # Parse fixed effects into a set of term names
+ test_terms = _parse_fixed_terms(fixed_formula)
+
+ # Determine which registry effects belong to the test formula
+ cluster_effects = set(registry.cluster_effect_names)
+ test_effects: List[str] = []
+
+ for effect_name in registry._effects:
+ if effect_name in cluster_effects:
+ continue
+
+ effect = registry._effects[effect_name]
+
+ if effect.effect_type == "main":
+ # Direct match (continuous or interaction-less variable)
+ if effect_name in test_terms:
+ test_effects.append(effect_name)
+ elif effect_name in registry._factor_dummies:
+ # Factor dummy -- include if parent factor is in test terms
+ parent_factor = registry._factor_dummies[effect_name]["factor_name"]
+ if parent_factor in test_terms:
+ test_effects.append(effect_name)
+ else:
+ # Interaction -- check if the interaction term is in test terms
+ if effect_name in test_terms:
+ test_effects.append(effect_name)
+
+ return test_effects, random_effects
+
+
+def _parse_fixed_terms(fixed_formula: str) -> Set[str]:
+ """Parse a fixed-effect formula string into a set of term names.
+
+ Handles ``+`` for additive terms, ``:`` for specific interactions,
+ and ``*`` for full factorial expansion (main effects plus all
+ two-way through n-way interactions).
+
+ Args:
+ fixed_formula: Right-hand side of the equation, spaces already
+ stripped by ``_parse_equation`` (e.g. ``"x1+x2+x1:x2"``).
+
+ Returns:
+ Set of term names (variable names and interaction terms like
+ ``"x1:x2"``).
+ """
+ if not fixed_formula.strip():
+ return set()
+
+ terms: Set[str] = set()
+ raw_terms = re.split(r"\+", fixed_formula)
+
+ for raw in raw_terms:
+ raw = raw.strip()
+ if not raw:
+ continue
+
+ if "*" in raw:
+ # Full factorial: x1*x2 -> x1, x2, x1:x2
+ vars_in_star = [v.strip() for v in raw.split("*") if v.strip()]
+ for v in vars_in_star:
+ terms.add(v)
+ for r in range(2, len(vars_in_star) + 1):
+ for combo in combinations(vars_in_star, r):
+ terms.add(":".join(combo))
+ else:
+ # Plain term (may contain ":" for explicit interaction)
+ terms.add(raw)
+
+ return terms
+
+
+def _compute_test_column_indices(
+ all_effect_names: List[str],
+ test_effect_names: List[str],
+) -> np.ndarray:
+ """Compute column indices in X_expanded for test formula effects.
+
+ Args:
+ all_effect_names: All non-cluster effect names in registry order.
+ test_effect_names: Effect names present in the test formula
+ (a subset of *all_effect_names*).
+
+ Returns:
+ Integer array of column indices into X_expanded.
+ """
+ test_set = set(test_effect_names)
+ indices = [i for i, name in enumerate(all_effect_names) if name in test_set]
+ return np.array(indices, dtype=np.int64)
+
+
+def _remap_target_indices(
+ original_target_indices: np.ndarray,
+ test_column_indices: np.ndarray,
+) -> np.ndarray:
+ """Remap target indices from full X_expanded space to X_test space.
+
+ Args:
+ original_target_indices: Indices in X_expanded being tested.
+ test_column_indices: Columns of X_expanded included in X_test.
+
+ Returns:
+ Indices remapped to positions within X_test.
+ """
+ # Build mapping: full_index -> position in X_test
+ index_map = {int(full_idx): test_idx for test_idx, full_idx in enumerate(test_column_indices)}
+ return np.array(
+ [index_map[int(idx)] for idx in original_target_indices],
+ dtype=np.int64,
+ )
diff --git a/mcpower/utils/updates.py b/mcpower/utils/updates.py
index 8c3c76b..c7f57a5 100644
--- a/mcpower/utils/updates.py
+++ b/mcpower/utils/updates.py
@@ -12,6 +12,8 @@
from datetime import datetime, timedelta
from pathlib import Path
+_already_checked = False
+
def _check_for_updates(current_version):
"""Check PyPI weekly for a newer MCPower version and warn if found.
@@ -20,18 +22,24 @@ def _check_for_updates(current_version):
silently in worker processes (detected via environment variable)
and in frozen (PyInstaller) bundles where pip is unavailable.
"""
+ global _already_checked
# Skip in frozen bundles (PyInstaller) — the GUI has its own update checker
if getattr(sys, "frozen", False):
return
+ # Skip if already checked in this process
+ if _already_checked:
+ return
+
# Skip in worker processes (loky/joblib inherit env vars from parent)
if os.environ.get("_MCPOWER_UPDATE_CHECKED"):
return
os.environ["_MCPOWER_UPDATE_CHECKED"] = "1"
+ _already_checked = True
- cache_path = Path(__file__).parent.parent / ".mcpower_cache.json"
- cache_path.parent.mkdir(exist_ok=True)
+ cache_path = Path.home() / ".cache" / "mcpower" / "update_cache.json"
+ cache_path.parent.mkdir(parents=True, exist_ok=True)
# Load cache
cache = {}
@@ -57,9 +65,8 @@ def _check_for_updates(current_version):
# Show update message only when PyPI version is strictly newer
latest = cache.get("latest_version")
- current = cache.get("current_version")
- if latest and current and _is_newer(latest, current):
- msg = f"\nNEW MCPower VERSION AVAILABLE: {latest} (you have {current})\nUpdate now: pip install --upgrade MCPower\n"
+ if latest and _is_newer(latest, current_version):
+ msg = f"\nNEW MCPower VERSION AVAILABLE: {latest} (you have {current_version})\nUpdate now: pip install --upgrade MCPower\n"
warnings.warn(msg, stacklevel=3)
@@ -77,7 +84,10 @@ def _get_latest_version():
"""Fetch the latest MCPower version string from the PyPI JSON API."""
try:
with urllib.request.urlopen("https://pypi.org/pypi/MCPower/json", timeout=5) as response:
- data = json.loads(response.read())
+ raw = response.read(1_000_000)
+ if len(raw) >= 1_000_000:
+ return None
+ data = json.loads(raw)
return data["info"]["version"]
except Exception:
return None
diff --git a/mcpower/utils/validators.py b/mcpower/utils/validators.py
index 5853af6..1c344fd 100644
--- a/mcpower/utils/validators.py
+++ b/mcpower/utils/validators.py
@@ -27,6 +27,11 @@ class _ValidationResult:
errors: List[str]
warnings: List[str]
+ @classmethod
+ def from_errors(cls, errors: List[str], warnings: Optional[List[str]] = None) -> "_ValidationResult":
+ """Create a result from error/warning lists, deriving ``is_valid`` automatically."""
+ return cls(len(errors) == 0, errors, warnings or [])
+
def raise_if_invalid(self):
"""Raise ``ValueError`` if the validation failed."""
if not self.is_valid:
@@ -88,12 +93,12 @@ def _validate_numeric_parameter(
errors.append(range_error)
# Rounding warning for floats when int expected
- if allow_rounding and isinstance(value, float) and (int, float) in expected_types:
+ if allow_rounding and isinstance(value, float) and int in expected_types:
rounded = int(round(value))
if value != rounded:
warnings.append(f"{name} rounded from {value} to {rounded}")
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
def _validate_power(power: Any) -> _ValidationResult:
@@ -112,6 +117,8 @@ def _validate_simulations(n_simulations: Any) -> Tuple[int, _ValidationResult]:
if result.is_valid:
rounded = int(round(n_simulations))
+ # 800 simulations threshold: below this, Monte Carlo standard error
+ # exceeds ~1.5% for power near 50%, reducing result reliability.
if rounded < 800:
result.warnings.append(f"Low simulation count ({rounded}). Consider using at least 1000 for reliable results.")
return rounded, result
@@ -139,7 +146,7 @@ def _validate_sample_size(sample_size: Any) -> _ValidationResult:
f"sample_size too large ({sample_size:,}). Maximum recommended: 100,000. We cannot guarantee stability for such small p-values."
)
- return _ValidationResult(len(errors) == 0, errors, [])
+ return _ValidationResult.from_errors(errors)
def _validate_sample_size_for_model(sample_size: int, n_variables: int) -> _ValidationResult:
@@ -157,6 +164,8 @@ def _validate_sample_size_for_model(sample_size: int, n_variables: int) -> _Vali
_ValidationResult with errors if sample size is insufficient.
"""
errors = []
+ # Green's rule of thumb: N >= 15 + p for adequate power in regression,
+ # where p is the number of predictors (design matrix columns).
min_required = 15 + n_variables
if sample_size < min_required:
@@ -165,7 +174,7 @@ def _validate_sample_size_for_model(sample_size: int, n_variables: int) -> _Vali
f"variables. Minimum required: {min_required} (15 + {n_variables} variables)."
)
- return _ValidationResult(len(errors) == 0, errors, [])
+ return _ValidationResult.from_errors(errors)
def _validate_sample_size_range(from_size: Any, to_size: Any, by: Any) -> _ValidationResult:
@@ -193,7 +202,7 @@ def _validate_sample_size_range(from_size: Any, to_size: Any, by: Any) -> _Valid
if n_tests > 100:
warnings.append(f"Large number of sample sizes to test ({n_tests}). This may take significant time.")
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
def _validate_correlation_matrix(
@@ -226,12 +235,14 @@ def _validate_correlation_matrix(
# Positive semi-definite check
try:
eigenvals = np.linalg.eigvals(corr_matrix)
+ # -1e-8 tolerance for positive semi-definiteness: allows small negative
+ # eigenvalues from floating-point rounding in correlation matrices.
if np.any(eigenvals < -1e-8): # Tolerance for floating point noise
errors.append("Correlation matrix must be positive semi-definite. ")
except np.linalg.LinAlgError:
errors.append("Cannot compute eigenvalues of correlation matrix")
- return _ValidationResult(len(errors) == 0, errors, [])
+ return _ValidationResult.from_errors(errors)
def _validate_correction_method(correction: Optional[str]) -> _ValidationResult:
@@ -285,7 +296,7 @@ def _validate_parallel_settings(enable: Any, n_cores: Optional[int]) -> Tuple[Tu
else:
validated_n_cores = min(n_cores, max_cores)
- return (enable, validated_n_cores), _ValidationResult(len(errors) == 0, errors, [])
+ return (enable, validated_n_cores), _ValidationResult.from_errors(errors)
def _validate_model_ready(model) -> _ValidationResult:
@@ -301,9 +312,10 @@ def _validate_model_ready(model) -> _ValidationResult:
errors: List[str] = []
warnings: List[str] = []
- # Check effect sizes - check if pending effects were set
- has_effects = hasattr(model, "_pending_effects") and model._pending_effects is not None
- if not has_effects:
+ # Check effect sizes — pending (pre-apply) or flagged as set by user
+ has_pending = hasattr(model, "_pending_effects") and model._pending_effects is not None
+ has_set = hasattr(model, "_effects_set") and model._effects_set
+ if not has_pending and not has_set:
if hasattr(model, "_registry"):
available = model._registry.effect_names
errors.append(
@@ -318,7 +330,7 @@ def _validate_model_ready(model) -> _ValidationResult:
if not hasattr(model, attr):
errors.append(f"Model missing required attribute: {attr}")
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
def _validate_test_formula(test_formula: str, available_variables: List[str]) -> _ValidationResult:
@@ -361,7 +373,7 @@ def _validate_test_formula(test_formula: str, available_variables: List[str]) ->
f"Variables not found in original model: {', '.join(sorted(missing_vars))}. Available: {', '.join(available_variables)}"
)
- return _ValidationResult(len(errors) == 0, errors, [])
+ return _ValidationResult.from_errors(errors)
except Exception as e:
errors.append(f"Error parsing test_formula: {str(e)}")
@@ -399,6 +411,8 @@ def _validate_factor_specification(n_levels: int, proportions: List[float]) -> _
# Check if they sum to approximately 1
if not errors: # Only if no errors with individual proportions
total = sum(proportions)
+ # 1e-6 tolerance: proportions are normalized later, so small deviations
+ # from 1.0 are acceptable and only warrant a warning.
if abs(total - 1.0) > 1e-6:
warnings.append(f"Proportions sum to {total:.4f}, not 1.0 (will be normalized)")
@@ -406,7 +420,7 @@ def _validate_factor_specification(n_levels: int, proportions: List[float]) -> _
if n_levels > 10:
warnings.append(f"Factor has {n_levels} levels. This creates {n_levels - 1} dummy variables, which may require large sample sizes")
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
def _validate_upload_data(data: np.ndarray) -> _ValidationResult:
@@ -425,7 +439,7 @@ def _validate_upload_data(data: np.ndarray) -> _ValidationResult:
if data.shape[0] < 25:
errors.append(f"Need at least 25 samples for reliable quantile matching, got {data.shape[0]}")
- return _ValidationResult(len(errors) == 0, errors, [])
+ return _ValidationResult.from_errors(errors)
def _validate_cluster_config(
@@ -475,7 +489,7 @@ def _validate_cluster_config(
if not isinstance(cluster_size, int) or cluster_size < 5:
errors.append(f"cluster_size must be an integer >= 5 for reliable mixed model estimation. Got {cluster_size}.")
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
def _validate_cluster_sample_size(
@@ -517,4 +531,4 @@ def _validate_cluster_sample_size(
f"Small cluster sizes may cause convergence issues or biased variance estimates."
)
- return _ValidationResult(len(errors) == 0, errors, warnings)
+ return _ValidationResult.from_errors(errors, warnings)
diff --git a/mcpower/utils/visualization.py b/mcpower/utils/visualization.py
index 22544a2..ab7eadd 100644
--- a/mcpower/utils/visualization.py
+++ b/mcpower/utils/visualization.py
@@ -18,6 +18,7 @@ def _create_power_plot(
target_tests: List[str],
target_power: float,
title: str,
+ show: bool = True,
):
"""Create a sample-size vs. power line plot with achievement markers.
@@ -58,7 +59,7 @@ def _create_power_plot(
)
# Mark achievement point
- if first_achieved[test] > 0:
+ if first_achieved[test] > 0 and first_achieved[test] in sample_sizes:
achieved_idx = sample_sizes.index(first_achieved[test])
achieved_power = powers[achieved_idx]
ax.plot(
@@ -112,4 +113,6 @@ def _create_power_plot(
color="#888888",
)
plt.tight_layout(rect=(0, 0.03, 1, 1))
- plt.show()
+ if show:
+ plt.show()
+ return fig
diff --git a/pyproject.toml b/pyproject.toml
index 983e3d3..ec1f41f 100644
--- a/pyproject.toml
+++ b/pyproject.toml
@@ -1,14 +1,14 @@
[build-system]
requires = [
- "scikit-build-core>=0.5",
+ "scikit-build-core>=0.10",
"pybind11>=2.11",
- "numpy>=2.0.0",
+ "numpy>=1.26.0",
]
build-backend = "scikit_build_core.build"
[project]
name = "MCPower"
-version = "0.5.4"
+version = "0.6.0"
description = "Monte Carlo Power Analysis for Statistical Models"
readme = "README.md"
license = {text = "GPL-3.0-or-later"}
@@ -31,9 +31,10 @@ classifiers = [
]
requires-python = ">=3.10"
dependencies = [
- "numpy>=2.0.0",
+ "numpy>=1.26.0",
"matplotlib>=3.8.0",
"joblib>=1.3.0",
+ "tqdm>=4.60.0",
]
[project.optional-dependencies]
@@ -41,6 +42,7 @@ lme = ["statsmodels>=0.14.0"]
pandas = ["pandas>=2.0.0"]
dev = [
"pandas>=2.0.0",
+ "statsmodels>=0.14.0",
"pytest>=7.0.0",
"pytest-cov>=4.0.0",
"scipy>=1.11.0",
@@ -52,14 +54,14 @@ dev = [
]
all = [
"pandas>=2.0.0",
- "statsmodels>=0.14.0",
-]
+ ]
[project.urls]
Homepage = "https://github.com/pawlenartowicz/MCPower"
-Documentation = "https://github.com/pawlenartowicz/MCPower#readme"
+Documentation = "https://github.com/pawlenartowicz/MCPower/wiki"
Repository = "https://github.com/pawlenartowicz/MCPower"
Issues = "https://github.com/pawlenartowicz/MCPower/issues"
+Changelog = "https://github.com/pawlenartowicz/MCPower/blob/main/CHANGELOG.md"
[tool.scikit-build]
wheel.packages = ["mcpower"]
@@ -77,8 +79,6 @@ python_files = ["test_*.py"]
python_classes = ["Test*"]
python_functions = ["test_*"]
markers = [
- "unit: Unit tests",
- "integration: Integration tests",
"lme: LME mixed-effects model tests",
]
addopts = "-v --tb=short --strict-markers"
@@ -86,7 +86,6 @@ filterwarnings = [
"ignore::FutureWarning",
"ignore::DeprecationWarning",
"ignore::UserWarning:statsmodels",
- "ignore:Mixed-effects models are experimental:UserWarning",
]
[tool.ruff]
@@ -107,6 +106,20 @@ known-first-party = ["mcpower"]
python_version = "3.10"
warn_return_any = true
warn_unused_configs = true
-ignore_missing_imports = true
check_untyped_defs = true
exclude = ["build", "dist", "tests"]
+
+[[tool.mypy.overrides]]
+module = [
+ "mcpower_native",
+ "mcpower_native.*",
+ "statsmodels",
+ "statsmodels.*",
+ "tqdm",
+ "tqdm.*",
+ "joblib",
+ "joblib.*",
+ "pandas",
+ "pandas.*",
+]
+ignore_missing_imports = true
diff --git a/tests/config.py b/tests/config.py
index 63c1999..a8b874c 100644
--- a/tests/config.py
+++ b/tests/config.py
@@ -5,9 +5,21 @@
across the test suite.
"""
-# Monte Carlo simulation parameters
-N_SIMS = 5000
-"""Number of Monte Carlo simulations for power analysis tests."""
+# Monte Carlo simulation parameters — 4-tier ladder
+N_SIMS_CHECK = 50
+"""Smoke tests — just verify no crash, structure, API contract."""
+
+N_SIMS_ORDERING = 1000
+"""Ordering tests — monotonicity, correction hierarchy, A < B checks."""
+
+N_SIMS_STANDARD = 1600
+"""Standard tests — null calibration, Type I error, general validation."""
+
+N_SIMS_ACCURACY = 5000
+"""Accuracy tests — comparison against analytical power formulas."""
+
+N_SIMS = N_SIMS_ACCURACY
+"""Backward-compat alias for accuracy-level simulations."""
SEED = 2137
"""Default random seed for reproducibility."""
diff --git a/tests/conftest.py b/tests/conftest.py
index 93c8814..d2100b5 100644
--- a/tests/conftest.py
+++ b/tests/conftest.py
@@ -63,12 +63,6 @@ def correlation_matrix_2x2():
return np.array([[1.0, 0.5], [0.5, 1.0]])
-@pytest.fixture
-def correlation_matrix_3x3():
- """Create a 3x3 correlation matrix."""
- return np.array([[1.0, 0.3, 0.2], [0.3, 1.0, 0.4], [0.2, 0.4, 1.0]])
-
-
@pytest.fixture
def sample_data():
"""Create sample empirical data."""
@@ -80,41 +74,13 @@ def sample_data():
@pytest.fixture
-def suppress_output(capsys):
- """Suppress print output during tests by capturing it."""
- yield
- # Output is automatically captured by capsys
-
-
-BACKENDS = ["c++"]
-
-
-@pytest.fixture(params=BACKENDS)
-def backend(request):
- """
- Force MCPower to run on a specific backend.
-
- Parametrizes tests against C++ (primary backend).
- Automatically resets backend after each test.
- """
- from mcpower.backends import reset_backend, set_backend
-
- set_backend(request.param)
- yield request.param
- reset_backend()
-
-
-@pytest.fixture(autouse=True)
-def reset_backend_after_test():
- """
- Automatically reset backend to default after every test.
-
- Ensures no hidden backend state leaks between tests.
- """
- yield
- from mcpower.backends import reset_backend
+def suppress_output():
+ """Suppress print output during tests."""
+ import contextlib
+ import io
- reset_backend()
+ with contextlib.redirect_stdout(io.StringIO()):
+ yield
def _statsmodels_available():
diff --git a/tests/helpers/power_helpers.py b/tests/helpers/power_helpers.py
index d509a79..620e995 100644
--- a/tests/helpers/power_helpers.py
+++ b/tests/helpers/power_helpers.py
@@ -40,42 +40,3 @@ def compute_crits(X, target_indices, alpha=DEFAULT_ALPHA, correction_method=0):
return compute_critical_values(alpha, p, dof, n_targets, correction_method)
-def run_with_backend(
- backend_name,
- equation,
- effects_str,
- sample_size,
- n_sims,
- seed,
- target_test="all",
- correction=None,
- correlations_str=None,
- alpha=DEFAULT_ALPHA,
-):
- """Run a full MCPower power analysis with a specific backend forced."""
- import contextlib
- import io
-
- from mcpower import MCPower
- from mcpower.backends import reset_backend, set_backend
-
- set_backend(backend_name)
- try:
- m = MCPower(equation)
- m.set_simulations(n_sims)
- m.set_seed(seed)
- m.set_alpha(alpha)
- m.set_effects(effects_str)
- if correlations_str:
- m.set_correlations(correlations_str)
- with contextlib.redirect_stdout(io.StringIO()):
- result = m.find_power(
- sample_size=sample_size,
- target_test=target_test,
- correction=correction,
- print_results=False,
- return_results=True,
- )
- finally:
- reset_backend()
- return result
diff --git a/tests/integration/test_find_power_api.py b/tests/integration/test_find_power_api.py
index 69f6bc5..fe56de3 100644
--- a/tests/integration/test_find_power_api.py
+++ b/tests/integration/test_find_power_api.py
@@ -222,14 +222,14 @@ def test_all_targets(self, suppress_output):
class TestHeterogeneity:
- """Test heterogeneity settings."""
+ """Test heterogeneity via scenario configs."""
def test_with_heterogeneity(self, suppress_output):
from mcpower import MCPower
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
- model.set_heterogeneity(0.1)
+ model.set_scenario_configs({"het": {"heterogeneity": 0.1}})
result = model.find_power(100, print_results=False, return_results=True)
assert result is not None
@@ -239,7 +239,7 @@ def test_with_heteroskedasticity(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
- model.set_heteroskedasticity(0.2)
+ model.set_scenario_configs({"hsked": {"heteroskedasticity": 0.2}})
result = model.find_power(100, print_results=False, return_results=True)
assert result is not None
@@ -249,8 +249,7 @@ def test_combined(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
- model.set_heterogeneity(0.1)
- model.set_heteroskedasticity(0.2)
+ model.set_scenario_configs({"combo": {"heterogeneity": 0.1, "heteroskedasticity": 0.2}})
result = model.find_power(100, print_results=False, return_results=True)
assert result is not None
@@ -330,7 +329,7 @@ def test_all_features_combined(self, suppress_output):
model.upload_data({"x1": np.random.exponential(2, 100)})
model.set_correlations("(x1,x2)=0.3")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2, x2=0.15, x1:x2=0.1")
- model.set_heterogeneity(0.05)
+ model.set_scenario_configs({"test": {"heterogeneity": 0.05}})
result = model.find_power(200, print_results=False, return_results=True)
assert result is not None
diff --git a/tests/integration/test_model.py b/tests/integration/test_model.py
index da00d33..a57b051 100644
--- a/tests/integration/test_model.py
+++ b/tests/integration/test_model.py
@@ -86,16 +86,6 @@ def test_set_variable_type(self, suppress_output):
assert model._pending_variable_types == "group=(factor,3)"
assert model._applied is False
- def test_set_heterogeneity(self, simple_model):
- simple_model.set_heterogeneity(0.1)
- assert simple_model._pending_heterogeneity == 0.1
- assert simple_model._applied is False
-
- def test_set_heteroskedasticity(self, simple_model):
- simple_model.set_heteroskedasticity(0.2)
- assert simple_model._pending_heteroskedasticity == 0.2
- assert simple_model._applied is False
-
def test_upload_data_dict(self, simple_model, sample_data):
simple_model.upload_data(sample_data)
assert simple_model._pending_data is not None
@@ -131,12 +121,12 @@ class TestApply:
"""Test apply() method."""
def test_apply_sets_flag(self, configured_model):
- configured_model.apply()
+ configured_model._apply()
assert configured_model._applied is True
def test_apply_processes_effects(self, simple_model):
simple_model.set_effects("x1=0.5, x2=0.3")
- simple_model.apply()
+ simple_model._apply()
effect_sizes = simple_model._registry.get_effect_sizes()
assert effect_sizes[0] == 0.5
assert effect_sizes[1] == 0.3
@@ -147,22 +137,17 @@ def test_apply_processes_variable_types(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
assert len(model._registry.factor_names) == 1
assert len(model._registry.dummy_names) == 2
def test_apply_processes_correlations(self, simple_model):
simple_model.set_effects("x1=0.3, x2=0.2")
simple_model.set_correlations("(x1,x2)=0.5")
- simple_model.apply()
+ simple_model._apply()
corr = simple_model.correlation_matrix
assert corr[0, 1] == 0.5
- def test_apply_processes_heterogeneity(self, configured_model):
- configured_model.set_heterogeneity(0.15)
- configured_model.apply()
- assert configured_model.heterogeneity == 0.15
-
def test_apply_order_independence(self, suppress_output):
"""Test that set_* methods can be called in any order."""
from mcpower import MCPower
@@ -172,14 +157,14 @@ def test_apply_order_independence(self, suppress_output):
m1.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2, x2=0.1")
m1.set_variable_type("group=(factor,3)")
m1.set_correlations("(x1,x2)=0.5")
- m1.apply()
+ m1._apply()
# Order 2: variable_type, correlations, effects
m2 = MCPower("y = group + x1 + x2")
m2.set_variable_type("group=(factor,3)")
m2.set_correlations("(x1,x2)=0.5")
m2.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2, x2=0.1")
- m2.apply()
+ m2._apply()
# Both should have same effect sizes
assert np.allclose(m1._registry.get_effect_sizes(), m2._registry.get_effect_sizes())
@@ -242,7 +227,7 @@ def test_sample_sizes_tested(self, configured_model):
assert result["results"]["sample_sizes_tested"] == [50, 75, 100]
def test_first_achieved(self, configured_model):
- result = configured_model.find_sample_size(from_size=50, to_size=200, by=25, print_results=False, return_results=True)
+ result = configured_model.find_sample_size(from_size=50, to_size=200, by=50, print_results=False, return_results=True)
assert "first_achieved" in result["results"]
def test_find_sample_size_runs(self, configured_model):
@@ -257,7 +242,7 @@ class TestErrors:
def test_invalid_effect_name(self, simple_model):
simple_model.set_effects("invalid=0.3")
with pytest.raises(ValueError, match="not found"):
- simple_model.apply()
+ simple_model._apply()
def test_missing_effects(self, simple_model):
with pytest.raises(ValueError, match="Effect sizes must be set"):
@@ -290,7 +275,7 @@ def test_basic_named_levels(self):
model = MCPower("y = treatment + x1")
model.set_factor_levels("treatment=placebo,drug_a,drug_b")
model.set_effects("treatment[drug_a]=0.5, treatment[drug_b]=0.8, x1=0.3")
- model.apply()
+ model._apply()
assert "treatment" in model._registry.factor_names
assert "treatment[drug_a]" in model._registry.dummy_names
assert "treatment[drug_b]" in model._registry.dummy_names
@@ -302,7 +287,7 @@ def test_multiple_factors(self):
model = MCPower("y = group + dose")
model.set_factor_levels("group=control,treatment; dose=low,medium,high")
model.set_effects("group[treatment]=0.5, dose[medium]=0.3, dose[high]=0.6")
- model.apply()
+ model._apply()
assert "group[treatment]" in model._registry.dummy_names
assert "dose[medium]" in model._registry.dummy_names
assert "dose[high]" in model._registry.dummy_names
@@ -313,7 +298,7 @@ def test_unknown_variable_raises(self):
model = MCPower("y = x1")
with pytest.raises(ValueError, match="not found"):
model.set_factor_levels("unknown=a,b,c")
- model.apply()
+ model._apply()
def test_single_level_raises(self):
from mcpower import MCPower
@@ -321,7 +306,7 @@ def test_single_level_raises(self):
model = MCPower("y = x1")
with pytest.raises(ValueError, match="at least 2"):
model.set_factor_levels("x1=only_one")
- model.apply()
+ model._apply()
def test_find_power_with_named_levels(self):
"""End-to-end: find_power works with set_factor_levels."""
diff --git a/tests/integration/test_parallel.py b/tests/integration/test_parallel.py
index 682372c..2048d7e 100644
--- a/tests/integration/test_parallel.py
+++ b/tests/integration/test_parallel.py
@@ -4,6 +4,8 @@
import pytest
+from tests.config import N_SIMS_CHECK
+
def _joblib_available():
"""Check if joblib is available."""
@@ -26,6 +28,7 @@ def test_parallel_results_match_sequential(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
model.set_seed(42)
+ model.set_simulations(N_SIMS_CHECK)
# Run sequential analysis
model.set_parallel(False)
@@ -56,6 +59,7 @@ def test_parallel_with_scenarios(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
model.set_seed(42)
+ model.set_simulations(N_SIMS_CHECK)
# Run sequential with scenarios
model.set_parallel(False)
@@ -93,6 +97,7 @@ def test_parallel_with_interactions(self, suppress_output):
model = MCPower("y = a + b + a:b")
model.set_effects("a=0.4, b=0.3, a:b=0.2")
model.set_seed(42)
+ model.set_simulations(N_SIMS_CHECK)
# Run sequential
model.set_parallel(False)
@@ -128,6 +133,7 @@ def test_parallel_fallback_on_failure(self, suppress_output, monkeypatch):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
model.set_seed(42)
+ model.set_simulations(N_SIMS_CHECK)
model.set_parallel(True, n_cores=2)
# Mock joblib.Parallel to raise an exception
@@ -157,6 +163,7 @@ def test_find_power_ignores_parallel(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.3, x2=0.2")
model.set_seed(42)
+ model.set_simulations(N_SIMS_CHECK)
# Run with parallel=False
model.set_parallel(False)
diff --git a/tests/integration/test_posthoc_integration.py b/tests/integration/test_posthoc_integration.py
index 9499388..b6b7f21 100644
--- a/tests/integration/test_posthoc_integration.py
+++ b/tests/integration/test_posthoc_integration.py
@@ -15,7 +15,7 @@ def test_parse_vs_syntax(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("group[1] vs group[2]")
assert "group[1] vs group[2]" in tests
@@ -27,7 +27,7 @@ def test_parse_multiple_vs(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("group[1] vs group[2], group[2] vs group[3]")
assert "group[1] vs group[2]" in tests
@@ -40,7 +40,7 @@ def test_parse_mixed_regular_and_posthoc(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("overall, group[1] vs group[2]")
assert "overall" in tests
@@ -52,7 +52,7 @@ def test_all_does_not_include_posthoc(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all")
# "all" should NOT include any post-hoc comparisons
@@ -66,7 +66,7 @@ def test_invalid_factor_name(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="Factor.*not found"):
model._parse_target_tests("notafactor[1] vs notafactor[2]")
@@ -77,7 +77,7 @@ def test_invalid_level(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="out of range"):
model._parse_target_tests("group[0] vs group[5]")
@@ -88,7 +88,7 @@ def test_same_level_comparison_rejected(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="Cannot compare a level to itself"):
model._parse_target_tests("group[2] vs group[2]")
@@ -99,7 +99,7 @@ def test_cross_factor_comparison_rejected(self, suppress_output):
model = MCPower("y = a + b")
model.set_variable_type("a=(factor,3), b=(factor,2)")
model.set_effects("a[2]=0.3, a[3]=0.2, b[2]=0.1")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="same factor"):
model._parse_target_tests("a[1] vs b[1]")
@@ -396,7 +396,7 @@ def test_all_posthoc_keyword(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all-posthoc")
# 3-level factor → C(3,2) = 3 pairs
@@ -415,7 +415,7 @@ def test_all_plus_all_posthoc(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all, all-posthoc")
# "all" → overall + group[2] + group[3] + x1 = 4
@@ -434,7 +434,7 @@ def test_all_posthoc_multiple_factors(self, suppress_output):
model = MCPower("y = a + b")
model.set_variable_type("a=(factor,3), b=(factor,2)")
model.set_effects("a[2]=0.3, a[3]=0.2, b[2]=0.1")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all-posthoc")
# a: C(3,2)=3, b: C(2,2)=1 → 4 total
@@ -448,7 +448,7 @@ def test_all_posthoc_no_factors_with_all(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all, all-posthoc")
assert "overall" in tests
@@ -461,7 +461,7 @@ def test_all_posthoc_alone_no_factors_raises(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="no factor variables"):
model._parse_target_tests("all-posthoc")
@@ -472,7 +472,7 @@ def test_exclusion_removes_test(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all, -overall")
assert "overall" not in tests
@@ -486,7 +486,7 @@ def test_exclusion_posthoc(self, suppress_output):
model = MCPower("y = group + x1")
model.set_variable_type("group=(factor,3)")
model.set_effects("group[2]=0.4, group[3]=0.3, x1=0.2")
- model.apply()
+ model._apply()
tests = model._parse_target_tests("all-posthoc, -group[1] vs group[2]")
assert "group[1] vs group[2]" not in tests
@@ -500,7 +500,7 @@ def test_exclusion_invalid_raises(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="does not match"):
model._parse_target_tests("all, -nonexistent")
@@ -511,7 +511,7 @@ def test_exclusion_all_raises(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="nothing left"):
model._parse_target_tests("all, -overall, -x1, -x2")
@@ -522,7 +522,7 @@ def test_duplicate_raises(self, suppress_output):
model = MCPower("y = x1 + x2")
model.set_effects("x1=0.5, x2=0.3")
- model.apply()
+ model._apply()
with pytest.raises(ValueError, match="Duplicate"):
model._parse_target_tests("all, x1")
diff --git a/tests/integration/test_scenarios.py b/tests/integration/test_scenarios.py
index 5172fb3..b0339cc 100644
--- a/tests/integration/test_scenarios.py
+++ b/tests/integration/test_scenarios.py
@@ -5,8 +5,10 @@
from unittest.mock import MagicMock
import numpy as np
+import pytest
from mcpower.core.scenarios import (
+ DEFAULT_SCENARIO_CONFIG,
ScenarioRunner,
apply_per_simulation_perturbations,
)
@@ -82,6 +84,152 @@ def test_create_scenario_plots_early_return(self):
runner._create_scenario_plots({"scenarios": {"optimistic": {}}})
+class TestSetScenarioConfigs:
+ """Test set_scenario_configs() merge behavior and KeyError prevention."""
+
+ # All keys that must exist in every scenario config
+ ALL_KEYS = sorted(DEFAULT_SCENARIO_CONFIG["optimistic"].keys())
+
+ def _make_model(self):
+ from mcpower import MCPower
+
+ m = MCPower("y = x1 + x2")
+ m.set_effects("x1=0.3, x2=0.2")
+ return m
+
+ # ── Merge semantics ──────────────────────────────────────────
+
+ def test_custom_scenario_inherits_all_optimistic_keys(self):
+ """New custom scenario with one key still has every required key."""
+ m = self._make_model()
+ m.set_scenario_configs({"extreme": {"heterogeneity": 0.6}})
+ cfg = m._scenario_configs["extreme"]
+ missing = set(self.ALL_KEYS) - set(cfg.keys())
+ assert not missing, f"Missing keys: {missing}"
+
+ def test_custom_scenario_overrides_value(self):
+ """Provided key overrides the optimistic default."""
+ m = self._make_model()
+ m.set_scenario_configs({"extreme": {"heterogeneity": 0.6}})
+ assert m._scenario_configs["extreme"]["heterogeneity"] == 0.6
+
+ def test_custom_scenario_non_overridden_keys_are_optimistic(self):
+ """Non-overridden keys equal the optimistic baseline."""
+ m = self._make_model()
+ m.set_scenario_configs({"extreme": {"heterogeneity": 0.6}})
+ opt = DEFAULT_SCENARIO_CONFIG["optimistic"]
+ cfg = m._scenario_configs["extreme"]
+ for key in self.ALL_KEYS:
+ if key != "heterogeneity":
+ assert cfg[key] == opt[key], f"Key {key}: {cfg[key]} != {opt[key]}"
+
+ def test_existing_scenario_update_preserves_other_keys(self):
+ """Updating one key on 'realistic' keeps the rest intact."""
+ m = self._make_model()
+ m.set_scenario_configs({"realistic": {"heterogeneity": 0.99}})
+ cfg = m._scenario_configs["realistic"]
+ assert cfg["heterogeneity"] == 0.99
+ # Other keys should match original realistic defaults
+ assert cfg["correlation_noise_sd"] == DEFAULT_SCENARIO_CONFIG["realistic"]["correlation_noise_sd"]
+
+ def test_defaults_still_present_after_adding_custom(self):
+ """Adding a custom scenario doesn't remove optimistic/realistic/doomer."""
+ m = self._make_model()
+ m.set_scenario_configs({"custom": {"heterogeneity": 0.1}})
+ for name in ("optimistic", "realistic", "doomer", "custom"):
+ assert name in m._scenario_configs
+
+ def test_multiple_custom_scenarios(self):
+ """Multiple custom scenarios each inherit independently."""
+ m = self._make_model()
+ m.set_scenario_configs({
+ "mild": {"heterogeneity": 0.05},
+ "severe": {"heterogeneity": 0.8, "heteroskedasticity": 0.5},
+ })
+ assert m._scenario_configs["mild"]["heterogeneity"] == 0.05
+ assert m._scenario_configs["mild"]["heteroskedasticity"] == 0.0 # optimistic default
+ assert m._scenario_configs["severe"]["heterogeneity"] == 0.8
+ assert m._scenario_configs["severe"]["heteroskedasticity"] == 0.5
+
+ def test_empty_custom_scenario_equals_optimistic(self):
+ """An empty custom config is identical to the optimistic baseline."""
+ m = self._make_model()
+ m.set_scenario_configs({"empty": {}})
+ opt = DEFAULT_SCENARIO_CONFIG["optimistic"]
+ for key in self.ALL_KEYS:
+ assert m._scenario_configs["empty"][key] == opt[key]
+
+ # ── Type validation ──────────────────────────────────────────
+
+ def test_non_dict_raises_type_error(self):
+ m = self._make_model()
+ with pytest.raises(TypeError):
+ m.set_scenario_configs("not_a_dict")
+
+ def test_returns_self_for_chaining(self):
+ m = self._make_model()
+ result = m.set_scenario_configs({"custom": {"heterogeneity": 0.1}})
+ assert result is m
+
+ # ── End-to-end: no KeyError during simulation ────────────────
+
+ def test_custom_partial_config_runs_without_error(self):
+ """Custom scenario with only one key runs find_power without KeyError."""
+ m = self._make_model()
+ m.set_scenario_configs({"partial": {"heterogeneity": 0.3}})
+ result = m.find_power(
+ 50, scenarios=True, print_results=False, return_results=True
+ )
+ assert "partial" in result["scenarios"]
+ power = result["scenarios"]["partial"]["results"]["individual_powers"]["overall"]
+ assert 0 <= power <= 100
+
+ def test_custom_residual_only_config_runs(self):
+ """Custom scenario with only residual keys runs without error."""
+ m = self._make_model()
+ m.set_scenario_configs({
+ "residual_test": {
+ "residual_change_prob": 1.0,
+ "residual_dists": ["heavy_tailed"],
+ "residual_df": 5,
+ }
+ })
+ result = m.find_power(
+ 50, scenarios=True, print_results=False, return_results=True
+ )
+ assert "residual_test" in result["scenarios"]
+
+ def test_custom_lme_keys_on_ols_model_ignored(self):
+ """LME-specific keys on an OLS model don't cause errors."""
+ m = self._make_model()
+ m.set_scenario_configs({
+ "lme_on_ols": {
+ "icc_noise_sd": 0.3,
+ "random_effect_dist": "heavy_tailed",
+ "random_effect_df": 3,
+ }
+ })
+ result = m.find_power(
+ 50, scenarios=True, print_results=False, return_results=True
+ )
+ assert "lme_on_ols" in result["scenarios"]
+
+ def test_overriding_all_three_defaults(self):
+ """Overriding optimistic, realistic, and doomer all at once."""
+ m = self._make_model()
+ m.set_scenario_configs({
+ "optimistic": {"heterogeneity": 0.01},
+ "realistic": {"heterogeneity": 0.5},
+ "doomer": {"heterogeneity": 0.9},
+ })
+ assert m._scenario_configs["optimistic"]["heterogeneity"] == 0.01
+ assert m._scenario_configs["realistic"]["heterogeneity"] == 0.5
+ assert m._scenario_configs["doomer"]["heterogeneity"] == 0.9
+ # Other keys preserved from defaults
+ assert m._scenario_configs["realistic"]["correlation_noise_sd"] == DEFAULT_SCENARIO_CONFIG["realistic"]["correlation_noise_sd"]
+ assert m._scenario_configs["doomer"]["correlation_noise_sd"] == DEFAULT_SCENARIO_CONFIG["doomer"]["correlation_noise_sd"]
+
+
class TestApplyPerSimulationPerturbations:
"""Test apply_per_simulation_perturbations function."""
@@ -122,3 +270,152 @@ def test_var_type_perturbation(self):
# All normal (type 0) vars should be changed to right_skewed (type 2)
assert np.all(p_types == 2)
+
+
+class TestScenarioConfigKeysE2E:
+ """End-to-end tests for each individual config key and mixed combinations.
+
+ Each test verifies that setting a single config key (or combination)
+ via set_scenario_configs() runs find_power(scenarios=True) without
+ error and produces valid power values.
+ """
+
+ N_SIMS = 50
+ SAMPLE_SIZE = 80
+
+ def _make_model(self):
+ from mcpower import MCPower
+
+ m = MCPower("y = x1 + x2")
+ m.set_effects("x1=0.3, x2=0.2")
+ m.set_simulations(self.N_SIMS)
+ return m
+
+ def _run(self, model, config, scenario_name="test_scenario"):
+ model.set_scenario_configs({scenario_name: config})
+ result = model.find_power(
+ self.SAMPLE_SIZE,
+ scenarios=True,
+ print_results=False,
+ return_results=True,
+ )
+ power = result["scenarios"][scenario_name]["results"]["individual_powers"]["overall"]
+ assert 0 <= power <= 100, f"Power out of range: {power}"
+ return result
+
+ # ── Individual general keys ───────────────────────────────────
+
+ def test_heterogeneity_only(self):
+ self._run(self._make_model(), {"heterogeneity": 0.3})
+
+ def test_heteroskedasticity_only(self):
+ self._run(self._make_model(), {"heteroskedasticity": 0.2})
+
+ def test_correlation_noise_sd_only(self):
+ m = self._make_model()
+ m.set_correlations("(x1,x2)=0.4")
+ self._run(m, {"correlation_noise_sd": 0.3})
+
+ def test_distribution_change_prob_only(self):
+ self._run(self._make_model(), {"distribution_change_prob": 0.5})
+
+ def test_new_distributions_with_change_prob(self):
+ self._run(self._make_model(), {
+ "distribution_change_prob": 1.0,
+ "new_distributions": ["uniform"],
+ })
+
+ # ── Individual residual keys ──────────────────────────────────
+
+ def test_residual_change_prob_only(self):
+ self._run(self._make_model(), {"residual_change_prob": 0.5})
+
+ def test_residual_df_only(self):
+ self._run(self._make_model(), {
+ "residual_change_prob": 1.0,
+ "residual_df": 3,
+ })
+
+ def test_residual_dists_only(self):
+ self._run(self._make_model(), {
+ "residual_change_prob": 1.0,
+ "residual_dists": ["heavy_tailed"],
+ })
+
+ # ── Mixed general combinations ────────────────────────────────
+
+ def test_heterogeneity_and_correlation_noise(self):
+ m = self._make_model()
+ m.set_correlations("(x1,x2)=0.3")
+ self._run(m, {
+ "heterogeneity": 0.25,
+ "correlation_noise_sd": 0.3,
+ })
+
+ def test_distribution_change_and_heteroskedasticity(self):
+ self._run(self._make_model(), {
+ "distribution_change_prob": 0.5,
+ "heteroskedasticity": 0.15,
+ })
+
+ def test_all_general_keys_together(self):
+ m = self._make_model()
+ m.set_correlations("(x1,x2)=0.3")
+ self._run(m, {
+ "heterogeneity": 0.2,
+ "heteroskedasticity": 0.1,
+ "correlation_noise_sd": 0.2,
+ "distribution_change_prob": 0.3,
+ })
+
+ # ── Mixed general + residual ──────────────────────────────────
+
+ def test_general_plus_residual_keys(self):
+ self._run(self._make_model(), {
+ "heterogeneity": 0.2,
+ "residual_change_prob": 0.5,
+ "residual_df": 5,
+ })
+
+ def test_all_ols_keys_together(self):
+ m = self._make_model()
+ m.set_correlations("(x1,x2)=0.3")
+ self._run(m, {
+ "heterogeneity": 0.3,
+ "heteroskedasticity": 0.15,
+ "correlation_noise_sd": 0.25,
+ "distribution_change_prob": 0.4,
+ "new_distributions": ["right_skewed", "uniform"],
+ "residual_change_prob": 0.5,
+ "residual_dists": ["heavy_tailed", "skewed"],
+ "residual_df": 6,
+ })
+
+ # ── Boundary values ───────────────────────────────────────────
+
+ def test_zero_perturbation_matches_optimistic(self):
+ """A custom scenario with all zeros should match optimistic power."""
+ m = self._make_model()
+ m.set_seed(42)
+ result = self._run(m, {
+ "heterogeneity": 0.0,
+ "heteroskedasticity": 0.0,
+ "correlation_noise_sd": 0.0,
+ "distribution_change_prob": 0.0,
+ "residual_change_prob": 0.0,
+ })
+ opt_power = result["scenarios"]["optimistic"]["results"]["individual_powers"]["overall"]
+ custom_power = result["scenarios"]["test_scenario"]["results"]["individual_powers"]["overall"]
+ # Same seed, same zero config → should be close (not exact due to seed offsets)
+ assert abs(opt_power - custom_power) < 15
+
+ def test_max_perturbation_runs(self):
+ """Extreme perturbation values should not crash."""
+ self._run(self._make_model(), {
+ "heterogeneity": 0.9,
+ "heteroskedasticity": 0.5,
+ "correlation_noise_sd": 0.8,
+ "distribution_change_prob": 1.0,
+ "residual_change_prob": 1.0,
+ "residual_df": 2,
+ })
diff --git a/tests/integration/test_test_formula.py b/tests/integration/test_test_formula.py
new file mode 100644
index 0000000..77b2e6e
--- /dev/null
+++ b/tests/integration/test_test_formula.py
@@ -0,0 +1,388 @@
+"""
+End-to-end integration tests for the test_formula feature.
+
+The test_formula feature generates data using one model formula but fits a
+different (reduced) model for statistical testing, enabling model
+misspecification analysis (e.g. omitted variable bias).
+"""
+
+import numpy as np
+import pandas as pd
+import pytest
+
+from mcpower import MCPower
+
+N_SIMS = 200
+SEED = 42
+
+
+# ---------------------------------------------------------------------------
+# Helpers
+# ---------------------------------------------------------------------------
+
+def _power_model(formula, effects, *, n_sims=N_SIMS, seed=SEED, **kwargs):
+ """Create a configured MCPower model ready for find_power."""
+ model = MCPower(formula)
+
+ # Apply optional configuration before effects
+ if "variable_types" in kwargs:
+ model.set_variable_type(kwargs.pop("variable_types"))
+ if "correlations" in kwargs:
+ model.set_correlations(kwargs.pop("correlations"))
+ if "cluster" in kwargs:
+ cluster_cfg = kwargs.pop("cluster")
+ model.set_cluster(**cluster_cfg)
+ if "max_failed" in kwargs:
+ model.set_max_failed_simulations(kwargs.pop("max_failed"))
+ if "upload_data" in kwargs:
+ model.upload_data(kwargs.pop("upload_data"))
+
+ model.set_effects(effects)
+ model.set_simulations(n_sims)
+ model.set_seed(seed)
+ return model
+
+
+def _run_power(model, sample_size, **kwargs):
+ """Run find_power with standard test defaults."""
+ return model.find_power(
+ sample_size,
+ print_results=False,
+ return_results=True,
+ progress_callback=False,
+ **kwargs,
+ )
+
+
+def _individual_powers(result):
+ """Extract individual_powers dict from a result."""
+ return result["results"]["individual_powers"]
+
+
+# ===========================================================================
+# Class 1: TestOLSSubset
+# ===========================================================================
+
+
+class TestOLSSubset:
+ """Test basic OLS test_formula subsetting scenarios."""
+
+ def test_omitted_variable_reduces_power(self):
+ """Omitting x3 from test formula excludes it from results."""
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.5",
+ )
+ result = _run_power(model, 100, test_formula="y = x1 + x2")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" in powers
+ assert "x3" not in powers
+
+ def test_omitted_interaction(self):
+ """Omitting interaction from test formula excludes it from results."""
+ model = _power_model(
+ "y = x1 + x2 + x1:x2",
+ "x1=0.5, x2=0.3, x1:x2=0.2",
+ )
+ result = _run_power(model, 100, test_formula="y = x1 + x2")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" in powers
+ assert "x1:x2" not in powers
+
+ def test_single_variable_test(self):
+ """Testing only x1 from a 3-variable generation model."""
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.2",
+ )
+ result = _run_power(model, 100, test_formula="y = x1")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "overall" in powers
+ assert "x2" not in powers
+ assert "x3" not in powers
+
+ def test_same_formula_matches_no_test_formula(self):
+ """Using test_formula identical to generation gives same powers."""
+ model_a = _power_model("y = x1 + x2", "x1=0.5, x2=0.3")
+ result_a = _run_power(model_a, 100, test_formula="y = x1 + x2")
+
+ model_b = _power_model("y = x1 + x2", "x1=0.5, x2=0.3")
+ result_b = _run_power(model_b, 100)
+
+ powers_a = _individual_powers(result_a)
+ powers_b = _individual_powers(result_b)
+
+ for key in powers_b:
+ assert abs(powers_a[key] - powers_b[key]) < 0.01, (
+ f"Power mismatch for {key}: {powers_a[key]} vs {powers_b[key]}"
+ )
+
+ def test_empty_test_formula_uses_generation(self):
+ """Empty test_formula string uses the generation formula (default)."""
+ model_a = _power_model("y = x1 + x2", "x1=0.5, x2=0.3")
+ result_a = _run_power(model_a, 100, test_formula="")
+
+ model_b = _power_model("y = x1 + x2", "x1=0.5, x2=0.3")
+ result_b = _run_power(model_b, 100)
+
+ powers_a = _individual_powers(result_a)
+ powers_b = _individual_powers(result_b)
+
+ for key in powers_b:
+ assert abs(powers_a[key] - powers_b[key]) < 0.01, (
+ f"Power mismatch for {key}: {powers_a[key]} vs {powers_b[key]}"
+ )
+
+
+# ===========================================================================
+# Class 2: TestFactorVariables
+# ===========================================================================
+
+
+class TestFactorVariables:
+ """Test test_formula with factor (categorical) variables."""
+
+ def test_omitted_factor(self):
+ """Omitting a factor variable from test formula excludes its dummies."""
+ model = _power_model(
+ "y = x1 + x2",
+ "x1=0.5, x2[2]=0.3, x2[3]=0.4",
+ variable_types="x2=(factor,3)",
+ )
+ result = _run_power(model, 150, test_formula="y = x1")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ # Factor dummies should not be in results
+ assert "x2[2]" not in powers
+ assert "x2[3]" not in powers
+
+ def test_factor_kept_continuous_dropped(self):
+ """Keeping factor but dropping continuous variable."""
+ model = _power_model(
+ "y = x1 + x2",
+ "x1=0.5, x2[2]=0.3, x2[3]=0.4",
+ variable_types="x2=(factor,3)",
+ )
+ result = _run_power(model, 150, test_formula="y = x2")
+
+ powers = _individual_powers(result)
+ # x1 excluded
+ assert "x1" not in powers
+ # Factor dummies should be present
+ assert "x2[2]" in powers
+ assert "x2[3]" in powers
+
+
+# ===========================================================================
+# Class 3: TestCorrelationStructures
+# ===========================================================================
+
+
+class TestCorrelationStructures:
+ """Test test_formula with correlated predictors."""
+
+ def test_correlated_variables_subset(self):
+ """Subsetting correlated variables runs without error."""
+ model = _power_model(
+ "y = x1 + x2",
+ "x1=0.5, x2=0.3",
+ correlations="(x1,x2)=0.5",
+ )
+ result = _run_power(model, 100, test_formula="y = x1")
+
+ assert result is not None
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" not in powers
+
+
+# ===========================================================================
+# Class 4: TestResultsStructure
+# ===========================================================================
+
+
+class TestResultsStructure:
+ """Test that result dict contains correct test_formula metadata."""
+
+ def test_results_contain_both_formulas(self):
+ """Result should have data_formula and test_formula fields."""
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.2",
+ )
+ result = _run_power(model, 100, test_formula="y = x1 + x2")
+
+ assert "data_formula" in result["model"]
+ assert "test_formula" in result["model"]
+ # data_formula should be the generation formula
+ assert "x3" in result["model"]["data_formula"]
+ # test_formula should be the reduced formula
+ assert result["model"]["test_formula"] == "y = x1 + x2"
+
+ def test_target_tests_reflect_test_formula(self):
+ """target_tests in results should not contain excluded effects."""
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.2",
+ )
+ result = _run_power(model, 100, test_formula="y = x1 + x2")
+
+ target_tests = result["model"]["target_tests"]
+ assert "x1" in target_tests
+ assert "x2" in target_tests
+ assert "x3" not in target_tests
+
+
+# ===========================================================================
+# Class 5: TestValidation
+# ===========================================================================
+
+
+class TestValidation:
+ """Test validation errors for invalid test_formula usage."""
+
+ def test_nonexistent_variable_raises(self):
+ """test_formula with unknown variable raises ValueError."""
+ model = _power_model(
+ "y = x1 + x2",
+ "x1=0.5, x2=0.3",
+ )
+ with pytest.raises(ValueError, match="not found"):
+ _run_power(model, 100, test_formula="y = x1 + x99")
+
+ def test_ols_to_lme_raises(self):
+ """test_formula with random effects on OLS model raises ValueError.
+
+ When the grouping variable (school) is not in the generation model,
+ validation fails with 'not found'. When it is present but has no
+ cluster config, it fails with 'random effects'.
+ """
+ # Case 1: grouping var not in model at all -> "not found"
+ model = _power_model(
+ "y = x1 + x2",
+ "x1=0.5, x2=0.3",
+ )
+ with pytest.raises(ValueError, match="not found"):
+ _run_power(model, 100, test_formula="y = x1 + (1|school)")
+
+ def test_ols_with_cluster_var_but_no_cluster_config_raises(self):
+ """test_formula with random effects when var exists but no cluster config.
+
+ When the generation model knows about 'school' as a variable but has
+ no cluster specification, the random effects check triggers.
+ """
+ # This would require a model that has 'school' as a predictor but
+ # no set_cluster call. The generation model includes school as a
+ # fixed effect, so it's a known variable.
+ model = _power_model(
+ "y = x1 + school",
+ "x1=0.5, school=0.3",
+ )
+ with pytest.raises(ValueError, match="random effects"):
+ _run_power(model, 100, test_formula="y = x1 + (1|school)")
+
+
+# ===========================================================================
+# Class 6: TestFindSampleSize
+# ===========================================================================
+
+
+class TestFindSampleSize:
+ """Test test_formula with find_sample_size."""
+
+ def test_subset_via_find_sample_size(self):
+ """find_sample_size with test_formula excludes omitted variable."""
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.2",
+ )
+ result = model.find_sample_size(
+ target_test="x1",
+ from_size=30,
+ to_size=100,
+ by=10,
+ test_formula="y = x1 + x2",
+ print_results=False,
+ return_results=True,
+ progress_callback=False,
+ )
+
+ assert result is not None
+ powers_by_test = result["results"]["powers_by_test"]
+ assert "x1" in powers_by_test
+ assert "x3" not in powers_by_test
+
+
+# ===========================================================================
+# Class 7: TestMixedModelCross (LME)
+# ===========================================================================
+
+
+@pytest.mark.lme
+class TestMixedModelCross:
+ """Test test_formula across mixed model boundaries."""
+
+ def test_lme_gen_ols_test(self):
+ """Generate with LME, test with OLS (drop random effects)."""
+ model = _power_model(
+ "y ~ x1 + x2 + (1|school)",
+ "x1=0.5, x2=0.3",
+ cluster={"grouping_var": "school", "ICC": 0.2, "n_clusters": 20},
+ max_failed=0.10,
+ )
+ result = _run_power(model, 1000, test_formula="y ~ x1 + x2")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" in powers
+
+ def test_lme_gen_lme_subset(self):
+ """Generate with LME full model, test with LME subset (drop x2)."""
+ model = _power_model(
+ "y ~ x1 + x2 + (1|school)",
+ "x1=0.5, x2=0.3",
+ cluster={"grouping_var": "school", "ICC": 0.2, "n_clusters": 20},
+ max_failed=0.10,
+ )
+ result = _run_power(model, 1000, test_formula="y ~ x1 + (1|school)")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" not in powers
+
+
+# ===========================================================================
+# Class 8: TestUploadedData
+# ===========================================================================
+
+
+class TestUploadedData:
+ """Test test_formula with uploaded empirical data."""
+
+ def test_upload_with_test_formula(self):
+ """Uploaded data with test_formula excludes omitted variable."""
+ np.random.seed(SEED)
+ data = pd.DataFrame({
+ "x1": np.random.normal(0, 1, 50),
+ "x2": np.random.normal(0, 1, 50),
+ "x3": np.random.normal(0, 1, 50),
+ })
+
+ model = _power_model(
+ "y = x1 + x2 + x3",
+ "x1=0.5, x2=0.3, x3=0.2",
+ upload_data=data,
+ )
+ result = _run_power(model, 100, test_formula="y = x1 + x2")
+
+ powers = _individual_powers(result)
+ assert "x1" in powers
+ assert "x2" in powers
+ assert "x3" not in powers
diff --git a/tests/integration/test_upload_data.py b/tests/integration/test_upload_data.py
index 93a7d8f..05602f0 100644
--- a/tests/integration/test_upload_data.py
+++ b/tests/integration/test_upload_data.py
@@ -71,7 +71,7 @@ def test_binary_auto_detection(self, cars_data):
model = MCPower("mpg = vs + am")
model.upload_data(_select(cars_data, ["vs", "am"]))
model.set_effects("vs=0.3, am=0.4")
- model.apply()
+ model._apply()
# Check that vs and am were detected as uploaded_binary
vs_pred = model._registry.get_predictor("vs")
@@ -85,7 +85,7 @@ def test_factor_auto_detection(self, cars_data):
model = MCPower("mpg = cyl + gear")
model.upload_data(_select(cars_data, ["cyl", "gear"]), preserve_factor_level_names=False)
model.set_effects("cyl[2]=0.3, cyl[3]=0.4, gear[2]=0.2, gear[3]=0.3")
- model.apply()
+ model._apply()
# Check that cyl and gear were detected as factor
# After expansion, check the factor names
@@ -103,7 +103,7 @@ def test_continuous_auto_detection(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]))
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# Check that hp and wt were detected as continuous (uploaded_data)
hp_pred = model._registry.get_predictor("hp")
@@ -122,14 +122,14 @@ def test_constant_column_dropped(self, cars_data):
# Should raise error because 'constant' will be dropped
with pytest.raises(ValueError, match="All uploaded columns were dropped"):
model.upload_data(_select(data, ["constant"]))
- model.apply()
+ model._apply()
def test_mixed_types_auto_detection(self, cars_data):
"""Test auto-detection with mixed variable types."""
model = MCPower("mpg = vs + cyl + hp")
model.upload_data(_select(cars_data, ["vs", "cyl", "hp"]), preserve_factor_level_names=False)
model.set_effects("vs=0.3, cyl[2]=0.2, cyl[3]=0.4, hp=0.5")
- model.apply()
+ model._apply()
vs_pred = model._registry.get_predictor("vs")
hp_pred = model._registry.get_predictor("hp")
@@ -147,7 +147,7 @@ def test_override_to_continuous(self, cars_data):
model = MCPower("mpg = cyl + hp")
model.upload_data(_select(cars_data, ["cyl", "hp"]), data_types={"cyl": "continuous"})
model.set_effects("cyl=0.4, hp=0.5")
- model.apply()
+ model._apply()
cyl_pred = model._registry.get_predictor("cyl")
# Should be uploaded_data (continuous) instead of factor
@@ -178,7 +178,7 @@ def test_override_to_binary(self, cars_data):
model_binary = MCPower("mpg = hp_binary + wt")
model_binary.upload_data(data, data_types={"hp_binary": "binary"})
model_binary.set_effects("hp_binary=0.4, wt=0.3")
- model_binary.apply()
+ model_binary._apply()
hp_pred = model_binary._registry.get_predictor("hp_binary")
assert hp_pred.var_type == "uploaded_binary"
@@ -206,7 +206,7 @@ def test_no_correlation_from_data(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="no")
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# Correlation matrix should be identity (or user-specified)
corr = model.correlation_matrix
@@ -219,7 +219,7 @@ def test_binary_uses_standard_generation(self, cars_data):
model = MCPower("mpg = vs + am")
model.upload_data(_select(cars_data, ["vs", "am"]), preserve_correlation="no")
model.set_effects("vs=0.3, am=0.4")
- model.apply()
+ model._apply()
# Should detect proportions from data
vs_pred = model._registry.get_predictor("vs")
@@ -231,7 +231,7 @@ def test_continuous_uses_lookup_tables(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="no")
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# Should have lookup tables populated
assert model.upload_normal_values.shape[0] > 0
@@ -246,7 +246,7 @@ def test_strict_is_default(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]))
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
assert model._preserve_correlation == "strict"
@@ -255,7 +255,7 @@ def test_correlations_computed_from_data(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="partial")
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# Correlation should match data correlation
hp_arr = np.array(cars_data["hp"])
@@ -278,7 +278,7 @@ def test_user_can_override_correlations(self, cars_data):
# This tests that user correlations can override data correlations
# For now, the implementation always uses data correlations
# TODO: Implement user override priority
- model.apply()
+ model._apply()
# Just verify it doesn't crash
assert model.correlation_matrix is not None
@@ -292,7 +292,7 @@ def test_strict_mode_sets_metadata(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="strict")
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
assert model._preserve_correlation == "strict"
assert model._uploaded_raw_data is not None
@@ -303,7 +303,7 @@ def test_strict_mode_warns_cross_correlations(self, cars_data, capsys):
model = MCPower("mpg = hp + wt + x1") # x1 is created, hp/wt uploaded
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="strict")
model.set_effects("hp=0.5, wt=0.3, x1=0.4")
- model.apply()
+ model._apply()
captured = capsys.readouterr()
# Should warn about cross-correlations
@@ -314,7 +314,7 @@ def test_strict_mode_bootstrap_preserves_relationships(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]), preserve_correlation="strict")
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# Should be able to run simulation without error
result = model.find_power(sample_size=50, print_results=False, return_results=True)
@@ -356,7 +356,7 @@ def test_strict_mode_with_binary(self, cars_data):
model = MCPower("mpg = vs + am")
model.upload_data(_select(cars_data, ["vs", "am"]), preserve_correlation="strict")
model.set_effects("vs=0.3, am=0.4")
- model.apply()
+ model._apply()
# Check metadata
assert "vs" in model._uploaded_var_metadata
@@ -373,7 +373,7 @@ def test_strict_mode_with_factor(self, cars_data):
preserve_factor_level_names=False,
)
model.set_effects("cyl[2]=0.3, cyl[3]=0.4, gear[2]=0.2, gear[3]=0.3")
- model.apply()
+ model._apply()
# Check metadata
assert "cyl" in model._uploaded_var_metadata
@@ -390,7 +390,7 @@ def test_warning_for_unmatched_columns(self, cars_data, capsys):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt", "vs"])) # vs not in model
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
captured = capsys.readouterr()
assert "Ignoring unmatched columns" in captured.out
@@ -401,7 +401,7 @@ def test_warning_for_large_sample_size(self, cars_data, capsys):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]))
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
# 32 samples * 3 = 96, so 100 should trigger warning
model.find_power(sample_size=100, print_results=False)
@@ -424,7 +424,7 @@ def test_warning_for_dropped_constant_columns(self, cars_data, capsys):
# This should raise an error because constant was dropped and no effect was set for it
# But the auto-detection output should show it was dropped
try:
- model.apply()
+ model._apply()
except ValueError:
pass # Expected to fail because constant column missing
@@ -442,7 +442,7 @@ def test_full_dict_with_unmatched_columns(self, cars_data):
model = MCPower("mpg = hp + wt")
model.upload_data(cars_data) # Full dict, not pre-filtered
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
hp_pred = model._registry.get_predictor("hp")
wt_pred = model._registry.get_predictor("wt")
@@ -463,7 +463,7 @@ def test_full_dict_with_mixed_var_types(self, cars_data):
model = MCPower("mpg = vs + cyl + hp")
model.upload_data(cars_data, preserve_factor_level_names=False) # Full dict
model.set_effects("vs=0.3, cyl[2]=0.2, cyl[3]=0.4, hp=0.5")
- model.apply()
+ model._apply()
vs_pred = model._registry.get_predictor("vs")
hp_pred = model._registry.get_predictor("hp")
@@ -493,7 +493,7 @@ def test_string_matched_column_auto_detected_as_factor(self):
model = MCPower("y = x")
model.upload_data(data)
model.set_effects("x[b]=0.3, x[c]=0.4")
- model.apply()
+ model._apply()
assert "x" in model._registry.factor_names
assert "x[b]" in model._registry.dummy_names
assert "x[c]" in model._registry.dummy_names
@@ -507,7 +507,7 @@ def test_no_matching_columns_ignores_data(self, cars_data, capsys):
model = MCPower("mpg = x1 + x2")
model.upload_data(_select(cars_data, ["hp", "wt"]))
model.set_effects("x1=0.3, x2=0.4")
- model.apply()
+ model._apply()
captured = capsys.readouterr()
assert "uploaded data ignored" in captured.out.lower()
@@ -557,7 +557,7 @@ def test_dict_format(self, cars_data):
}
model.upload_data(data_dict)
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
assert model._applied is True
@@ -566,10 +566,10 @@ def test_sample_size_warning_in_find_sample_size(self, cars_data, capsys):
model = MCPower("mpg = hp + wt")
model.upload_data(_select(cars_data, ["hp", "wt"]))
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
- # 32 * 3 = 96, so to_size=150 should trigger warning
- model.find_sample_size(from_size=30, to_size=150, by=20, print_results=False)
+ # 32 * 3 = 96, so size=110 > 96 triggers warning
+ model.find_sample_size(from_size=50, to_size=110, by=30, print_results=False)
captured = capsys.readouterr()
assert "Warning" in captured.out
@@ -583,14 +583,14 @@ def test_string_column_auto_detected_as_factor(self, cars_data):
model = MCPower("mpg = origin + hp")
model.upload_data(_select(cars_data, ["origin", "hp"]))
model.set_effects("origin[Japan]=0.3, origin[USA]=0.4, hp=0.5")
- model.apply()
+ model._apply()
assert "origin" in model._registry.factor_names
def test_string_column_creates_named_dummies(self, cars_data):
model = MCPower("mpg = origin + hp")
model.upload_data(_select(cars_data, ["origin", "hp"]))
model.set_effects("origin[Japan]=0.3, origin[USA]=0.4, hp=0.5")
- model.apply()
+ model._apply()
dummy_names = model._registry.dummy_names
assert "origin[Japan]" in dummy_names
assert "origin[USA]" in dummy_names
@@ -600,7 +600,7 @@ def test_string_column_no_mode(self, cars_data):
model = MCPower("mpg = origin + hp")
model.upload_data(_select(cars_data, ["origin", "hp"]), preserve_correlation="no")
model.set_effects("origin[Japan]=0.3, origin[USA]=0.4, hp=0.5")
- model.apply()
+ model._apply()
assert "origin" in model._registry.factor_names
def test_too_many_string_levels_raises(self):
@@ -611,7 +611,7 @@ def test_too_many_string_levels_raises(self):
model = MCPower("y = name + x1")
with pytest.raises(ValueError, match="too many unique"):
model.upload_data(_select(data, ["name", "x1"]))
- model.apply()
+ model._apply()
class TestPreserveFactorLevelNames:
@@ -621,7 +621,7 @@ def test_numeric_factor_uses_original_values(self, cars_data):
model = MCPower("mpg = cyl + hp")
model.upload_data(_select(cars_data, ["cyl", "hp"]))
model.set_effects("cyl[6]=0.3, cyl[8]=0.4, hp=0.5")
- model.apply()
+ model._apply()
dummy_names = model._registry.dummy_names
assert "cyl[6]" in dummy_names
assert "cyl[8]" in dummy_names
@@ -631,7 +631,7 @@ def test_preserve_false_uses_integer_indices(self, cars_data):
model = MCPower("mpg = cyl + hp")
model.upload_data(_select(cars_data, ["cyl", "hp"]), preserve_factor_level_names=False)
model.set_effects("cyl[2]=0.3, cyl[3]=0.4, hp=0.5")
- model.apply()
+ model._apply()
dummy_names = model._registry.dummy_names
assert "cyl[2]" in dummy_names
assert "cyl[3]" in dummy_names
@@ -640,7 +640,7 @@ def test_custom_reference_via_data_types_tuple(self, cars_data):
model = MCPower("mpg = cyl + hp")
model.upload_data(_select(cars_data, ["cyl", "hp"]), data_types={"cyl": ("factor", 6)})
model.set_effects("cyl[4]=0.3, cyl[8]=0.4, hp=0.5")
- model.apply()
+ model._apply()
dummy_names = model._registry.dummy_names
assert "cyl[4]" in dummy_names
assert "cyl[8]" in dummy_names
@@ -650,7 +650,7 @@ def test_invalid_reference_level_raises(self, cars_data):
model = MCPower("mpg = cyl + hp")
with pytest.raises(ValueError, match="not found in"):
model.upload_data(_select(cars_data, ["cyl", "hp"]), data_types={"cyl": ("factor", 99)})
- model.apply()
+ model._apply()
def test_string_custom_reference(self, cars_data):
model = MCPower("mpg = origin + hp")
@@ -658,7 +658,7 @@ def test_string_custom_reference(self, cars_data):
_select(cars_data, ["origin", "hp"]), data_types={"origin": ("factor", "Japan")}
)
model.set_effects("origin[Europe]=0.3, origin[USA]=0.4, hp=0.5")
- model.apply()
+ model._apply()
dummy_names = model._registry.dummy_names
assert "origin[Europe]" in dummy_names
assert "origin[USA]" in dummy_names
@@ -737,7 +737,7 @@ def test_origin_as_factor(self, cars_data):
model = MCPower("mpg = origin + hp")
model.upload_data(_select(cars_data, ["origin", "hp"]))
model.set_effects("origin[Japan]=0.3, origin[USA]=0.5, hp=0.4")
- model.apply()
+ model._apply()
assert "origin" in model._registry.factor_names
assert "origin[Japan]" in model._registry.dummy_names
@@ -762,7 +762,7 @@ def test_origin_with_cyl_mixed(self, cars_data):
model = MCPower("mpg = origin + cyl")
model.upload_data(_select(cars_data, ["origin", "cyl"]))
model.set_effects("origin[Japan]=0.3, origin[USA]=0.5, cyl[6]=0.2, cyl[8]=0.4")
- model.apply()
+ model._apply()
assert "origin[Japan]" in model._registry.dummy_names
assert "cyl[6]" in model._registry.dummy_names
@@ -822,7 +822,7 @@ def test_dataframe_upload(self):
model = MCPower("mpg = hp + wt")
model.upload_data(df[["hp", "wt"]])
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
hp_pred = model._registry.get_predictor("hp")
assert hp_pred.var_type == "uploaded_data"
@@ -833,7 +833,7 @@ def test_dataframe_with_string_index_column(self):
model = MCPower("mpg = hp + wt")
model.upload_data(df)
model.set_effects("hp=0.5, wt=0.3")
- model.apply()
+ model._apply()
hp_pred = model._registry.get_predictor("hp")
assert hp_pred.var_type == "uploaded_data"
diff --git a/tests/mixed_models/test_cluster_validators.py b/tests/mixed_models/test_cluster_validators.py
index 0cca25e..27540d2 100644
--- a/tests/mixed_models/test_cluster_validators.py
+++ b/tests/mixed_models/test_cluster_validators.py
@@ -102,7 +102,7 @@ def test_sufficient_observations_per_cluster(self):
model.set_cluster("cluster", ICC=0.2, n_clusters=5)
model.set_effects("x=0.5")
model.set_simulations(10)
- model.apply()
+ model._apply()
# 50 / 5 = 10 (above warning band)
result = model.find_power(sample_size=50, return_results=True)
@@ -118,7 +118,7 @@ def test_insufficient_observations_per_cluster_rejected(self):
model.set_cluster("cluster", ICC=0.2, n_clusters=5)
model.set_effects("x=0.5")
model.set_simulations(10)
- model.apply()
+ model._apply()
# 20 / 5 = 4 (below minimum)
with pytest.raises(ValueError, match="Insufficient observations per cluster"):
@@ -134,7 +134,7 @@ def test_validation_message_suggestions(self):
model.set_cluster("cluster", ICC=0.2, n_clusters=10)
model.set_effects("x=0.5")
model.set_simulations(10)
- model.apply()
+ model._apply()
with pytest.raises(ValueError) as exc_info:
model.find_power(sample_size=30) # 30/10 = 3 < 5
@@ -155,7 +155,7 @@ def test_valid_config_runs_successfully(self):
model.set_cluster("cluster", ICC=0.2, n_clusters=5)
model.set_effects("x=0.5")
model.set_simulations(10)
- model.apply()
+ model._apply()
result = model.find_power(sample_size=50, return_results=True) # 10 per cluster
@@ -170,7 +170,7 @@ def test_edge_case_exactly_5_per_cluster(self):
model.set_effects("x=0.5")
model.set_simulations(10)
model.set_max_failed_simulations(0.30) # Allow more failures at edge
- model.apply()
+ model._apply()
result = model.find_power(sample_size=20, return_results=True) # 20/4 = 5
@@ -182,7 +182,7 @@ def test_icc_zero_no_convergence_issues(self):
model.set_cluster("cluster", ICC=0.0, n_clusters=5)
model.set_effects("x=0.5")
model.set_simulations(20)
- model.apply()
+ model._apply()
result = model.find_power(sample_size=250, return_results=True)
diff --git a/tests/mixed_models/test_integration_phase2.py b/tests/mixed_models/test_integration_phase2.py
index 42d8621..0d04919 100644
--- a/tests/mixed_models/test_integration_phase2.py
+++ b/tests/mixed_models/test_integration_phase2.py
@@ -23,7 +23,7 @@ def test_slope_model_setup(self):
slope_intercept_corr=0.3,
)
model.set_effects("x1=0.5")
- model.apply()
+ model._apply()
# Verify cluster spec was configured correctly
spec = model._registry._cluster_specs["school"]
@@ -106,7 +106,7 @@ def test_nested_model_setup(self):
model.set_cluster("school", ICC=0.15, n_clusters=10)
model.set_cluster("classroom", ICC=0.10, n_per_parent=3)
model.set_effects("treatment=0.5")
- model.apply()
+ model._apply()
assert "school" in model._registry._cluster_specs
assert "school:classroom" in model._registry._cluster_specs
diff --git a/tests/mixed_models/test_mixed_models.py b/tests/mixed_models/test_mixed_models.py
index de4b422..e0a867c 100644
--- a/tests/mixed_models/test_mixed_models.py
+++ b/tests/mixed_models/test_mixed_models.py
@@ -405,7 +405,6 @@ def test_unknown_backend_raises(self):
np.zeros(10),
np.array([0]),
np.zeros(10, dtype=int),
- [],
0,
0.05,
backend="nonexistent",
diff --git a/tests/mixed_models/test_mixed_models_validation.py b/tests/mixed_models/test_mixed_models_validation.py
index fbe2ecf..5502bfe 100644
--- a/tests/mixed_models/test_mixed_models_validation.py
+++ b/tests/mixed_models/test_mixed_models_validation.py
@@ -90,7 +90,7 @@ def test_icc_recovery_medium(self):
from mcpower.stats.data_generation import _generate_cluster_effects
- sample_size = 500
+ sample_size = 1000
n_clusters = 20
icc_target = ICC_MODERATE_HIGH
@@ -270,7 +270,7 @@ def test_diagnostics_available(self):
y=y,
target_indices=np.array([0]),
cluster_ids=cluster_ids,
- cluster_column_indices=[],
+
correction_method=0,
alpha=0.05,
backend="statsmodels",
diff --git a/tests/mixed_models/test_scenarios_lme.py b/tests/mixed_models/test_scenarios_lme.py
index f3d5ad6..41597d3 100644
--- a/tests/mixed_models/test_scenarios_lme.py
+++ b/tests/mixed_models/test_scenarios_lme.py
@@ -14,7 +14,6 @@
from mcpower.core.scenarios import (
DEFAULT_SCENARIO_CONFIG,
apply_lme_perturbations,
- apply_lme_residual_perturbations,
)
from mcpower.stats.data_generation import (
_generate_cluster_effects,
@@ -35,7 +34,7 @@ class TestDefaultConfig:
"random_effect_dist",
"random_effect_df",
"icc_noise_sd",
- "residual_dist",
+ "residual_dists",
"residual_change_prob",
"residual_df",
]
@@ -49,22 +48,37 @@ def test_doomer_has_lme_keys(self):
assert key in DEFAULT_SCENARIO_CONFIG["doomer"], f"Missing key: {key}"
def test_realistic_values(self):
+ """Realistic scenario has non-zero LME perturbation values."""
cfg = DEFAULT_SCENARIO_CONFIG["realistic"]
assert cfg["random_effect_dist"] == "heavy_tailed"
- assert cfg["random_effect_df"] == 5
- assert cfg["icc_noise_sd"] == 0.15
- assert cfg["residual_dist"] == "heavy_tailed"
- assert cfg["residual_change_prob"] == 0.3
- assert cfg["residual_df"] == 10
+ assert cfg["random_effect_df"] > 0
+ assert cfg["icc_noise_sd"] > 0
+ assert cfg["residual_dists"] == ["heavy_tailed", "skewed"]
+ assert cfg["residual_change_prob"] > 0
+ assert cfg["residual_df"] > 2
def test_doomer_values(self):
- cfg = DEFAULT_SCENARIO_CONFIG["doomer"]
- assert cfg["random_effect_dist"] == "heavy_tailed"
- assert cfg["random_effect_df"] == 3
- assert cfg["icc_noise_sd"] == 0.30
- assert cfg["residual_dist"] == "heavy_tailed"
- assert cfg["residual_change_prob"] == 0.8
- assert cfg["residual_df"] == 5
+ """Doomer scenario has more severe perturbation than realistic."""
+ real = DEFAULT_SCENARIO_CONFIG["realistic"]
+ doom = DEFAULT_SCENARIO_CONFIG["doomer"]
+ assert doom["random_effect_dist"] == "heavy_tailed"
+ assert doom["random_effect_df"] <= real["random_effect_df"]
+ assert doom["icc_noise_sd"] >= real["icc_noise_sd"]
+ assert doom["residual_dists"] == ["heavy_tailed", "skewed"]
+ assert doom["residual_change_prob"] >= real["residual_change_prob"]
+ assert doom["residual_df"] <= real["residual_df"]
+
+ def test_optimistic_has_lme_keys(self):
+ for key in self.LME_KEYS:
+ assert key in DEFAULT_SCENARIO_CONFIG["optimistic"], f"Missing key: {key}"
+
+ def test_optimistic_values_are_zero(self):
+ cfg = DEFAULT_SCENARIO_CONFIG["optimistic"]
+ assert cfg["heterogeneity"] == 0.0
+ assert cfg["heteroskedasticity"] == 0.0
+ assert cfg["residual_change_prob"] == 0.0
+ assert cfg["icc_noise_sd"] == 0.0
+ assert cfg["random_effect_dist"] == "normal"
# ---------------------------------------------------------------------------
@@ -309,73 +323,3 @@ def test_slopes_without_perturbations(self):
assert result.intercept_columns.shape == (1000, 1)
-# ---------------------------------------------------------------------------
-# apply_lme_residual_perturbations
-# ---------------------------------------------------------------------------
-class TestApplyLmeResidualPerturbations:
- """Test apply_lme_residual_perturbations() function."""
-
- def _make_y(self, seed=42):
- """Generate a deterministic y vector with known errors."""
- rng = np.random.RandomState(seed + 2)
- return rng.standard_normal(500)
-
- def test_normal_dist_returns_unchanged(self):
- y = self._make_y()
- config = {"residual_dist": "normal", "residual_change_prob": 1.0, "residual_df": 5}
- result = apply_lme_residual_perturbations(y.copy(), config, 42)
- np.testing.assert_array_equal(result, y)
-
- def test_zero_prob_returns_unchanged(self):
- y = self._make_y()
- config = {"residual_dist": "heavy_tailed", "residual_change_prob": 0.0, "residual_df": 5}
- result = apply_lme_residual_perturbations(y.copy(), config, 42)
- np.testing.assert_array_equal(result, y)
-
- def test_prob_1_always_applies(self):
- y = self._make_y()
- config = {"residual_dist": "heavy_tailed", "residual_change_prob": 1.0, "residual_df": 5}
- result = apply_lme_residual_perturbations(y.copy(), config, 42)
- # Should be different from original
- assert not np.array_equal(result, y)
-
- def test_heavy_tailed_residuals_have_excess_kurtosis(self):
- """When residuals are replaced with t(5), the diff should have heavy tails."""
- y_orig = self._make_y()
- config = {"residual_dist": "heavy_tailed", "residual_change_prob": 1.0, "residual_df": 5}
- y_perturbed = apply_lme_residual_perturbations(y_orig.copy(), config, 42)
- diff = y_perturbed - y_orig
- # The diff = new_errors - original_errors. Both have finite variance,
- # but the new_errors are t(5) which has excess kurtosis.
- # For large enough N, the kurtosis of the difference should be positive.
- sp_stats.kurtosis(diff + y_orig, fisher=True)
- # Just check it ran without error and output differs
- assert not np.array_equal(y_perturbed, y_orig)
-
- def test_skewed_residuals_applied(self):
- y_orig = self._make_y()
- config = {"residual_dist": "skewed", "residual_change_prob": 1.0, "residual_df": 5}
- y_perturbed = apply_lme_residual_perturbations(y_orig.copy(), config, 42)
- assert not np.array_equal(y_perturbed, y_orig)
-
- def test_coin_flip_seed_reproducible(self):
- y = self._make_y()
- config = {"residual_dist": "heavy_tailed", "residual_change_prob": 0.5, "residual_df": 5}
- r1 = apply_lme_residual_perturbations(y.copy(), config, 42)
- r2 = apply_lme_residual_perturbations(y.copy(), config, 42)
- np.testing.assert_array_equal(r1, r2)
-
- def test_coin_flip_prob_respected(self):
- """With prob=0.3, roughly 30% of simulations should be perturbed."""
- config = {"residual_dist": "heavy_tailed", "residual_change_prob": 0.3, "residual_df": 5}
- n_perturbed = 0
- n_trials = 200
- y_template = np.ones(100)
- for i in range(n_trials):
- y = y_template.copy()
- result = apply_lme_residual_perturbations(y, config, i * 100)
- if not np.array_equal(result, y_template):
- n_perturbed += 1
- # Should be roughly 30% ± some tolerance
- pct = n_perturbed / n_trials
- assert 0.10 < pct < 0.55, f"Expected ~30% perturbed, got {pct:.1%}"
diff --git a/tests/specs/test_alpha_levels.py b/tests/specs/test_alpha_levels.py
index 529e9db..c1f0908 100644
--- a/tests/specs/test_alpha_levels.py
+++ b/tests/specs/test_alpha_levels.py
@@ -1,9 +1,8 @@
"""
-Non-default alpha level tests — backend-agnostic.
+Non-default alpha level tests.
Validates that the full alpha pipeline (power accuracy, corrections,
null calibration) works correctly at alpha != 0.05.
-Tests run on ALL available backends via the backend fixture.
"""
import contextlib
@@ -12,7 +11,7 @@
import numpy as np
import pytest
-from tests.config import N_SIMS, SEED
+from tests.config import N_SIMS, N_SIMS_ORDERING, N_SIMS_STANDARD, SEED
from tests.helpers.analytical import analytical_f_power, analytical_t_power
from tests.helpers.mc_margins import mc_accuracy_margin, mc_margin
from tests.helpers.power_helpers import get_power, get_power_corrected, make_null_model
@@ -44,7 +43,7 @@ class TestAlphaAccuracyVsAnalytical:
(0.5, 100),
],
)
- def test_single_predictor_t_test_alpha(self, backend, alpha, beta, n):
+ def test_single_predictor_t_test_alpha(self, alpha, beta, n):
"""t-test power matches analytical non-central t at non-default alpha."""
from mcpower import MCPower
@@ -63,7 +62,7 @@ def test_single_predictor_t_test_alpha(self, backend, alpha, beta, n):
exact_power = analytical_t_power(beta, n, p=1, sigma_eps=1.0, vif_j=1.0, alpha=alpha)
margin = mc_accuracy_margin(exact_power, N_SIMS)
assert abs(mc_power - exact_power) < margin, (
- f"[{backend}] alpha={alpha}, β={beta}, n={n}: MC={mc_power:.2f}%, analytical={exact_power:.2f}% ± {margin:.2f}%"
+ f"alpha={alpha}, β={beta}, n={n}: MC={mc_power:.2f}%, analytical={exact_power:.2f}% ± {margin:.2f}%"
)
@pytest.mark.parametrize("alpha", [0.01, 0.10])
@@ -74,7 +73,7 @@ def test_single_predictor_t_test_alpha(self, backend, alpha, beta, n):
(0.5, 0.3, 80),
],
)
- def test_two_predictors_uncorrelated_alpha(self, backend, alpha, b1, b2, n):
+ def test_two_predictors_uncorrelated_alpha(self, alpha, b1, b2, n):
"""Each t-test and F-test with Σ = I at non-default alpha."""
from mcpower import MCPower
@@ -103,14 +102,14 @@ def test_two_predictors_uncorrelated_alpha(self, backend, alpha, b1, b2, n):
)
margin = mc_accuracy_margin(exact, N_SIMS)
assert abs(mc_power - exact) < margin, (
- f"[{backend}] alpha={alpha}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
+ f"alpha={alpha}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
)
mc_f = get_power(result, "overall")
exact_f = analytical_f_power([b1, b2], n, Sigma, sigma_eps=1.0, alpha=alpha)
margin_f = mc_accuracy_margin(exact_f, N_SIMS)
assert abs(mc_f - exact_f) < margin_f, (
- f"[{backend}] alpha={alpha}, F-test: MC={mc_f:.2f}%, analytical={exact_f:.2f}% ± {margin_f:.2f}%"
+ f"alpha={alpha}, F-test: MC={mc_f:.2f}%, analytical={exact_f:.2f}% ± {margin_f:.2f}%"
)
@pytest.mark.parametrize("alpha", [0.01, 0.10])
@@ -121,7 +120,7 @@ def test_two_predictors_uncorrelated_alpha(self, backend, alpha, b1, b2, n):
(0.5, 0.3, 0.5, 80),
],
)
- def test_two_predictors_correlated_alpha(self, backend, alpha, b1, b2, rho, n):
+ def test_two_predictors_correlated_alpha(self, alpha, b1, b2, rho, n):
"""VIF-corrected t-tests with correlated predictors at non-default alpha."""
from mcpower import MCPower
@@ -154,7 +153,7 @@ def test_two_predictors_correlated_alpha(self, backend, alpha, b1, b2, rho, n):
)
margin = mc_accuracy_margin(exact, N_SIMS)
assert abs(mc_power - exact) < margin, (
- f"[{backend}] alpha={alpha}, rho={rho}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
+ f"alpha={alpha}, rho={rho}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
)
@@ -170,9 +169,9 @@ class TestAlphaCorrectionAccuracy:
@pytest.mark.parametrize("alpha", [0.01, 0.10])
@pytest.mark.parametrize("correction", ["bonferroni", "holm", "fdr"])
- def test_corrected_leq_uncorrected_at_alpha(self, backend, alpha, correction):
+ def test_corrected_leq_uncorrected_at_alpha(self, alpha, correction):
"""Corrected power <= uncorrected power when all effects = 0."""
- m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS, alpha=alpha, seed=SEED)
+ m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS_ORDERING, alpha=alpha, seed=SEED)
result = m.find_power(
sample_size=100,
target_test="x1, x2, x3",
@@ -184,14 +183,14 @@ def test_corrected_leq_uncorrected_at_alpha(self, backend, alpha, correction):
uncorr = get_power(result, var)
corr = get_power_corrected(result, var)
assert corr <= uncorr + 0.5, (
- f"[{backend}] alpha={alpha}, {correction}: corrected {corr:.2f}% > uncorrected {uncorr:.2f}% for {var}"
+ f"alpha={alpha}, {correction}: corrected {corr:.2f}% > uncorrected {uncorr:.2f}% for {var}"
)
@pytest.mark.parametrize("alpha", [0.01, 0.10])
@pytest.mark.parametrize("correction", ["bonferroni", "holm"])
- def test_fwer_controlled_at_alpha(self, backend, alpha, correction):
+ def test_fwer_controlled_at_alpha(self, alpha, correction):
"""FWER-controlling methods keep per-test rejection below nominal alpha."""
- m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS, alpha=alpha, seed=SEED)
+ m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS_ORDERING, alpha=alpha, seed=SEED)
result = m.find_power(
sample_size=100,
target_test="x1, x2, x3",
@@ -201,17 +200,17 @@ def test_fwer_controlled_at_alpha(self, backend, alpha, correction):
)
for var in ["x1", "x2", "x3"]:
corr = get_power_corrected(result, var)
- assert corr < alpha * 100 + mc_margin(alpha, N_SIMS), (
- f"[{backend}] alpha={alpha}, {correction} FWER violation for {var}: corrected power = {corr:.2f}%"
+ assert corr < alpha * 100 + mc_margin(alpha, N_SIMS_ORDERING), (
+ f"alpha={alpha}, {correction} FWER violation for {var}: corrected power = {corr:.2f}%"
)
@pytest.mark.parametrize("alpha", [0.01, 0.10])
- def test_bonferroni_more_conservative_than_fdr_at_alpha(self, backend, alpha):
+ def test_bonferroni_more_conservative_than_fdr_at_alpha(self, alpha):
"""Bonferroni should reject <= FDR (BH) under non-null at non-default alpha."""
from mcpower import MCPower
m = MCPower("y = x1 + x2 + x3")
- m.set_simulations(N_SIMS)
+ m.set_simulations(N_SIMS_ORDERING)
m.set_seed(SEED)
m.set_alpha(alpha)
m.set_effects("x1=0.3, x2=0.2, x3=0.1")
@@ -233,7 +232,7 @@ def test_bonferroni_more_conservative_than_fdr_at_alpha(self, backend, alpha):
for var in ["x1", "x2", "x3"]:
bonf = get_power_corrected(result_bonf, var)
fdr = get_power_corrected(result_fdr, var)
- assert bonf <= fdr + 2.0, f"[{backend}] alpha={alpha}: Bonferroni ({bonf:.2f}%) > FDR ({fdr:.2f}%) for {var}"
+ assert bonf <= fdr + 2.0, f"alpha={alpha}: Bonferroni ({bonf:.2f}%) > FDR ({fdr:.2f}%) for {var}"
# ── Class 3: Null calibration at alpha != 0.05 (multi-predictor) ────
@@ -245,29 +244,29 @@ class TestAlphaCalibrationExtended:
to multi-predictor models and corrected rejection under the null.
"""
- @pytest.mark.parametrize("alpha", [0.01, 0.05, 0.10])
- def test_null_rejection_multi_predictor(self, backend, alpha):
+ @pytest.mark.parametrize("alpha", [0.01, 0.10])
+ def test_null_rejection_multi_predictor(self, alpha):
"""Two-predictor null: each t-test and overall F-test reject at ~alpha."""
- m = make_null_model("y = x1 + x2", n_sims=N_SIMS, alpha=alpha, seed=SEED)
+ m = make_null_model("y = x1 + x2", n_sims=N_SIMS_STANDARD, alpha=alpha, seed=SEED)
result = m.find_power(
sample_size=100,
target_test="all",
print_results=False,
return_results=True,
)
- margin = mc_margin(alpha, N_SIMS)
+ margin = mc_margin(alpha, N_SIMS_STANDARD)
expected = alpha * 100
for test_name in ["x1", "x2", "overall"]:
power = get_power(result, test_name)
assert abs(power - expected) < margin, (
- f"[{backend}] alpha={alpha}, {test_name}: observed {power:.2f}%, expected {expected}% ± {margin:.2f}%"
+ f"alpha={alpha}, {test_name}: observed {power:.2f}%, expected {expected}% ± {margin:.2f}%"
)
- @pytest.mark.parametrize("alpha", [0.01, 0.05, 0.10])
+ @pytest.mark.parametrize("alpha", [0.01, 0.10])
@pytest.mark.parametrize("correction", ["bonferroni", "holm"])
- def test_null_rejection_corrected_at_alpha(self, backend, alpha, correction):
+ def test_null_rejection_corrected_at_alpha(self, alpha, correction):
"""Corrected null rejection stays below alpha + MC margin for 3 predictors."""
- m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS, alpha=alpha, seed=SEED)
+ m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS_STANDARD, alpha=alpha, seed=SEED)
result = m.find_power(
sample_size=100,
target_test="x1, x2, x3",
@@ -275,9 +274,9 @@ def test_null_rejection_corrected_at_alpha(self, backend, alpha, correction):
print_results=False,
return_results=True,
)
- margin = mc_margin(alpha, N_SIMS)
+ margin = mc_margin(alpha, N_SIMS_STANDARD)
for var in ["x1", "x2", "x3"]:
corr = get_power_corrected(result, var)
assert corr < alpha * 100 + margin, (
- f"[{backend}] alpha={alpha}, {correction}, {var}: corrected rejection {corr:.2f}% exceeds {alpha * 100}% + {margin:.2f}%"
+ f"alpha={alpha}, {correction}, {var}: corrected rejection {corr:.2f}% exceeds {alpha * 100}% + {margin:.2f}%"
)
diff --git a/tests/specs/test_corrections.py b/tests/specs/test_corrections.py
index b26b8e1..025aee7 100644
--- a/tests/specs/test_corrections.py
+++ b/tests/specs/test_corrections.py
@@ -1,7 +1,5 @@
"""
-Multiple comparison correction tests — backend-agnostic.
-
-Tests run on ALL available backends via the backend fixture.
+Multiple comparison correction tests.
"""
import contextlib
@@ -9,7 +7,7 @@
import pytest
-from tests.config import N_SIMS, SEED
+from tests.config import N_SIMS_ORDERING as N_SIMS, SEED
from tests.helpers.mc_margins import mc_margin
from tests.helpers.power_helpers import get_power, get_power_corrected, make_null_model
@@ -28,7 +26,7 @@ class TestCorrectionConservativeness:
"""
@pytest.mark.parametrize("correction", ["bonferroni", "holm", "fdr"])
- def test_corrected_leq_uncorrected_under_null(self, backend, correction):
+ def test_corrected_leq_uncorrected_under_null(self, correction):
"""Corrected power ≤ uncorrected power when all effects = 0."""
m = make_null_model("y = x1 + x2 + x3", n_sims=N_SIMS, seed=SEED)
result = m.find_power(
@@ -42,11 +40,11 @@ def test_corrected_leq_uncorrected_under_null(self, backend, correction):
uncorr = get_power(result, var)
corr = get_power_corrected(result, var)
assert corr <= uncorr + 0.5, ( # tiny tolerance for MC noise
- f"[{backend}] {correction}: corrected {corr:.2f}% > uncorrected {uncorr:.2f}% for {var}"
+ f"{correction}: corrected {corr:.2f}% > uncorrected {uncorr:.2f}% for {var}"
)
@pytest.mark.parametrize("correction", ["bonferroni", "holm"])
- def test_fwer_controlled_under_null(self, backend, correction):
+ def test_fwer_controlled_under_null(self, correction):
"""
Family-wise error rate under H0 should be ≤ alpha.
@@ -65,10 +63,10 @@ def test_fwer_controlled_under_null(self, backend, correction):
# Under complete null, FWER-controlling methods should have
# per-test rejection well below the nominal alpha
assert corr < m.alpha * 100 + mc_margin(m.alpha, m.n_simulations), (
- f"[{backend}] {correction} FWER violation for {var}: corrected power = {corr:.2f}%"
+ f"{correction} FWER violation for {var}: corrected power = {corr:.2f}%"
)
- def test_bonferroni_more_conservative_than_fdr(self, backend):
+ def test_bonferroni_more_conservative_than_fdr(self):
"""Bonferroni should reject ≤ FDR (BH) under non-null."""
from mcpower import MCPower
@@ -95,4 +93,4 @@ def test_bonferroni_more_conservative_than_fdr(self, backend):
bonf = get_power_corrected(result_bonf, var)
fdr = get_power_corrected(result_fdr, var)
# Bonferroni ≤ BH-FDR (with MC tolerance)
- assert bonf <= fdr + 2.0, f"[{backend}] Bonferroni ({bonf:.2f}%) > FDR ({fdr:.2f}%) for {var}"
+ assert bonf <= fdr + 2.0, f"Bonferroni ({bonf:.2f}%) > FDR ({fdr:.2f}%) for {var}"
diff --git a/tests/specs/test_monotonicity.py b/tests/specs/test_monotonicity.py
index 4261b99..bac2fe9 100644
--- a/tests/specs/test_monotonicity.py
+++ b/tests/specs/test_monotonicity.py
@@ -1,8 +1,7 @@
"""
-Power monotonicity tests — backend-agnostic.
+Power monotonicity tests.
Power must increase with effect size, sample size, and alpha.
-Tests run on ALL available backends via the backend fixture.
"""
import contextlib
@@ -10,7 +9,7 @@
import pytest
-from tests.config import N_SIMS, SEED
+from tests.config import N_SIMS_ORDERING as N_SIMS, SEED
from tests.helpers.power_helpers import get_power
@@ -24,7 +23,7 @@ def _quiet():
class TestPowerMonotonicity:
"""Power must increase with effect size, sample size, and alpha."""
- def test_power_increases_with_effect_size(self, backend):
+ def test_power_increases_with_effect_size(self):
"""Larger standardised beta → higher power."""
from mcpower import MCPower
@@ -43,9 +42,9 @@ def test_power_increases_with_effect_size(self, backend):
powers.append(get_power(result, "x1"))
for i in range(len(powers) - 1):
- assert powers[i] < powers[i + 1], f"[{backend}] Power not monotonic in effect size: {powers}"
+ assert powers[i] < powers[i + 1], f"Power not monotonic in effect size: {powers}"
- def test_power_increases_with_sample_size(self, backend):
+ def test_power_increases_with_sample_size(self):
"""Larger N → higher power (for non-zero effect)."""
from mcpower import MCPower
@@ -64,9 +63,9 @@ def test_power_increases_with_sample_size(self, backend):
powers.append(get_power(result, "x1"))
for i in range(len(powers) - 1):
- assert powers[i] < powers[i + 1], f"[{backend}] Power not monotonic in N: {powers}"
+ assert powers[i] < powers[i + 1], f"Power not monotonic in N: {powers}"
- def test_power_increases_with_alpha(self, backend):
+ def test_power_increases_with_alpha(self):
"""Less stringent alpha → higher power."""
from mcpower import MCPower
@@ -86,13 +85,13 @@ def test_power_increases_with_alpha(self, backend):
powers.append(get_power(result, "x1"))
for i in range(len(powers) - 1):
- assert powers[i] < powers[i + 1], f"[{backend}] Power not monotonic in alpha: {powers}"
+ assert powers[i] < powers[i + 1], f"Power not monotonic in alpha: {powers}"
class TestPowerConvergence:
"""Power must approach 100% when signal is overwhelming."""
- def test_large_effect_high_power(self, backend):
+ def test_large_effect_high_power(self):
"""Very large effect → power near 100%."""
from mcpower import MCPower
@@ -107,9 +106,9 @@ def test_large_effect_high_power(self, backend):
return_results=True,
)
power = get_power(result, "x1")
- assert power > 99.0, f"[{backend}] Large-effect power should be ~100%, got {power:.2f}%"
+ assert power > 99.0, f"Large-effect power should be ~100%, got {power:.2f}%"
- def test_large_n_moderate_effect(self, backend):
+ def test_large_n_moderate_effect(self):
"""Large N with moderate effect → power near 100%."""
from mcpower import MCPower
@@ -124,4 +123,4 @@ def test_large_n_moderate_effect(self, backend):
return_results=True,
)
power = get_power(result, "x1")
- assert power > 99.0, f"[{backend}] Large-N power should be ~100%, got {power:.2f}%"
+ assert power > 99.0, f"Large-N power should be ~100%, got {power:.2f}%"
diff --git a/tests/specs/test_power_accuracy.py b/tests/specs/test_power_accuracy.py
index 755d0bb..fff7235 100644
--- a/tests/specs/test_power_accuracy.py
+++ b/tests/specs/test_power_accuracy.py
@@ -1,9 +1,8 @@
"""
-Power accuracy tests — backend-agnostic.
+Power accuracy tests.
Compare MC power estimates against exact analytical power from
non-central t / F distributions.
-Tests run on ALL available backends via the backend fixture.
"""
import contextlib
@@ -42,7 +41,7 @@ class TestAccuracyVsAnalytical:
(0.5, 150),
],
)
- def test_single_predictor_t_test(self, backend, beta, n):
+ def test_single_predictor_t_test(self, beta, n):
"""t-test power matches analytical non-central t."""
from mcpower import MCPower
@@ -60,7 +59,7 @@ def test_single_predictor_t_test(self, backend, beta, n):
exact_power = analytical_t_power(beta, n, p=1, sigma_eps=1.0, vif_j=1.0)
margin = mc_accuracy_margin(exact_power, N_SIMS)
assert abs(mc_power - exact_power) < margin, (
- f"[{backend}] β={beta}, n={n}: MC={mc_power:.2f}%, analytical={exact_power:.2f}% ± {margin:.2f}%"
+ f"β={beta}, n={n}: MC={mc_power:.2f}%, analytical={exact_power:.2f}% ± {margin:.2f}%"
)
@pytest.mark.parametrize(
@@ -71,7 +70,7 @@ def test_single_predictor_t_test(self, backend, beta, n):
(0.2, 0.2, 200),
],
)
- def test_two_predictors_uncorrelated(self, backend, b1, b2, n):
+ def test_two_predictors_uncorrelated(self, b1, b2, n):
"""Each t-test and F-test with Σ = I."""
from mcpower import MCPower
@@ -91,12 +90,12 @@ def test_two_predictors_uncorrelated(self, backend, b1, b2, n):
mc_power = get_power(result, var)
exact = analytical_t_power(beta, n, p=2, sigma_eps=1.0, vif_j=1.0)
margin = mc_accuracy_margin(exact, N_SIMS)
- assert abs(mc_power - exact) < margin, f"[{backend}] {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
+ assert abs(mc_power - exact) < margin, f"{var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
mc_f = get_power(result, "overall")
exact_f = analytical_f_power([b1, b2], n, Sigma, sigma_eps=1.0)
margin_f = mc_accuracy_margin(exact_f, N_SIMS)
- assert abs(mc_f - exact_f) < margin_f, f"[{backend}] F-test: MC={mc_f:.2f}%, analytical={exact_f:.2f}% ± {margin_f:.2f}%"
+ assert abs(mc_f - exact_f) < margin_f, f"F-test: MC={mc_f:.2f}%, analytical={exact_f:.2f}% ± {margin_f:.2f}%"
@pytest.mark.parametrize(
"b1,b2,rho,n",
@@ -107,7 +106,7 @@ def test_two_predictors_uncorrelated(self, backend, b1, b2, n):
(0.5, 0.3, 0.5, 80),
],
)
- def test_two_predictors_correlated_t_tests(self, backend, b1, b2, rho, n):
+ def test_two_predictors_correlated_t_tests(self, b1, b2, rho, n):
"""Individual t-tests with correlated predictors: VIF matters."""
from mcpower import MCPower
@@ -132,5 +131,5 @@ def test_two_predictors_correlated_t_tests(self, backend, b1, b2, rho, n):
exact = analytical_t_power(beta, n, p=2, sigma_eps=1.0, vif_j=vif)
margin = mc_accuracy_margin(exact, N_SIMS)
assert abs(mc_power - exact) < margin, (
- f"[{backend}] rho={rho}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
+ f"rho={rho}, {var}: MC={mc_power:.2f}%, analytical={exact:.2f}% ± {margin:.2f}%"
)
diff --git a/tests/specs/test_type1_error.py b/tests/specs/test_type1_error.py
index c56b4fe..565d80e 100644
--- a/tests/specs/test_type1_error.py
+++ b/tests/specs/test_type1_error.py
@@ -1,8 +1,7 @@
"""
-Type I error control tests — backend-agnostic.
+Type I error control tests.
Under H0 (effect = 0), rejection rate must equal alpha.
-Tests run on ALL available backends via the backend fixture.
"""
import contextlib
@@ -10,7 +9,7 @@
import pytest
-from tests.config import N_SIMS, SEED
+from tests.config import N_SIMS_STANDARD as N_SIMS, SEED
from tests.helpers.mc_margins import mc_margin
from tests.helpers.power_helpers import get_power, make_null_model
@@ -25,7 +24,7 @@ def _quiet():
class TestTypeIErrorControl:
"""Under H0 (effect = 0), rejection rate must equal alpha."""
- def test_single_predictor_null_overall(self, backend):
+ def test_single_predictor_null_overall(self):
"""F-test rejection rate ≈ alpha with one predictor at zero effect."""
m = make_null_model("y = x1", n_sims=N_SIMS, seed=SEED)
result = m.find_power(
@@ -37,9 +36,9 @@ def test_single_predictor_null_overall(self, backend):
power = get_power(result, "overall")
margin = mc_margin(m.alpha, m.n_simulations)
expected = m.alpha * 100
- assert abs(power - expected) < margin, f"[{backend}] F-test power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
+ assert abs(power - expected) < margin, f"F-test power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
- def test_single_predictor_null_individual(self, backend):
+ def test_single_predictor_null_individual(self):
"""t-test rejection rate ≈ alpha for a single zero-effect predictor."""
m = make_null_model("y = x1", n_sims=N_SIMS, seed=SEED)
result = m.find_power(
@@ -51,9 +50,9 @@ def test_single_predictor_null_individual(self, backend):
power = get_power(result, "x1")
margin = mc_margin(m.alpha, m.n_simulations)
expected = m.alpha * 100
- assert abs(power - expected) < margin, f"[{backend}] t-test power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
+ assert abs(power - expected) < margin, f"t-test power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
- def test_two_predictors_null_each(self, backend):
+ def test_two_predictors_null_each(self):
"""Both predictors at zero → each t-test rejects at ~alpha."""
m = make_null_model("y = x1 + x2", n_sims=N_SIMS, seed=SEED)
result = m.find_power(
@@ -66,9 +65,9 @@ def test_two_predictors_null_each(self, backend):
expected = m.alpha * 100
for var in ["x1", "x2"]:
power = get_power(result, var)
- assert abs(power - expected) < margin, f"[{backend}] {var} power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
+ assert abs(power - expected) < margin, f"{var} power under H0: {power:.2f}%, expected {expected}% ± {margin:.2f}%"
- def test_large_sample_null(self, backend):
+ def test_large_sample_null(self):
"""
Large N with zero effect must NOT inflate Type I error.
@@ -76,7 +75,7 @@ def test_large_sample_null(self, backend):
"""
m = make_null_model("y = x1", n_sims=N_SIMS, seed=SEED)
result = m.find_power(
- sample_size=1000,
+ sample_size=500,
target_test="x1",
print_results=False,
return_results=True,
@@ -85,7 +84,7 @@ def test_large_sample_null(self, backend):
margin = mc_margin(m.alpha, m.n_simulations)
expected = m.alpha * 100
assert abs(power - expected) < margin, (
- f"[{backend}] Large-N null power: {power:.2f}%, expected {expected}% ± {margin:.2f}% (Type I error inflated with N?)"
+ f"Large-N null power: {power:.2f}%, expected {expected}% ± {margin:.2f}% (Type I error inflated with N?)"
)
@@ -93,7 +92,7 @@ class TestAlphaCalibration:
"""Rejection rate tracks the nominal alpha across levels."""
@pytest.mark.parametrize("alpha", [0.01, 0.05, 0.10])
- def test_null_rejection_matches_alpha(self, backend, alpha):
+ def test_null_rejection_matches_alpha(self, alpha):
m = make_null_model("y = x1", n_sims=N_SIMS, alpha=alpha, seed=SEED)
result = m.find_power(
sample_size=100,
@@ -104,4 +103,4 @@ def test_null_rejection_matches_alpha(self, backend, alpha):
power = get_power(result, "x1")
margin = mc_margin(alpha, m.n_simulations)
expected = alpha * 100
- assert abs(power - expected) < margin, f"[{backend}] alpha={alpha}: observed {power:.2f}%, expected {expected}% ± {margin:.2f}%"
+ assert abs(power - expected) < margin, f"alpha={alpha}: observed {power:.2f}%, expected {expected}% ± {margin:.2f}%"
diff --git a/tests/unit/test_distributions.py b/tests/unit/test_distributions.py
index d3e77f9..693d1ff 100644
--- a/tests/unit/test_distributions.py
+++ b/tests/unit/test_distributions.py
@@ -24,7 +24,6 @@
import pytest
from mcpower.stats.distributions import (
- _BACKEND,
chi2_cdf,
chi2_ppf,
compute_critical_values_lme,
@@ -587,29 +586,6 @@ def test_studentized_range_k_too_large_returns_inf(self):
# ===========================================================================
# 15. Backend detection
-# ===========================================================================
-class TestBackendDetection:
- """Verify the distribution backend is correctly detected."""
-
- def test_backend_is_set(self):
- assert _BACKEND is not None
-
- def test_backend_is_string(self):
- assert isinstance(_BACKEND, str)
-
- def test_backend_is_known_value(self):
- assert _BACKEND in ("native", "scipy")
-
- def test_native_backend_when_compiled(self):
- """When the C++ extension is compiled, backend should be 'native'."""
- try:
- import mcpower.backends.mcpower_native # noqa: F401
-
- assert _BACKEND == "native"
- except ImportError:
- pytest.skip("C++ native backend not compiled")
-
-
# ===========================================================================
# Cross-consistency checks
# ===========================================================================
diff --git a/tests/unit/test_distributions_coverage.py b/tests/unit/test_distributions_coverage.py
new file mode 100644
index 0000000..224c8e6
--- /dev/null
+++ b/tests/unit/test_distributions_coverage.py
@@ -0,0 +1,42 @@
+"""Tests for distributions.py — optimizer functions and edge cases."""
+
+import numpy as np
+import pytest
+
+from mcpower.stats.distributions import minimize_lbfgsb, minimize_scalar_brent
+
+
+class TestOptimizerLBFGSB:
+ """L-BFGS-B optimizer via native backend."""
+
+ def test_finds_correct_minimum(self):
+ # Simple quadratic: f(x) = (x-2)^2
+ result = minimize_lbfgsb(
+ lambda x: float((x[0] - 2) ** 2),
+ x0=np.array([0.0]),
+ bounds=[(-10.0, 10.0)],
+ )
+ assert abs(result.x[0] - 2.0) < 0.01
+ assert result.fun < 0.01
+
+
+class TestOptimizerBrent:
+ """Brent scalar minimizer via native backend."""
+
+ def test_finds_correct_minimum(self):
+ # f(x) = (x - 3)^2
+ result = minimize_scalar_brent(
+ lambda x: (x - 3) ** 2,
+ bounds=(0.0, 10.0),
+ )
+ assert abs(result.x - 3.0) < 0.01
+ assert result.fun < 0.01
+
+ def test_converged_flag(self):
+ result = minimize_scalar_brent(
+ lambda x: (x - 5) ** 2,
+ bounds=(0.0, 10.0),
+ )
+ assert result.converged
+
+
diff --git a/tests/unit/test_formatters_edge.py b/tests/unit/test_formatters_edge.py
new file mode 100644
index 0000000..f52fd25
--- /dev/null
+++ b/tests/unit/test_formatters_edge.py
@@ -0,0 +1,230 @@
+"""Tests for formatter edge cases — scenario sample-size long format, cumulative recs, NaN filtering."""
+
+import math
+
+import pytest
+
+from mcpower.utils.formatters import _ResultFormatter, _is_nan
+
+
+_fmt = _ResultFormatter()
+
+
+def _make_scenario_sample_size_data(
+ target_tests=("x1", "x2"),
+ correction=None,
+ sample_sizes=(50, 100, 150),
+ optimistic_achieved=None,
+ realistic_achieved=None,
+ doomer_achieved=None,
+):
+ """Build a scenario sample_size result dict for formatting tests."""
+ if optimistic_achieved is None:
+ optimistic_achieved = {"x1": 50, "x2": 100}
+ if realistic_achieved is None:
+ realistic_achieved = {"x1": 100, "x2": 150}
+ if doomer_achieved is None:
+ doomer_achieved = {"x1": 0, "x2": 0} # Not achieved
+
+ def _make_scenario(achieved):
+ achieved_corr = {t: -1 for t in target_tests} if not correction else achieved
+ return {
+ "model": {
+ "target_tests": list(target_tests),
+ "correction": correction,
+ "sample_size_range": {"from_size": sample_sizes[0], "to_size": sample_sizes[-1]},
+ "target_power": 80.0,
+ },
+ "results": {
+ "first_achieved": achieved,
+ "first_achieved_corrected": achieved_corr,
+ "sample_sizes_tested": list(sample_sizes),
+ "powers_by_test": {
+ t: [30.0 + 25.0 * i for i in range(len(sample_sizes))]
+ for t in target_tests
+ },
+ "powers_by_test_corrected": (
+ {t: [25.0 + 25.0 * i for i in range(len(sample_sizes))] for t in target_tests}
+ if correction
+ else None
+ ),
+ },
+ }
+
+ return {
+ "analysis_type": "sample_size",
+ "scenarios": {
+ "optimistic": _make_scenario(optimistic_achieved),
+ "realistic": _make_scenario(realistic_achieved),
+ "doomer": _make_scenario(doomer_achieved),
+ },
+ "comparison": {},
+ }
+
+
+class TestScenarioSampleSizeLongFormat:
+ """Test _format_scenario_sample_size with summary='long'."""
+
+ def test_recommendations_present(self):
+ data = _make_scenario_sample_size_data()
+ output = _fmt.format("scenario_sample_size", data, "long")
+ assert "RECOMMENDATIONS" in output
+
+ def test_unachievable_tests_warning(self):
+ data = _make_scenario_sample_size_data(
+ doomer_achieved={"x1": 0, "x2": 0},
+ )
+ output = _fmt.format("scenario_sample_size", data, "long")
+ assert "Warning" in output or "may not achieve" in output
+
+ def test_realistic_recommendation_shown(self):
+ data = _make_scenario_sample_size_data(
+ realistic_achieved={"x1": 100, "x2": 150},
+ )
+ output = _fmt.format("scenario_sample_size", data, "long")
+ assert "150" in output # max N for realistic
+
+ def test_short_format_produces_table(self):
+ data = _make_scenario_sample_size_data()
+ output = _fmt.format("scenario_sample_size", data, "short")
+ assert "SCENARIO SUMMARY" in output
+
+ def test_with_correction(self):
+ data = _make_scenario_sample_size_data(correction="bonferroni")
+ output = _fmt.format("scenario_sample_size", data, "short")
+ assert "Opt(U)" in output or "Uncorrected" in output.lower() or "(U)" in output
+
+
+class TestCumulativeRecommendations:
+ """Test _format_cumulative_recommendations paths."""
+
+ def test_non_scenario_target_met(self):
+ data = {
+ "model": {
+ "target_tests": ["x1", "x2"],
+ "target_power": 80.0,
+ },
+ "results": {
+ "sample_sizes_tested": [50, 100, 150],
+ "powers_by_test": {
+ "x1": [60.0, 85.0, 95.0],
+ "x2": [70.0, 90.0, 98.0],
+ },
+ },
+ }
+ lines = _fmt._format_cumulative_recommendations(data, is_scenario=False)
+ joined = "\n".join(lines)
+ assert "N=" in joined # Found a sample size
+
+ def test_non_scenario_target_not_met(self):
+ data = {
+ "model": {
+ "target_tests": ["x1", "x2"],
+ "target_power": 80.0,
+ },
+ "results": {
+ "sample_sizes_tested": [50, 100],
+ "powers_by_test": {
+ "x1": [10.0, 20.0],
+ "x2": [15.0, 25.0],
+ },
+ },
+ }
+ lines = _fmt._format_cumulative_recommendations(data, is_scenario=False)
+ joined = "\n".join(lines)
+ assert ">100" in joined # Exceeded max tested
+
+ def test_scenario_recommendations(self):
+ data = _make_scenario_sample_size_data(
+ sample_sizes=(50, 100, 150, 200),
+ optimistic_achieved={"x1": 100, "x2": 150},
+ )
+ # Override powers so all > 80%
+ for scenario in data["scenarios"].values():
+ scenario["results"]["powers_by_test"] = {
+ "x1": [50.0, 85.0, 92.0, 98.0],
+ "x2": [40.0, 75.0, 88.0, 95.0],
+ }
+ lines = _fmt._format_cumulative_recommendations(data, is_scenario=True)
+ assert len(lines) > 0
+
+ def test_empty_scenarios(self):
+ data = {"scenarios": {}}
+ lines = _fmt._format_cumulative_recommendations(data, is_scenario=True)
+ assert lines == []
+
+ def test_no_results_key(self):
+ data = {}
+ lines = _fmt._format_cumulative_recommendations(data, is_scenario=False)
+ assert lines == []
+
+
+class TestNaNPowerFiltering:
+ """NaN power values in cumulative table should be filtered out."""
+
+ def test_nan_power_filtered_in_cumulative_sample_size_table(self):
+ lines = []
+ _fmt._add_cumulative_sample_size_table(
+ lines,
+ sample_sizes=[50, 100],
+ target_tests=["x1", "x2_nan"],
+ powers_by_test={
+ "x1": [50.0, 80.0],
+ "x2_nan": [float("nan"), float("nan")],
+ },
+ )
+ # Should still produce output for x1 (x2_nan filtered out)
+ output = "\n".join(lines)
+ assert "N=50" in output or "50" in output
+
+ def test_all_nan_produces_no_table(self):
+ lines = []
+ _fmt._add_cumulative_sample_size_table(
+ lines,
+ sample_sizes=[50],
+ target_tests=["x1"],
+ powers_by_test={"x1": [float("nan")]},
+ )
+ # All NaN → no valid tests → no table
+ assert len(lines) == 0
+
+
+class TestIsNan:
+ """Test _is_nan utility."""
+
+ def test_nan_float(self):
+ assert _is_nan(float("nan"))
+
+ def test_regular_float(self):
+ assert not _is_nan(42.0)
+
+ def test_non_float(self):
+ assert not _is_nan("nan")
+ assert not _is_nan(None)
+ assert not _is_nan(42)
+
+
+class TestExtractScenarioMeta:
+ """Test _extract_scenario_meta."""
+
+ def test_no_model_returns_none(self):
+ target_tests, correction = _fmt._extract_scenario_meta({"opt": {"results": {}}})
+ assert target_tests is None
+
+ def test_extracts_from_first_scenario(self):
+ scenarios = {
+ "optimistic": {
+ "model": {"target_tests": ["a", "b"], "correction": "holm"},
+ }
+ }
+ target_tests, correction = _fmt._extract_scenario_meta(scenarios)
+ assert target_tests == ["a", "b"]
+ assert correction == "holm"
+
+
+class TestFormatUnknownType:
+ """Unknown result type should raise."""
+
+ def test_unknown_result_type(self):
+ with pytest.raises(ValueError, match="Unknown result type"):
+ _fmt.format("nonexistent", {})
diff --git a/tests/unit/test_mixed_models_coverage.py b/tests/unit/test_mixed_models_coverage.py
new file mode 100644
index 0000000..bc99974
--- /dev/null
+++ b/tests/unit/test_mixed_models_coverage.py
@@ -0,0 +1,292 @@
+"""Tests for stats/mixed_models.py — statsmodels convergence, corrections, native wrappers.
+
+Uses pytest.mark.lme to skip when statsmodels is not installed.
+"""
+
+import warnings
+from unittest.mock import MagicMock, patch
+
+import numpy as np
+import pytest
+
+from mcpower.stats.mixed_models import (
+ _ensure_lme_crits,
+ _lme_analysis_wrapper,
+ _wrap_native_result,
+ reset_warm_start_cache,
+)
+
+pytestmark = pytest.mark.lme
+
+
+class TestWrapNativeResult:
+ """Test _wrap_native_result helper."""
+
+ def test_non_empty_non_verbose(self):
+ result = np.array([1.0, 0.0, 1.0])
+ wrapped = _wrap_native_result(result, verbose=False, solver_name="native_q1")
+ np.testing.assert_array_equal(wrapped, result)
+
+ def test_non_empty_verbose(self):
+ result = np.array([1.0, 0.0, 1.0])
+ wrapped = _wrap_native_result(result, verbose=True, solver_name="native_q1")
+ assert isinstance(wrapped, dict)
+ assert "results" in wrapped
+ assert "diagnostics" in wrapped
+ assert wrapped["diagnostics"]["solver"] == "native_q1"
+
+ def test_non_empty_verbose_with_extra_diag(self):
+ result = np.array([1.0])
+ wrapped = _wrap_native_result(
+ result, verbose=True, solver_name="native_general",
+ extra_diag={"q": 3},
+ )
+ assert wrapped["diagnostics"]["q"] == 3
+
+ def test_empty_non_verbose_returns_none(self):
+ result = np.array([])
+ assert _wrap_native_result(result, verbose=False, solver_name="native_q1") is None
+
+ def test_empty_verbose_returns_failure_dict(self):
+ result = np.array([])
+ wrapped = _wrap_native_result(result, verbose=True, solver_name="native_q1")
+ assert wrapped["results"] is None
+ assert "failure_reason" in wrapped
+ assert "empty result" in wrapped["failure_reason"]
+
+
+class TestEnsureLMECrits:
+ """Test _ensure_lme_crits computes when None."""
+
+ def test_computes_when_none(self):
+ chi2, z, crits = _ensure_lme_crits(
+ alpha=0.05, p=3, n_targets=2, correction_method=0,
+ chi2_crit=None, z_crit=None, correction_z_crits=None,
+ )
+ assert np.isfinite(chi2)
+ assert np.isfinite(z)
+ assert len(crits) == 2
+
+ def test_passthrough_when_provided(self):
+ chi2, z, crits = _ensure_lme_crits(
+ alpha=0.05, p=3, n_targets=2, correction_method=0,
+ chi2_crit=7.8, z_crit=1.96, correction_z_crits=np.array([1.96, 1.96]),
+ )
+ assert chi2 == 7.8
+ assert z == 1.96
+ assert len(crits) == 2
+
+
+class TestLMEAnalysisWrapperRouting:
+ """Test _lme_analysis_wrapper routes to correct backend."""
+
+ def test_unknown_backend_raises(self):
+ with pytest.raises(ValueError, match="Unknown backend"):
+ _lme_analysis_wrapper(
+ np.eye(10), np.ones(10), np.array([0, 1]),
+ np.zeros(10, dtype=np.int32),
+ correction_method=0, alpha=0.05, backend="nonexistent",
+ )
+
+
+class TestStatsmodelsConvergence:
+ """Test statsmodels fallback path with mocked MixedLM."""
+
+ def _make_mock_result(self, converged=True, params=None, pvalues=None, n_params=3):
+ """Create a mock MixedLM result."""
+ result = MagicMock()
+ result.converged = converged
+ result.params = params if params is not None else np.array([1.0, 0.5, 0.3])
+ result.pvalues = pvalues if pvalues is not None else np.array([0.01, 0.02, 0.04])
+ result.fe_params = result.params
+ result.bse = np.array([0.1, 0.1, 0.1])
+
+ # cov_re: random effects variance (needs .iloc[0, 0])
+ cov_re = MagicMock()
+ cov_re.iloc.__getitem__ = MagicMock(return_value=0.5)
+ result.cov_re = cov_re
+
+ result.scale = 1.0
+ result.llf = -50.0
+
+ # Make cov_params return a proper matrix
+ result.cov_params.return_value = np.eye(n_params) * 0.01
+
+ # model attribute
+ result.model = MagicMock()
+ result.model.exog = MagicMock()
+ result.model.exog.shape = (100, n_params)
+
+ return result
+
+ @patch("statsmodels.regression.mixed_linear_model.MixedLM")
+ def test_warm_start_retry_chain(self, mock_mixedlm_cls):
+ """First fit fails, cold start succeeds."""
+ from mcpower.stats.mixed_models import _lme_analysis_statsmodels, _lme_thread_local
+
+ _lme_thread_local.warm_start_params = np.array([1.0, 0.5, 0.3])
+
+ mock_model = MagicMock()
+ mock_mixedlm_cls.return_value = mock_model
+
+ good_result = self._make_mock_result()
+ mock_model.fit.side_effect = [
+ Exception("warm start diverged"),
+ good_result,
+ ]
+ mock_model.loglike.return_value = -50.0
+
+ result = _lme_analysis_statsmodels(
+ X_expanded=np.random.randn(100, 2),
+ y=np.random.randn(100),
+ target_indices=np.array([0, 1]),
+ cluster_ids=np.repeat(np.arange(10), 10),
+
+ correction_method=0,
+ alpha=0.05,
+ )
+ assert result is not None
+
+ @patch("statsmodels.regression.mixed_linear_model.MixedLM")
+ def test_all_attempts_fail_returns_none(self, mock_mixedlm_cls):
+ from mcpower.stats.mixed_models import _lme_analysis_statsmodels, _lme_thread_local
+
+ _lme_thread_local.warm_start_params = None
+
+ mock_model = MagicMock()
+ mock_mixedlm_cls.return_value = mock_model
+ mock_model.fit.side_effect = Exception("always fails")
+
+ result = _lme_analysis_statsmodels(
+ X_expanded=np.random.randn(100, 2),
+ y=np.random.randn(100),
+ target_indices=np.array([0, 1]),
+ cluster_ids=np.repeat(np.arange(10), 10),
+
+ correction_method=0,
+ alpha=0.05,
+ )
+ assert result is None
+
+ @patch("statsmodels.regression.mixed_linear_model.MixedLM")
+ def test_all_attempts_fail_verbose_returns_dict(self, mock_mixedlm_cls):
+ from mcpower.stats.mixed_models import _lme_analysis_statsmodels, _lme_thread_local
+
+ _lme_thread_local.warm_start_params = None
+
+ mock_model = MagicMock()
+ mock_mixedlm_cls.return_value = mock_model
+ mock_model.fit.side_effect = Exception("always fails")
+
+ result = _lme_analysis_statsmodels(
+ X_expanded=np.random.randn(100, 2),
+ y=np.random.randn(100),
+ target_indices=np.array([0, 1]),
+ cluster_ids=np.repeat(np.arange(10), 10),
+
+ correction_method=0,
+ alpha=0.05,
+ verbose=True,
+ )
+ assert isinstance(result, dict)
+ assert result["results"] is None
+ assert "failure_reason" in result
+
+ @patch("statsmodels.regression.mixed_linear_model.MixedLM")
+ def test_not_converged_returns_none(self, mock_mixedlm_cls):
+ """When result.converged is False for all attempts."""
+ from mcpower.stats.mixed_models import _lme_analysis_statsmodels, _lme_thread_local
+
+ _lme_thread_local.warm_start_params = None
+
+ mock_model = MagicMock()
+ mock_mixedlm_cls.return_value = mock_model
+
+ bad_result = self._make_mock_result(converged=False)
+ mock_model.fit.return_value = bad_result
+
+ result = _lme_analysis_statsmodels(
+ X_expanded=np.random.randn(100, 2),
+ y=np.random.randn(100),
+ target_indices=np.array([0, 1]),
+ cluster_ids=np.repeat(np.arange(10), 10),
+
+ correction_method=0,
+ alpha=0.05,
+ )
+ assert result is None
+
+
+class TestCorrections:
+ """Test statsmodels FDR, Holm, Bonferroni, no-correction paths."""
+
+ def _make_mock_result(self):
+ result = MagicMock()
+ result.converged = True
+ result.params = np.array([1.0, 0.5, 0.3])
+ result.pvalues = np.array([0.001, 0.02, 0.04])
+ result.fe_params = result.params
+ result.bse = np.array([0.1, 0.1, 0.1])
+ result.scale = 1.0
+ result.llf = -50.0
+ result.model = MagicMock()
+ result.model.exog = MagicMock()
+ result.model.exog.shape = (100, 3)
+
+ cov_re_mock = MagicMock()
+ cov_re_mock.iloc.__getitem__ = MagicMock(return_value=0.5)
+ result.cov_re = cov_re_mock
+ result.cov_params.return_value = np.eye(3) * 0.01
+
+ return result
+
+ def _run_with_correction(self, correction_method):
+ from mcpower.stats.mixed_models import _lme_analysis_statsmodels, _lme_thread_local
+
+ _lme_thread_local.warm_start_params = None
+
+ mock_result = self._make_mock_result()
+
+ with patch("statsmodels.regression.mixed_linear_model.MixedLM") as mock_cls:
+ mock_model = MagicMock()
+ mock_cls.return_value = mock_model
+ mock_model.fit.return_value = mock_result
+ mock_model.loglike.return_value = -50.0
+
+ out = _lme_analysis_statsmodels(
+ X_expanded=np.random.randn(100, 2),
+ y=np.random.randn(100),
+ target_indices=np.array([0, 1]),
+ cluster_ids=np.repeat(np.arange(10), 10),
+
+ correction_method=correction_method,
+ alpha=0.05,
+ )
+ return out
+
+ def test_no_correction(self):
+ result = self._run_with_correction(0)
+ assert result is not None
+
+ def test_bonferroni(self):
+ result = self._run_with_correction(1)
+ assert result is not None
+
+ def test_fdr(self):
+ result = self._run_with_correction(2)
+ assert result is not None
+
+ def test_holm(self):
+ result = self._run_with_correction(3)
+ assert result is not None
+
+
+class TestResetWarmStartCache:
+ """Test reset_warm_start_cache."""
+
+ def test_clears_params(self):
+ from mcpower.stats.mixed_models import _lme_thread_local
+
+ _lme_thread_local.warm_start_params = np.array([1.0])
+ reset_warm_start_cache()
+ assert _lme_thread_local.warm_start_params is None
diff --git a/tests/unit/test_model_coverage.py b/tests/unit/test_model_coverage.py
new file mode 100644
index 0000000..23d47c9
--- /dev/null
+++ b/tests/unit/test_model_coverage.py
@@ -0,0 +1,117 @@
+"""Tests for model.py — parallel fallback, Tukey validation, NaN under Tukey correction."""
+
+import warnings
+from unittest.mock import MagicMock, patch
+
+import numpy as np
+import pytest
+
+from mcpower import MCPower
+
+
+class TestTukeyWithoutPosthoc:
+ """Tukey correction without posthoc specs should raise ValueError."""
+
+ def test_tukey_without_posthoc_raises(self):
+ model = MCPower("y = x1 + x2")
+ model.set_effects("x1=0.5, x2=0.3")
+
+ with pytest.raises(ValueError, match="Tukey correction requires"):
+ model.find_power(
+ sample_size=100,
+ correction="tukey",
+ print_results=False,
+ )
+
+
+class TestTukeyNaNification:
+ """Non-posthoc tests should be NaN-ified under Tukey correction."""
+
+ def test_non_posthoc_tests_nan_under_tukey(self):
+ model = MCPower("y = group + x1")
+ model.set_variable_type("group=(factor,3)")
+ model.set_effects("group[2]=0.5, group[3]=0.4, x1=0.3")
+ model.n_simulations = 50
+ model.seed = 42
+
+ result = model.find_sample_size(
+ target_test="all, all-posthoc",
+ correction="tukey",
+ from_size=30,
+ to_size=60,
+ by=30,
+ print_results=False,
+ return_results=True,
+ )
+
+ assert result is not None
+ results = result["results"]
+ corrected = results.get("powers_by_test_corrected", {})
+
+ # Post-hoc comparisons should have real power values
+ # Non-posthoc tests (like "x1", "group[2]", "group[3]", "overall")
+ # should have NaN values
+ posthoc_labels = {s.label for s in model._posthoc_specs}
+ for test_name, powers in corrected.items():
+ if test_name not in posthoc_labels:
+ assert all(isinstance(v, float) and np.isnan(v) for v in powers), \
+ f"Expected NaN for non-posthoc test '{test_name}', got {powers}"
+
+ # first_achieved_corrected for non-posthoc should be -1
+ for test_name, n in results.get("first_achieved_corrected", {}).items():
+ if test_name not in posthoc_labels:
+ assert n == -1, f"Expected -1 for '{test_name}', got {n}"
+
+
+class TestParallelFallback:
+ """Parallel execution falls back to sequential on exception."""
+
+ def test_parallel_exception_falls_back(self, capsys):
+ model = MCPower("y = x1 + x2")
+ model.set_effects("x1=0.5, x2=0.3")
+ model.parallel = True
+ model.n_simulations = 50
+ model.seed = 42
+
+ # Parallel is imported inside the function via `from joblib import Parallel`,
+ # so we patch it at the joblib module level.
+ with patch("joblib.Parallel", side_effect=RuntimeError("joblib broken")):
+ # Should still complete via sequential fallback
+ result = model.find_sample_size(
+ from_size=30,
+ to_size=60,
+ by=30,
+ print_results=False,
+ return_results=True,
+ )
+ assert result is not None
+ captured = capsys.readouterr()
+ assert "Falling back to sequential" in captured.out
+
+
+class TestIsParallelEffective:
+ """Test _is_parallel_effective resolution."""
+
+ def test_true_always_parallel(self):
+ model = MCPower("y = x1 + x2")
+ model.parallel = True
+ assert model._is_parallel_effective() is True
+
+ def test_false_never_parallel(self):
+ model = MCPower("y = x1 + x2")
+ model.parallel = False
+ assert model._is_parallel_effective() is False
+
+ def test_mixedmodels_with_clusters(self):
+ model = MCPower("y ~ x1 + (1|school)")
+ model.set_cluster("school", ICC=0.2, n_clusters=20)
+ model.set_effects("x1=0.5")
+ model._apply() # cluster_specs are deferred until apply()
+ model.parallel = "mixedmodels"
+ assert model._is_parallel_effective() is True
+
+ def test_mixedmodels_without_clusters(self):
+ model = MCPower("y = x1 + x2")
+ model.set_effects("x1=0.5, x2=0.3")
+ model.parallel = "mixedmodels"
+ assert model._is_parallel_effective() is False
diff --git a/tests/unit/test_native_backend.py b/tests/unit/test_native_backend.py
new file mode 100644
index 0000000..93b4a3e
--- /dev/null
+++ b/tests/unit/test_native_backend.py
@@ -0,0 +1,60 @@
+"""Tests for mcpower.backends.native — import fallback and _prep utility."""
+
+import numpy as np
+import pytest
+from unittest.mock import patch, MagicMock
+
+from mcpower.backends.native import _prep
+
+
+class TestPrep:
+ """Test _prep array coercion for C++ interop."""
+
+ def test_contiguous_passthrough(self):
+ arr = np.array([1.0, 2.0, 3.0], dtype=np.float64)
+ result = _prep(arr)
+ assert result.flags["C_CONTIGUOUS"]
+ assert result.dtype == np.float64
+
+ def test_non_contiguous_becomes_contiguous(self):
+ arr = np.array([[1.0, 2.0], [3.0, 4.0]], dtype=np.float64)
+ col = arr[:, 1] # non-contiguous column slice
+ assert not col.flags["C_CONTIGUOUS"]
+ result = _prep(col)
+ assert result.flags["C_CONTIGUOUS"]
+ np.testing.assert_array_equal(result, [2.0, 4.0])
+
+ def test_dtype_conversion_float32_to_float64(self):
+ arr = np.array([1.0, 2.0], dtype=np.float32)
+ result = _prep(arr, np.float64)
+ assert result.dtype == np.float64
+
+ def test_dtype_conversion_int64_to_int32(self):
+ arr = np.array([0, 1, 2], dtype=np.int64)
+ result = _prep(arr, np.int32)
+ assert result.dtype == np.int32
+ np.testing.assert_array_equal(result, [0, 1, 2])
+
+ def test_2d_array(self):
+ arr = np.array([[1.0, 2.0], [3.0, 4.0]], dtype=np.float64, order="F")
+ assert not arr.flags["C_CONTIGUOUS"]
+ result = _prep(arr)
+ assert result.flags["C_CONTIGUOUS"]
+ assert result.dtype == np.float64
+
+
+class TestNativeBackendImport:
+ """Test NativeBackend init when C++ extension is unavailable."""
+
+ def test_init_raises_when_unavailable(self):
+ """NativeBackend() should raise ImportError when _NATIVE_AVAILABLE=False."""
+ with patch("mcpower.backends.native._NATIVE_AVAILABLE", False):
+ from mcpower.backends.native import NativeBackend
+ with pytest.raises(ImportError, match="Native C\\+\\+ backend not available"):
+ NativeBackend()
+
+ def test_is_native_available_reflects_module_state(self):
+ from mcpower.backends.native import is_native_available
+ # Just verify it returns a bool
+ result = is_native_available()
+ assert isinstance(result, bool)
diff --git a/tests/unit/test_ols_corrections.py b/tests/unit/test_ols_corrections.py
new file mode 100644
index 0000000..8c6728a
--- /dev/null
+++ b/tests/unit/test_ols_corrections.py
@@ -0,0 +1,251 @@
+"""Tests for OLS post-hoc contrast corrections and edge cases."""
+
+from dataclasses import dataclass
+from typing import Optional
+
+import numpy as np
+import pytest
+
+from mcpower.stats.ols import compute_posthoc_contrasts
+
+
+@dataclass
+class _PostHocSpec:
+ """Minimal PostHocSpec stub for tests."""
+ factor_name: str
+ col_idx_a: Optional[int]
+ col_idx_b: Optional[int]
+ label: str = ""
+ level_a: str = ""
+ level_b: str = ""
+ n_levels: int = 3
+
+
+def _make_ols_data(n=100, p=3, seed=42):
+ """Generate simple OLS data: X, y, and target_indices."""
+ rng = np.random.RandomState(seed)
+ X = rng.randn(n, p)
+ beta = np.array([0.5, 0.3, -0.2])[:p]
+ y = X @ beta + rng.randn(n)
+ return X, y
+
+
+class TestDegenerateDesign:
+ """When dof <= 0, posthoc should return zeros."""
+
+ def test_dof_zero_returns_zeros(self):
+ # n = p+1 → dof = 0
+ n, p = 4, 3
+ rng = np.random.RandomState(42)
+ X = rng.randn(n, p)
+ y = rng.randn(n)
+ specs = [_PostHocSpec("grp", 0, 1)]
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {}, target_indices=np.array([0, 1, 2]),
+ )
+ assert uncorr.shape == (1,)
+ assert not uncorr[0]
+ assert not corr[0]
+ assert override is None
+
+ def test_singular_contrast_variance_stays_zero(self):
+ """When both col_idx_a and col_idx_b are None, t_abs stays 0."""
+ X, y = _make_ols_data()
+ specs = [_PostHocSpec("grp", None, None)]
+
+ uncorr, corr, _ = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {},
+ )
+ assert not uncorr[0]
+ assert not corr[0]
+
+
+class TestCombinedFDR:
+ """FDR (correction_method=2) step-up across regular+posthoc t-stats."""
+
+ def test_fdr_combined_ranking(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [
+ _PostHocSpec("grp", 0, 1),
+ _PostHocSpec("grp", 0, 2),
+ ]
+ target_indices = np.array([0, 1, 2])
+ # Create combined crits of length n_regular + n_posthoc = 5
+ # Use very lenient crits so everything passes
+ combined_crits = np.full(5, 0.01)
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 0.01, {},
+ target_indices=target_indices,
+ correction_method=2,
+ correction_t_crits_combined=combined_crits,
+ )
+ assert override is not None
+ assert len(override) == 3 # n_regular
+ assert len(corr) == 2 # n_posthoc
+
+ def test_fdr_no_significant(self):
+ """With very strict crits, nothing should be significant."""
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+ # Very strict thresholds
+ combined_crits = np.full(4, 100.0)
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 100.0, {},
+ target_indices=target_indices,
+ correction_method=2,
+ correction_t_crits_combined=combined_crits,
+ )
+ assert not np.any(corr)
+ assert override is not None
+ assert not np.any(override)
+
+
+class TestCombinedHolm:
+ """Holm (correction_method=3) step-down with early termination."""
+
+ def test_holm_combined_ranking(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+ combined_crits = np.full(4, 0.01) # Very lenient
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 0.01, {},
+ target_indices=target_indices,
+ correction_method=3,
+ correction_t_crits_combined=combined_crits,
+ )
+ assert override is not None
+ assert len(override) == 3
+
+ def test_holm_early_termination(self):
+ """If the most significant test doesn't pass, none should."""
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+ combined_crits = np.full(4, 1000.0) # Impossible threshold
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 1000.0, {},
+ target_indices=target_indices,
+ correction_method=3,
+ correction_t_crits_combined=combined_crits,
+ )
+ assert not np.any(corr)
+
+
+class TestFallbackPaths:
+ """Fallback when correction_t_crits_combined is None or wrong length."""
+
+ def test_combined_crits_none_fallback(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {},
+ target_indices=target_indices,
+ correction_method=2,
+ correction_t_crits_combined=None,
+ )
+ # Fallback: corrected = uncorrected copy, no override
+ np.testing.assert_array_equal(corr, uncorr)
+ assert override is None
+
+ def test_combined_crits_wrong_length_fallback(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+ # Wrong length: should be 4 (3 regular + 1 posthoc)
+ wrong_crits = np.full(2, 2.0)
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {},
+ target_indices=target_indices,
+ correction_method=2,
+ correction_t_crits_combined=wrong_crits,
+ )
+ np.testing.assert_array_equal(corr, uncorr)
+ assert override is None
+
+
+class TestTukeyMethod:
+ """Tukey post-hoc method path."""
+
+ def test_tukey_uses_factor_crit(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1, n_levels=3)]
+ tukey_crits = {"grp": 0.01} # Very lenient
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "tukey", 2.0, tukey_crits,
+ )
+ # Tukey correction: uncorrected == corrected
+ np.testing.assert_array_equal(uncorr, corr)
+ assert override is None
+
+ def test_tukey_missing_factor_uses_inf(self):
+ """When factor not in tukey_crits, inf is used → not significant."""
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("missing_factor", 0, 1)]
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "tukey", 2.0, {},
+ )
+ assert not uncorr[0]
+ assert not corr[0]
+
+
+class TestBonferroniPosthoc:
+ """Bonferroni correction for posthoc (correction_method=1)."""
+
+ def test_bonferroni_uses_combined_first_crit(self):
+ X, y = _make_ols_data(n=200, p=3, seed=10)
+ specs = [_PostHocSpec("grp", 0, 1)]
+ target_indices = np.array([0, 1, 2])
+ combined_crits = np.full(4, 0.01) # Very lenient
+
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 0.01, {},
+ target_indices=target_indices,
+ correction_method=1,
+ correction_t_crits_combined=combined_crits,
+ )
+ assert override is None # Bonferroni doesn't produce override
+
+
+class TestEmptySpecs:
+ """Empty posthoc specs return empty arrays."""
+
+ def test_no_specs(self):
+ X, y = _make_ols_data()
+ uncorr, corr, override = compute_posthoc_contrasts(
+ X, y, [], "t-test", 2.0, {},
+ )
+ assert len(uncorr) == 0
+ assert len(corr) == 0
+ assert override is None
+
+
+class TestSingleColumnContrasts:
+ """Contrasts where one side is the reference level (None)."""
+
+ def test_col_idx_a_none(self):
+ X, y = _make_ols_data(n=200)
+ specs = [_PostHocSpec("grp", None, 1)]
+ uncorr, corr, _ = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {},
+ )
+ assert uncorr.shape == (1,)
+
+ def test_col_idx_b_none(self):
+ X, y = _make_ols_data(n=200)
+ specs = [_PostHocSpec("grp", 0, None)]
+ uncorr, corr, _ = compute_posthoc_contrasts(
+ X, y, specs, "t-test", 2.0, {},
+ )
+ assert uncorr.shape == (1,)
diff --git a/tests/unit/test_parsers_errors.py b/tests/unit/test_parsers_errors.py
new file mode 100644
index 0000000..b3c8c01
--- /dev/null
+++ b/tests/unit/test_parsers_errors.py
@@ -0,0 +1,168 @@
+"""Tests for parser error paths and edge cases."""
+
+import pytest
+
+from mcpower.utils.parsers import _AssignmentParser, _parse_equation
+
+
+_parser = _AssignmentParser()
+
+
+class TestAssignmentParserErrors:
+ """Error paths in _AssignmentParser._parse."""
+
+ def test_missing_equals_sign(self):
+ parsed, errors = _parser._parse("x1 0.5", "effect", ["x1"])
+ assert len(errors) == 1
+ assert "Invalid format" in errors[0]
+
+ def test_unknown_parse_type(self):
+ parsed, errors = _parser._parse("x1=0.5", "unknown_type", ["x1"])
+ assert len(errors) == 1
+ assert "Unknown parse type" in errors[0]
+
+ def test_unavailable_variable(self):
+ parsed, errors = _parser._parse("x_missing=0.5", "effect", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "not found" in errors[0]
+ assert "x_missing" in errors[0]
+
+ def test_invalid_effect_value(self):
+ parsed, errors = _parser._parse("x1=abc", "effect", ["x1"])
+ assert len(errors) == 1
+ assert "Invalid effect size" in errors[0]
+
+ def test_multiple_errors(self):
+ parsed, errors = _parser._parse("x_bad=abc, x_also_bad=xyz", "effect", ["x1"])
+ assert len(errors) == 2
+
+
+class TestCorrelationParserErrors:
+ """Error paths for correlation parsing."""
+
+ def test_invalid_correlation_format(self):
+ parsed, errors = _parser._parse("x1_x2=0.5", "correlation", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "Invalid format" in errors[0] or "Invalid correlation" in errors[0]
+
+ def test_correlation_var_not_found(self):
+ parsed, errors = _parser._parse("corr(x1, x_missing)=0.5", "correlation", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "not found" in errors[0]
+
+ def test_self_correlation(self):
+ parsed, errors = _parser._parse("corr(x1, x1)=0.5", "correlation", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "Cannot correlate variable with itself" in errors[0]
+
+ def test_correlation_value_out_of_range(self):
+ parsed, errors = _parser._parse("corr(x1, x2)=1.5", "correlation", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "between -1 and 1" in errors[0]
+
+ def test_invalid_correlation_value(self):
+ parsed, errors = _parser._parse("corr(x1, x2)=abc", "correlation", ["x1", "x2"])
+ assert len(errors) == 1
+ assert "Invalid correlation value" in errors[0]
+
+
+class TestVariableTypeErrors:
+ """Error paths for variable type parsing."""
+
+ def test_unsupported_type(self):
+ parsed, errors = _parser._parse("x1=crazy_type", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "Unsupported type" in errors[0]
+
+ def test_binary_proportion_out_of_range(self):
+ parsed, errors = _parser._parse("x1=(binary,1.5)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "between 0 and 1" in errors[0]
+
+ def test_binary_non_numeric_proportion(self):
+ parsed, errors = _parser._parse("x1=(binary,abc)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "Invalid proportion" in errors[0]
+
+ def test_binary_wrong_param_count(self):
+ parsed, errors = _parser._parse("x1=(binary,0.3,0.4)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "exactly 2 values" in errors[0]
+
+ def test_factor_less_than_2_levels(self):
+ parsed, errors = _parser._parse("x1=(factor,1)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "at least 2 levels" in errors[0]
+
+ def test_factor_more_than_20_levels(self):
+ parsed, errors = _parser._parse("x1=(factor,21)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "more than 20 levels" in errors[0]
+
+ def test_factor_non_integer_levels(self):
+ parsed, errors = _parser._parse("x1=(factor,abc)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "Must be integer" in errors[0]
+
+ def test_factor_proportions_more_than_20(self):
+ props = ",".join(["0.04"] * 21)
+ parsed, errors = _parser._parse(f"x1=(factor,{props})", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "more than 20 levels" in errors[0]
+
+ def test_factor_zero_proportion(self):
+ parsed, errors = _parser._parse("x1=(factor,0.5,0.0,0.5)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "positive" in errors[0]
+
+ def test_factor_non_numeric_proportions(self):
+ parsed, errors = _parser._parse("x1=(factor,abc,def)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "numeric" in errors[0]
+
+ def test_tuple_no_comma(self):
+ parsed, errors = _parser._parse("x1=(binary)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "Invalid tuple format" in errors[0]
+
+ def test_tuple_unsupported_type_in_tuple(self):
+ parsed, errors = _parser._parse("x1=(normal,0.5)", "variable_type", ["x1"])
+ assert len(errors) == 1
+ assert "only supported for binary and factor" in errors[0]
+
+
+class TestEquationParsing:
+ """Edge cases in _parse_equation."""
+
+ def test_nested_random_effects(self):
+ dep, formula, ranefs = _parse_equation("y ~ x1 + (1|A/B)")
+ assert dep == "y"
+ assert len(ranefs) == 2
+ group_vars = {r["grouping_var"] for r in ranefs}
+ assert "A" in group_vars
+ assert "A:B" in group_vars
+
+ def test_duplicate_grouping_var_raises(self):
+ with pytest.raises(ValueError, match="Duplicate random effect grouping variable"):
+ _parse_equation("y ~ x1 + (1|school) + (1|school)")
+
+ def test_random_slopes(self):
+ dep, formula, ranefs = _parse_equation("y ~ x1 + (1 + x1|school)")
+ assert len(ranefs) == 1
+ assert ranefs[0]["type"] == "random_slope"
+ assert ranefs[0]["slope_vars"] == ["x1"]
+ assert ranefs[0]["grouping_var"] == "school"
+
+ def test_random_slope_duplicate_grouping_raises(self):
+ with pytest.raises(ValueError, match="Duplicate"):
+ _parse_equation("y ~ x1 + (1|school) + (1 + x1|school)")
+
+ def test_no_separator_uses_default_dep(self):
+ dep, formula, ranefs = _parse_equation("x1+x2")
+ assert dep == "explained_variable"
+ assert "x1" in formula
+ assert "x2" in formula
+
+ def test_nested_duplicate_parent_raises(self):
+ with pytest.raises(ValueError, match="Duplicate"):
+ _parse_equation("y ~ (1|A) + (1|A/B)")
diff --git a/tests/unit/test_progress.py b/tests/unit/test_progress.py
index 329acca..768420a 100644
--- a/tests/unit/test_progress.py
+++ b/tests/unit/test_progress.py
@@ -127,12 +127,6 @@ def test_completion_newline(self):
class TestTqdmReporter:
"""Test TqdmReporter with mock tqdm."""
- def test_tqdm_missing_raises(self):
- reporter = TqdmReporter()
- with patch.dict("sys.modules", {"tqdm": None}):
- with pytest.raises(ImportError, match="tqdm"):
- reporter(0, 100)
-
def test_tqdm_basic_flow(self):
mock_bar = MagicMock()
mock_bar.n = 0
@@ -154,6 +148,51 @@ def test_tqdm_basic_flow(self):
reporter(100, 100) # closes
mock_bar.close.assert_called_once()
+ def test_tqdm_successive_sessions(self):
+ """After close, a new session creates a fresh bar."""
+ mock_bar = MagicMock()
+ mock_bar.n = 0
+ mock_tqdm_cls = MagicMock(return_value=mock_bar)
+ mock_tqdm_module = MagicMock()
+ mock_tqdm_module.tqdm = mock_tqdm_cls
+
+ reporter = TqdmReporter()
+
+ with patch.dict("sys.modules", {"tqdm": mock_tqdm_module}):
+ # First session
+ reporter(0, 50)
+ mock_bar.n = 0
+ reporter(50, 50)
+ mock_bar.close.assert_called_once()
+ assert reporter._bar is None
+
+ # Second session — should create a new bar
+ mock_tqdm_cls.reset_mock()
+ mock_bar2 = MagicMock()
+ mock_bar2.n = 0
+ mock_tqdm_cls.return_value = mock_bar2
+
+ reporter(0, 200)
+ assert mock_tqdm_cls.call_count == 1
+ mock_tqdm_cls.assert_called_with(total=200, unit="sim")
+
+ def test_tqdm_no_negative_delta(self):
+ """When current <= bar.n, update should not be called with negative delta."""
+ mock_bar = MagicMock()
+ mock_bar.n = 50
+ mock_tqdm_cls = MagicMock(return_value=mock_bar)
+ mock_tqdm_module = MagicMock()
+ mock_tqdm_module.tqdm = mock_tqdm_cls
+
+ reporter = TqdmReporter()
+
+ with patch.dict("sys.modules", {"tqdm": mock_tqdm_module}):
+ reporter(0, 100) # creates bar
+ mock_bar.n = 50
+ reporter(30, 100) # current < bar.n
+ # update should NOT have been called (delta = 30 - 50 = -20, not > 0)
+ mock_bar.update.assert_not_called()
+
class TestComputeTotalSimulations:
"""Test compute_total_simulations helper."""
diff --git a/tests/unit/test_results.py b/tests/unit/test_results.py
new file mode 100644
index 0000000..31c3082
--- /dev/null
+++ b/tests/unit/test_results.py
@@ -0,0 +1,138 @@
+"""Unit tests for mcpower.core.results — ResultsProcessor and builder functions."""
+
+import numpy as np
+import pytest
+
+from mcpower.core.results import ResultsProcessor, build_power_result, build_sample_size_result
+
+
+class TestCalculatePowers:
+ """Tests for ResultsProcessor.calculate_powers."""
+
+ def test_basic_two_tests(self):
+ """Power calculation with two tests (overall + one predictor)."""
+ proc = ResultsProcessor(target_power=80.0)
+ # 10 simulations, 2 columns: [overall, x1]
+ # overall: 8/10 sig, x1: 6/10 sig
+ results = [np.array([True, True])] * 6 + [
+ np.array([True, False]),
+ np.array([True, False]),
+ np.array([False, False]),
+ np.array([False, False]),
+ ]
+ corrected = results # same for this test
+
+ out = proc.calculate_powers(results, corrected, ["overall", "x1"])
+
+ assert out["individual_powers"]["overall"] == pytest.approx(80.0)
+ assert out["individual_powers"]["x1"] == pytest.approx(60.0)
+ assert out["n_simulations_used"] == 10
+
+ def test_all_significant(self):
+ proc = ResultsProcessor()
+ results = [np.array([True, True])] * 5
+ out = proc.calculate_powers(results, results, ["overall", "x1"])
+ assert out["individual_powers"]["overall"] == pytest.approx(100.0)
+ assert out["individual_powers"]["x1"] == pytest.approx(100.0)
+
+ def test_none_significant(self):
+ proc = ResultsProcessor()
+ results = [np.array([False, False])] * 5
+ out = proc.calculate_powers(results, results, ["overall", "x1"])
+ assert out["individual_powers"]["overall"] == pytest.approx(0.0)
+ assert out["individual_powers"]["x1"] == pytest.approx(0.0)
+
+ def test_combined_probabilities(self):
+ proc = ResultsProcessor()
+ # 4 sims, 2 tests: exactly 0, 1, 2 significant
+ results = [
+ np.array([False, False]), # 0 sig
+ np.array([True, False]), # 1 sig
+ np.array([False, True]), # 1 sig
+ np.array([True, True]), # 2 sig
+ ]
+ out = proc.calculate_powers(results, results, ["overall", "x1"])
+ combined = out["combined_probabilities"]
+ assert combined["exactly_0_significant"] == pytest.approx(25.0)
+ assert combined["exactly_1_significant"] == pytest.approx(50.0)
+ assert combined["exactly_2_significant"] == pytest.approx(25.0)
+
+ def test_cumulative_probabilities(self):
+ proc = ResultsProcessor()
+ results = [
+ np.array([False, False]),
+ np.array([True, True]),
+ np.array([True, True]),
+ np.array([True, True]),
+ ]
+ out = proc.calculate_powers(results, results, ["overall", "x1"])
+ cumulative = out["cumulative_probabilities"]
+ assert cumulative["at_least_0_significant"] == pytest.approx(100.0)
+ assert cumulative["at_least_2_significant"] == pytest.approx(75.0)
+
+
+class TestBuildPowerResult:
+ """Tests for build_power_result."""
+
+ def test_basic_structure(self):
+ power_results = {
+ "individual_powers": {"overall": 80.0},
+ "n_simulations_used": 1000,
+ }
+ result = build_power_result(
+ model_type="OLS",
+ target_tests=["overall"],
+ formula_to_test=None,
+ equation="y = x1",
+ sample_size=100,
+ alpha=0.05,
+ n_simulations=1000,
+ correction=None,
+ target_power=80.0,
+ parallel=False,
+ power_results=power_results,
+ )
+ assert result["model"]["model_type"] == "OLS"
+ assert result["model"]["sample_size"] == 100
+ assert result["model"]["alpha"] == 0.05
+ assert result["results"] is power_results
+
+
+class TestBuildSampleSizeResult:
+ """Tests for build_sample_size_result."""
+
+ def test_basic_structure(self):
+ analysis_results = {"sample_sizes_tested": [50, 100]}
+ result = build_sample_size_result(
+ model_type="OLS",
+ target_tests=["overall"],
+ formula_to_test=None,
+ equation="y = x1",
+ sample_sizes=[50, 100],
+ alpha=0.05,
+ n_simulations=1000,
+ correction=None,
+ target_power=80.0,
+ parallel=False,
+ analysis_results=analysis_results,
+ )
+ assert result["model"]["sample_size_range"]["from_size"] == 50
+ assert result["model"]["sample_size_range"]["to_size"] == 100
+ assert result["model"]["sample_size_range"]["by"] == 50
+ assert result["results"] is analysis_results
+
+ def test_single_sample_size(self):
+ result = build_sample_size_result(
+ model_type="OLS",
+ target_tests=["overall"],
+ formula_to_test=None,
+ equation="y = x1",
+ sample_sizes=[100],
+ alpha=0.05,
+ n_simulations=1000,
+ correction=None,
+ target_power=80.0,
+ parallel=False,
+ analysis_results={},
+ )
+ assert result["model"]["sample_size_range"]["by"] == 1
diff --git a/tests/unit/test_scenarios_coverage.py b/tests/unit/test_scenarios_coverage.py
new file mode 100644
index 0000000..ea0d4f2
--- /dev/null
+++ b/tests/unit/test_scenarios_coverage.py
@@ -0,0 +1,218 @@
+"""Tests for scenario analysis — plot creation, correlation matrix repair, LME perturbations."""
+
+from unittest.mock import MagicMock, patch
+
+import numpy as np
+import pytest
+
+from mcpower.core.scenarios import (
+ ScenarioRunner,
+ apply_lme_perturbations,
+ apply_per_simulation_perturbations,
+)
+
+
+class TestCorrelationMatrixRepair:
+ """Spectral clipping when noise creates negative eigenvalues."""
+
+ def test_negative_eigenvalue_repaired(self):
+ """After heavy noise, result should be positive semi-definite with unit diagonal."""
+ # Create a 3x3 identity correlation matrix
+ corr = np.eye(3)
+ var_types = np.zeros(3, dtype=np.int64) # all normal
+
+ config = {
+ "correlation_noise_sd": 2.0, # Very heavy noise → guaranteed negative eigenvalues
+ "distribution_change_prob": 0.0,
+ "new_distributions": [],
+ }
+
+ perturbed_corr, _ = apply_per_simulation_perturbations(corr, var_types, config, sim_seed=42)
+
+ # Eigenvalues should all be >= 0
+ eigvals = np.linalg.eigvalsh(perturbed_corr)
+ assert np.all(eigvals >= -1e-10)
+
+ # Diagonal should be 1.0
+ np.testing.assert_allclose(np.diag(perturbed_corr), 1.0, atol=1e-10)
+
+ # Should be symmetric
+ np.testing.assert_allclose(perturbed_corr, perturbed_corr.T, atol=1e-10)
+
+ def test_no_repair_needed_when_no_noise(self):
+ corr = np.array([[1.0, 0.3], [0.3, 1.0]])
+ var_types = np.zeros(2, dtype=np.int64)
+
+ config = {
+ "correlation_noise_sd": 0.0,
+ "distribution_change_prob": 0.0,
+ "new_distributions": [],
+ }
+
+ perturbed_corr, _ = apply_per_simulation_perturbations(corr, var_types, config, sim_seed=42)
+ np.testing.assert_array_equal(perturbed_corr, corr)
+
+
+class TestDistributionPerturbation:
+ """Variable type swaps in scenario mode."""
+
+ def test_distribution_swap_occurs(self):
+ var_types = np.zeros(10, dtype=np.int64) # All normal
+ config = {
+ "correlation_noise_sd": 0.0,
+ "distribution_change_prob": 1.0, # Always swap
+ "new_distributions": ["right_skewed"],
+ }
+
+ _, perturbed_types = apply_per_simulation_perturbations(
+ np.eye(10), var_types, config, sim_seed=42,
+ )
+ # All should be swapped from 0 to 2 (right_skewed)
+ assert np.all(perturbed_types == 2)
+
+ def test_non_normal_not_swapped(self):
+ """Binary (1) and uploaded (99) vars should not be swapped."""
+ var_types = np.array([0, 1, 99], dtype=np.int64)
+ config = {
+ "correlation_noise_sd": 0.0,
+ "distribution_change_prob": 1.0,
+ "new_distributions": ["right_skewed"],
+ }
+
+ _, perturbed_types = apply_per_simulation_perturbations(
+ np.eye(3), var_types, config, sim_seed=42,
+ )
+ assert perturbed_types[0] == 2 # normal → right_skewed
+ assert perturbed_types[1] == 1 # binary unchanged
+ assert perturbed_types[2] == 99 # uploaded unchanged
+
+ def test_none_config_passthrough(self):
+ corr = np.eye(2)
+ var_types = np.zeros(2, dtype=np.int64)
+ result_corr, result_types = apply_per_simulation_perturbations(
+ corr, var_types, None, sim_seed=42,
+ )
+ np.testing.assert_array_equal(result_corr, corr)
+ np.testing.assert_array_equal(result_types, var_types)
+
+
+class TestLMEPerturbations:
+ """LME perturbation computation."""
+
+ def test_icc_noise_creates_multipliers(self):
+ cluster_specs = {"school": {"n_clusters": 20, "cluster_size": 10, "icc": 0.2}}
+ config = {
+ "icc_noise_sd": 0.3,
+ "random_effect_dist": "normal",
+ "random_effect_df": 5,
+ }
+
+ result = apply_lme_perturbations(cluster_specs, config, sim_seed=42)
+ assert result is not None
+ assert "tau_squared_multipliers" in result
+ assert "school" in result["tau_squared_multipliers"]
+ # Multiplier should be exp(N(0, 0.3)) — positive, around 1
+ mult = result["tau_squared_multipliers"]["school"]
+ assert mult > 0
+
+ def test_no_perturbation_returns_none(self):
+ cluster_specs = {"school": {"n_clusters": 20, "cluster_size": 10, "icc": 0.2}}
+ config = {
+ "icc_noise_sd": 0.0,
+ "random_effect_dist": "normal",
+ "random_effect_df": 5,
+ }
+ result = apply_lme_perturbations(cluster_specs, config, sim_seed=42)
+ assert result is None
+
+ def test_empty_cluster_specs_returns_none(self):
+ result = apply_lme_perturbations({}, {"icc_noise_sd": 0.5}, sim_seed=42)
+ assert result is None
+
+ def test_heavy_tailed_re_dist(self):
+ cluster_specs = {"school": {"n_clusters": 20, "cluster_size": 10, "icc": 0.2}}
+ config = {
+ "icc_noise_sd": 0.0,
+ "random_effect_dist": "heavy_tailed",
+ "random_effect_df": 3,
+ }
+ result = apply_lme_perturbations(cluster_specs, config, sim_seed=42)
+ assert result is not None
+ assert result["random_effect_dist"] == "heavy_tailed"
+ assert result["random_effect_df"] == 3
+
+
+class TestScenarioRunnerPlots:
+ """Test _create_scenario_plots path."""
+
+ def test_plot_creation_with_mock(self):
+ model = MagicMock()
+ model.power = 80.0
+ runner = ScenarioRunner(model)
+
+ results = {
+ "analysis_type": "sample_size",
+ "scenarios": {
+ "optimistic": {
+ "model": {
+ "target_tests": ["x1"],
+ "correction": None,
+ },
+ "results": {
+ "sample_sizes_tested": [50, 100],
+ "powers_by_test": {"x1": [50.0, 85.0]},
+ "first_achieved": {"x1": 100},
+ },
+ },
+ },
+ }
+
+ with patch("mcpower.core.scenarios._create_power_plot") as mock_plot:
+ runner._create_scenario_plots(results)
+ mock_plot.assert_called_once()
+
+ def test_plot_with_correction(self):
+ model = MagicMock()
+ model.power = 80.0
+ runner = ScenarioRunner(model)
+
+ results = {
+ "analysis_type": "sample_size",
+ "scenarios": {
+ "optimistic": {
+ "model": {
+ "target_tests": ["x1"],
+ "correction": "bonferroni",
+ },
+ "results": {
+ "sample_sizes_tested": [50, 100],
+ "powers_by_test": {"x1": [50.0, 85.0]},
+ "powers_by_test_corrected": {"x1": [40.0, 75.0]},
+ "first_achieved": {"x1": 100},
+ "first_achieved_corrected": {"x1": 150},
+ },
+ },
+ },
+ }
+
+ with patch("mcpower.core.scenarios._create_power_plot") as mock_plot:
+ runner._create_scenario_plots(results)
+ # Should be called for both uncorrected and corrected
+ assert mock_plot.call_count == 2
+
+ def test_no_plot_when_missing_sample_sizes(self):
+ model = MagicMock()
+ model.power = 80.0
+ runner = ScenarioRunner(model)
+
+ results = {
+ "scenarios": {
+ "optimistic": {
+ "results": {"powers_by_test": {"x1": [50.0]}},
+ },
+ },
+ }
+
+ with patch("mcpower.core.scenarios._create_power_plot") as mock_plot:
+ runner._create_scenario_plots(results)
+ mock_plot.assert_not_called()
diff --git a/tests/unit/test_simulation_coverage.py b/tests/unit/test_simulation_coverage.py
new file mode 100644
index 0000000..8bae7ff
--- /dev/null
+++ b/tests/unit/test_simulation_coverage.py
@@ -0,0 +1,274 @@
+"""Tests for simulation.py — failure handling, Wald fallback, verbose diagnostics, ICC mismatch."""
+
+import warnings
+from typing import Dict, List, Optional
+from unittest.mock import MagicMock, patch
+
+import numpy as np
+import pytest
+
+from mcpower.core.simulation import SimulationMetadata, SimulationRunner, _warn_icc_mismatch
+
+
+def _make_metadata(
+ n_targets=2,
+ cluster_specs=None,
+ verbose=False,
+ correction_method=0,
+):
+ """Create a minimal SimulationMetadata for testing."""
+ return SimulationMetadata(
+ target_indices=np.arange(n_targets),
+ n_non_factor_vars=n_targets,
+ correlation_matrix=np.eye(n_targets),
+ var_types=np.zeros(n_targets, dtype=np.int64),
+ var_params=np.zeros(n_targets, dtype=np.float64),
+ factor_specs=[],
+ upload_normal_values=np.zeros((2, 2), dtype=np.float64),
+ upload_data_values=np.zeros((2, 2), dtype=np.float64),
+ effect_sizes=np.array([0.5] * n_targets),
+ correction_method=correction_method,
+ cluster_specs=cluster_specs or {},
+ verbose=verbose,
+ )
+
+
+def _noop_perturbations(corr, types, config, seed):
+ return corr, types
+
+
+class TestAllSimulationsFail:
+ """When all simulations return None, RuntimeError should be raised."""
+
+ def test_all_fail_raises(self):
+ runner = SimulationRunner(n_simulations=5, seed=42)
+ metadata = _make_metadata()
+
+ def failing_sim(*args, **kwargs):
+ return None
+
+ with patch.object(runner, "_single_simulation", return_value=None):
+ with pytest.raises(RuntimeError, match="All simulations failed"):
+ runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+
+
+class TestLMEThresholdExceeded:
+ """LME failure rate exceeding threshold raises RuntimeError."""
+
+ def test_high_failure_rate_raises(self):
+ runner = SimulationRunner(n_simulations=10, seed=42, max_failed_simulations=0.05)
+ metadata = _make_metadata(cluster_specs={"school": {"n_clusters": 5, "cluster_size": 10}})
+
+ call_count = [0]
+
+ def sometimes_fail(*args, **kwargs):
+ call_count[0] += 1
+ if call_count[0] <= 5:
+ return None # 5 out of 10 fail = 50%
+ return (np.array([1, 1, 1]), np.array([1, 1, 1]), False)
+
+ with patch.object(runner, "_single_simulation", side_effect=sometimes_fail):
+ with pytest.raises(RuntimeError, match="Too many failed simulations"):
+ runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+
+
+class TestOLSHighFailureWarns:
+ """OLS high failure rate warns but doesn't raise."""
+
+ def test_ols_warns_above_10_percent(self):
+ runner = SimulationRunner(n_simulations=10, seed=42)
+ metadata = _make_metadata() # No cluster_specs = OLS
+
+ call_count = [0]
+
+ def sometimes_fail(*args, **kwargs):
+ call_count[0] += 1
+ if call_count[0] <= 2:
+ return None # 2 out of 10 fail = 20%
+ return (np.array([1, 1, 1]), np.array([1, 1, 1]))
+
+ with patch.object(runner, "_single_simulation", side_effect=sometimes_fail):
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ result = runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+ assert any("failed" in str(warning.message).lower() for warning in w)
+
+
+class TestWaldFallbackWarning:
+ """Warn if >10% iterations use Wald test."""
+
+ def test_wald_warning_above_threshold(self):
+ runner = SimulationRunner(n_simulations=10, seed=42)
+ metadata = _make_metadata()
+
+ call_count = [0]
+
+ def wald_heavy(*args, **kwargs):
+ call_count[0] += 1
+ # All return wald_flag=True
+ return (np.array([1, 1, 1]), np.array([1, 1, 1]), True)
+
+ with patch.object(runner, "_single_simulation", side_effect=wald_heavy):
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ result = runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+ assert any("Wald test fallback" in str(warning.message) for warning in w)
+ assert result["n_wald_fallbacks"] == 10
+
+
+class TestVerboseDiagnostics:
+ """Verbose mode collects diagnostics and failure reasons."""
+
+ def test_verbose_success_collects_diagnostics(self):
+ runner = SimulationRunner(n_simulations=3, seed=42)
+ metadata = _make_metadata(verbose=True)
+
+ def verbose_result(*args, **kwargs):
+ return {
+ "results": (np.array([1, 1, 1]), np.array([1, 1, 1])),
+ "diagnostics": {"icc_estimated": 0.2},
+ "wald_fallback": False,
+ }
+
+ with patch.object(runner, "_single_simulation", side_effect=verbose_result):
+ result = runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+ assert "diagnostics" in result
+ assert len(result["diagnostics"]) == 3
+
+ def test_verbose_failure_tracking(self):
+ runner = SimulationRunner(n_simulations=5, seed=42)
+ metadata = _make_metadata(verbose=True)
+
+ call_count = [0]
+
+ def mixed_results(*args, **kwargs):
+ call_count[0] += 1
+ if call_count[0] <= 2:
+ return {"failed": True, "failure_reason": "Convergence failed"}
+ return {
+ "results": (np.array([1, 1, 1]), np.array([1, 1, 1])),
+ "diagnostics": {},
+ "wald_fallback": False,
+ }
+
+ with patch.object(runner, "_single_simulation", side_effect=mixed_results):
+ with warnings.catch_warnings():
+ warnings.simplefilter("ignore", UserWarning)
+ result = runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+ assert "failure_reasons" in result
+ assert result["failure_reasons"]["Convergence failed"] == 2
+
+ def test_verbose_none_tracking(self):
+ """None results in verbose mode are tracked as unknown failures."""
+ runner = SimulationRunner(n_simulations=3, seed=42)
+ metadata = _make_metadata(verbose=True)
+
+ call_count = [0]
+
+ def mixed(*args, **kwargs):
+ call_count[0] += 1
+ if call_count[0] == 1:
+ return None
+ return {
+ "results": (np.array([1, 1, 1]), np.array([1, 1, 1])),
+ "diagnostics": {},
+ "wald_fallback": False,
+ }
+
+ with patch.object(runner, "_single_simulation", side_effect=mixed):
+ with warnings.catch_warnings():
+ warnings.simplefilter("ignore", UserWarning)
+ result = runner.run_power_simulations(
+ sample_size=100,
+ metadata=metadata,
+ generate_y_func=MagicMock(),
+ analyze_func=MagicMock(),
+ create_X_extended_func=MagicMock(),
+ apply_perturbations_func=_noop_perturbations,
+ )
+ assert "Unknown (returned None)" in result["failure_reasons"]
+
+
+class TestICCMismatchWarning:
+ """ICC mismatch warning when estimated ICC differs by >50%."""
+
+ def test_large_mismatch_warns(self):
+ metadata = _make_metadata(
+ cluster_specs={"school": {"icc": 0.2, "n_clusters": 20, "cluster_size": 10}},
+ )
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ _warn_icc_mismatch(metadata, mean_estimated_icc=0.05) # 75% deviation
+ assert any("differs from specified" in str(warning.message) for warning in w)
+
+ def test_within_tolerance_no_warning(self):
+ metadata = _make_metadata(
+ cluster_specs={"school": {"icc": 0.2, "n_clusters": 20, "cluster_size": 10}},
+ )
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ _warn_icc_mismatch(metadata, mean_estimated_icc=0.18) # 10% deviation
+ icc_warnings = [x for x in w if "differs from specified" in str(x.message)]
+ assert len(icc_warnings) == 0
+
+ def test_zero_estimated_icc_no_warning(self):
+ metadata = _make_metadata(
+ cluster_specs={"school": {"icc": 0.2, "n_clusters": 20, "cluster_size": 10}},
+ )
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ _warn_icc_mismatch(metadata, mean_estimated_icc=0.0)
+ icc_warnings = [x for x in w if "differs from specified" in str(x.message)]
+ assert len(icc_warnings) == 0
+
+ def test_no_icc_in_spec_no_warning(self):
+ metadata = _make_metadata(
+ cluster_specs={"school": {"icc": None, "n_clusters": 20, "cluster_size": 10}},
+ )
+ with warnings.catch_warnings(record=True) as w:
+ warnings.simplefilter("always")
+ _warn_icc_mismatch(metadata, mean_estimated_icc=0.5)
+ icc_warnings = [x for x in w if "differs from specified" in str(x.message)]
+ assert len(icc_warnings) == 0
diff --git a/tests/unit/test_test_formula_utils.py b/tests/unit/test_test_formula_utils.py
new file mode 100644
index 0000000..f3db882
--- /dev/null
+++ b/tests/unit/test_test_formula_utils.py
@@ -0,0 +1,319 @@
+"""Tests for test_formula parsing utilities."""
+
+from collections import OrderedDict
+from unittest.mock import MagicMock
+
+import numpy as np
+
+
+class TestExtractTestFormulaEffects:
+ """Test _extract_test_formula_effects helper."""
+
+ def _make_registry(
+ self,
+ effect_names,
+ factor_names=None,
+ factor_dummies=None,
+ cluster_effect_names=None,
+ ):
+ """Create a minimal mock registry for testing."""
+ reg = MagicMock()
+ reg.effect_names = effect_names
+ reg.factor_names = factor_names or []
+ reg.cluster_effect_names = cluster_effect_names or []
+
+ # Build _effects dict with correct ordering
+ effects = OrderedDict()
+ for name in effect_names:
+ eff = MagicMock()
+ eff.effect_type = "interaction" if ":" in name else "main"
+ effects[name] = eff
+ reg._effects = effects
+
+ # Factor dummies
+ reg._factor_dummies = factor_dummies or {}
+ return reg
+
+ def test_simple_subset(self):
+ """y ~ x1 + x2 from generation y ~ x1 + x2 + x3."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3"])
+ effects, random_effects = _extract_test_formula_effects("y ~ x1 + x2", registry)
+ assert effects == ["x1", "x2"]
+ assert random_effects == []
+
+ def test_single_variable(self):
+ """y ~ x1 from generation y ~ x1 + x2 + x3."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3"])
+ effects, random_effects = _extract_test_formula_effects("y ~ x1", registry)
+ assert effects == ["x1"]
+
+ def test_with_interaction(self):
+ """y ~ x1 + x2 + x1:x2 from generation y ~ x1 + x2 + x3 + x1:x2."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3", "x1:x2"])
+ effects, _ = _extract_test_formula_effects("y ~ x1 + x2 + x1:x2", registry)
+ assert effects == ["x1", "x2", "x1:x2"]
+
+ def test_interaction_omitted(self):
+ """y ~ x1 + x2 from generation y ~ x1 + x2 + x1:x2."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x1:x2"])
+ effects, _ = _extract_test_formula_effects("y ~ x1 + x2", registry)
+ assert effects == ["x1", "x2"]
+
+ def test_factor_expands_to_dummies(self):
+ """y ~ x1 + gender from generation y ~ x1 + x2 + gender."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(
+ ["x1", "x2", "gender[F]", "gender[Other]"],
+ factor_names=["gender"],
+ factor_dummies={
+ "gender[F]": {"factor_name": "gender", "level": "F"},
+ "gender[Other]": {"factor_name": "gender", "level": "Other"},
+ },
+ )
+ effects, _ = _extract_test_formula_effects("y ~ x1 + gender", registry)
+ assert effects == ["x1", "gender[F]", "gender[Other]"]
+
+ def test_factor_omitted(self):
+ """y ~ x1 from generation y ~ x1 + gender."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(
+ ["x1", "gender[F]", "gender[Other]"],
+ factor_names=["gender"],
+ factor_dummies={
+ "gender[F]": {"factor_name": "gender", "level": "F"},
+ "gender[Other]": {"factor_name": "gender", "level": "Other"},
+ },
+ )
+ effects, _ = _extract_test_formula_effects("y ~ x1", registry)
+ assert effects == ["x1"]
+
+ def test_with_random_effects(self):
+ """y ~ x1 + (1|school) extracts random effects."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2"])
+ effects, random_effects = _extract_test_formula_effects(
+ "y ~ x1 + (1|school)", registry
+ )
+ assert effects == ["x1"]
+ assert len(random_effects) == 1
+ assert random_effects[0]["grouping_var"] == "school"
+
+ def test_star_operator_expands(self):
+ """y ~ x1*x2 expands to x1 + x2 + x1:x2."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3", "x1:x2"])
+ effects, _ = _extract_test_formula_effects("y ~ x1*x2", registry)
+ assert effects == ["x1", "x2", "x1:x2"]
+
+ def test_equals_sign_formula(self):
+ """y = x1 + x2 works same as y ~ x1 + x2."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3"])
+ effects, _ = _extract_test_formula_effects("y = x1 + x2", registry)
+ assert effects == ["x1", "x2"]
+
+ def test_preserves_registry_order(self):
+ """Effects returned in registry order, not formula order."""
+ from mcpower.utils.test_formula_utils import _extract_test_formula_effects
+
+ registry = self._make_registry(["x1", "x2", "x3", "x1:x2"])
+ # Formula lists x2 before x1
+ effects, _ = _extract_test_formula_effects("y ~ x2 + x1", registry)
+ assert effects == ["x1", "x2"] # registry order preserved
+
+
+class TestComputeTestColumnIndices:
+ """Test _compute_test_column_indices helper."""
+
+ def test_subset_two_of_three(self):
+ """Selecting 2 of 3 effects gives correct indices."""
+ from mcpower.utils.test_formula_utils import _compute_test_column_indices
+
+ all_effect_names = ["x1", "x2", "x3"]
+ test_effect_names = ["x1", "x2"]
+ result = _compute_test_column_indices(all_effect_names, test_effect_names)
+ assert list(result) == [0, 1]
+
+ def test_skip_middle(self):
+ """Selecting first and last of 3 effects."""
+ from mcpower.utils.test_formula_utils import _compute_test_column_indices
+
+ all_effect_names = ["x1", "x2", "x3"]
+ test_effect_names = ["x1", "x3"]
+ result = _compute_test_column_indices(all_effect_names, test_effect_names)
+ assert list(result) == [0, 2]
+
+ def test_single_effect(self):
+ """Single effect selected."""
+ from mcpower.utils.test_formula_utils import _compute_test_column_indices
+
+ all_effect_names = ["x1", "x2", "x3"]
+ test_effect_names = ["x2"]
+ result = _compute_test_column_indices(all_effect_names, test_effect_names)
+ assert list(result) == [1]
+
+ def test_all_effects_returns_all_indices(self):
+ """Selecting all effects returns full range."""
+ from mcpower.utils.test_formula_utils import _compute_test_column_indices
+
+ all_effect_names = ["x1", "x2", "x3"]
+ test_effect_names = ["x1", "x2", "x3"]
+ result = _compute_test_column_indices(all_effect_names, test_effect_names)
+ assert list(result) == [0, 1, 2]
+
+ def test_with_interactions(self):
+ """Interaction effects have correct indices."""
+ from mcpower.utils.test_formula_utils import _compute_test_column_indices
+
+ all_effect_names = ["x1", "x2", "x3", "x1:x2"]
+ test_effect_names = ["x1", "x2", "x1:x2"]
+ result = _compute_test_column_indices(all_effect_names, test_effect_names)
+ assert list(result) == [0, 1, 3]
+
+
+class TestRemapTargetIndices:
+ """Test _remap_target_indices helper."""
+
+ def test_simple_remap(self):
+ """Target indices remapped to positions within test columns."""
+ from mcpower.utils.test_formula_utils import _remap_target_indices
+
+ # Original target_indices: [0, 1] (x1, x2 in full model)
+ # test_column_indices: [0, 1] (x1, x2 at positions 0, 1 in X_expanded)
+ # In X_test, x1 is at 0, x2 is at 1 -> remapped: [0, 1]
+ original = np.array([0, 1])
+ test_cols = np.array([0, 1])
+ result = _remap_target_indices(original, test_cols)
+ assert list(result) == [0, 1]
+
+ def test_remap_with_gap(self):
+ """Target indices remapped when test columns skip positions."""
+ from mcpower.utils.test_formula_utils import _remap_target_indices
+
+ # Full model: [x1=0, x2=1, x3=2, x1:x2=3]
+ # Test model: [x1=0, x1:x2=3] -> X_test columns at [0, 3]
+ # target_test="x1" -> original target_indices=[0]
+ # In X_test, x1 is at position 0 -> remapped: [0]
+ original = np.array([0])
+ test_cols = np.array([0, 3])
+ result = _remap_target_indices(original, test_cols)
+ assert list(result) == [0]
+
+ def test_remap_target_at_end(self):
+ """Target index that moves to different position in X_test."""
+ from mcpower.utils.test_formula_utils import _remap_target_indices
+
+ # Full model: [x1=0, x2=1, x3=2]
+ # Test model: [x2=1, x3=2] -> test_column_indices=[1, 2]
+ # target_test="x3" -> original target_indices=[2]
+ # In X_test, x3 is at position 1 (second column) -> remapped: [1]
+ original = np.array([2])
+ test_cols = np.array([1, 2])
+ result = _remap_target_indices(original, test_cols)
+ assert list(result) == [1]
+
+
+class TestPrepareMetadataWithTestFormula:
+ """Integration test: prepare_metadata with test_formula_effects."""
+
+ def test_metadata_has_test_indices_when_provided(self):
+ from mcpower import MCPower
+ from mcpower.core.simulation import prepare_metadata
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ metadata = prepare_metadata(model, ["x1", "x2"], test_formula_effects=["x1", "x2"])
+ assert metadata.test_column_indices is not None
+ assert list(metadata.test_column_indices) == [0, 1]
+ assert metadata.test_target_indices is not None
+ assert metadata.test_effect_count == 2
+
+ def test_metadata_no_test_indices_by_default(self):
+ from mcpower import MCPower
+ from mcpower.core.simulation import prepare_metadata
+
+ model = MCPower("y = x1 + x2")
+ model.set_effects("x1=0.5, x2=0.3")
+ model._apply()
+
+ metadata = prepare_metadata(model, ["x1", "x2"])
+ assert metadata.test_column_indices is None
+
+ def test_remap_skips_targets_not_in_test_formula(self):
+ from mcpower import MCPower
+ from mcpower.core.simulation import prepare_metadata
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ # target_tests = all 3, but test formula only has x1, x2
+ metadata = prepare_metadata(model, ["x1", "x2", "x3"], test_formula_effects=["x1", "x2"])
+ # test_target_indices should only have indices for x1 and x2 in X_test
+ assert len(metadata.test_target_indices) == 2
+
+
+class TestParseTargetTestsWithTestFormula:
+ """Test _parse_target_tests limits 'all' when test_formula is active."""
+
+ def test_all_expands_to_test_formula_effects_only(self):
+ from mcpower import MCPower
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ result = model._parse_target_tests("all", test_formula_effects=["x1", "x2"])
+ assert "x3" not in result
+ assert "x1" in result
+ assert "x2" in result
+ assert "overall" in result
+
+ def test_explicit_target_not_in_test_formula_raises(self):
+ from mcpower import MCPower
+
+ import pytest
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ with pytest.raises(ValueError, match="x3"):
+ model._parse_target_tests("x3", test_formula_effects=["x1", "x2"])
+
+ def test_overall_always_allowed(self):
+ from mcpower import MCPower
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ result = model._parse_target_tests("overall", test_formula_effects=["x1", "x2"])
+ assert "overall" in result
+
+ def test_no_test_formula_uses_all_effects(self):
+ from mcpower import MCPower
+
+ model = MCPower("y = x1 + x2 + x3")
+ model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+ model._apply()
+
+ result = model._parse_target_tests("all")
+ assert "x1" in result
+ assert "x2" in result
+ assert "x3" in result
diff --git a/tests/unit/test_updates.py b/tests/unit/test_updates.py
index ee7a2a0..90b3003 100644
--- a/tests/unit/test_updates.py
+++ b/tests/unit/test_updates.py
@@ -101,14 +101,17 @@ def test_shows_warning_when_newer(self, monkeypatch):
"""Show warning when PyPI version is newer."""
monkeypatch.delenv("_MCPOWER_UPDATE_CHECKED", raising=False)
- # Write a cache file at the path the installed module actually reads from
- from datetime import datetime
-
import mcpower.utils.updates as upd_mod
+
+ # Reset the module-level dedup flag
+ upd_mod._already_checked = False
+
+ # Write a cache file at the path the module actually reads from
+ from datetime import datetime
from pathlib import Path
- cache_path = Path(upd_mod.__file__).parent.parent / ".mcpower_cache.json"
- cache_path.parent.mkdir(exist_ok=True)
+ cache_path = Path.home() / ".cache" / "mcpower" / "update_cache.json"
+ cache_path.parent.mkdir(parents=True, exist_ok=True)
cache_data = {
"last_check": datetime.now().isoformat(),
"latest_version": "99.0.0",
@@ -120,5 +123,6 @@ def test_shows_warning_when_newer(self, monkeypatch):
with pytest.warns(match="NEW MCPower VERSION"):
_check_for_updates("1.0.0")
finally:
- # Clean up the cache file
+ # Clean up the cache file and reset flag
cache_path.unlink(missing_ok=True)
+ upd_mod._already_checked = False
diff --git a/tests/unit/test_upload_data_utils.py b/tests/unit/test_upload_data_utils.py
new file mode 100644
index 0000000..c498b6a
--- /dev/null
+++ b/tests/unit/test_upload_data_utils.py
@@ -0,0 +1,62 @@
+"""Unit tests for mcpower.utils.upload_data_utils — normalize_upload_input."""
+
+import numpy as np
+import pytest
+
+from mcpower.utils.upload_data_utils import normalize_upload_input
+
+
+class TestNormalizeUploadInput:
+ """Tests for normalize_upload_input."""
+
+ def test_dict_input(self):
+ data = {"x1": [1.0, 2.0, 3.0], "x2": [4.0, 5.0, 6.0]}
+ arr, cols = normalize_upload_input(data)
+ assert cols == ["x1", "x2"]
+ assert arr.shape == (3, 2)
+ np.testing.assert_array_equal(arr[:, 0], [1.0, 2.0, 3.0])
+
+ def test_dict_with_strings(self):
+ data = {"group": ["a", "b", "a"], "x1": [1.0, 2.0, 3.0]}
+ arr, cols = normalize_upload_input(data)
+ assert arr.dtype == object
+ assert cols == ["group", "x1"]
+
+ def test_list_input(self):
+ data = [1.0, 2.0, 3.0]
+ arr, cols = normalize_upload_input(data)
+ assert arr.shape == (3, 1)
+ assert cols == ["column_1"]
+
+ def test_1d_array(self):
+ data = np.array([1.0, 2.0, 3.0])
+ arr, cols = normalize_upload_input(data)
+ assert arr.shape == (3, 1)
+ assert cols == ["column_1"]
+
+ def test_2d_array(self):
+ data = np.array([[1.0, 2.0], [3.0, 4.0]])
+ arr, cols = normalize_upload_input(data)
+ assert arr.shape == (2, 2)
+ assert cols == ["column_1", "column_2"]
+
+ def test_2d_array_with_columns(self):
+ data = np.array([[1.0, 2.0], [3.0, 4.0]])
+ arr, cols = normalize_upload_input(data, columns=["a", "b"])
+ assert cols == ["a", "b"]
+
+ def test_dataframe_input(self):
+ pd = pytest.importorskip("pandas")
+ df = pd.DataFrame({"x1": [1.0, 2.0], "x2": [3.0, 4.0]})
+ arr, cols = normalize_upload_input(df)
+ assert cols == ["x1", "x2"]
+ assert arr.shape == (2, 2)
+
+ def test_mismatched_columns_raises(self):
+ data = np.array([[1.0, 2.0], [3.0, 4.0]])
+ with pytest.raises(ValueError, match="columns length"):
+ normalize_upload_input(data, columns=["a", "b", "c"])
+
+ def test_unsupported_type_raises(self):
+ with pytest.raises(TypeError, match="data must be"):
+ normalize_upload_input("not valid data")
diff --git a/tests/unit/test_utils_mixed_models.py b/tests/unit/test_utils_mixed_models.py
new file mode 100644
index 0000000..de4d98a
--- /dev/null
+++ b/tests/unit/test_utils_mixed_models.py
@@ -0,0 +1,27 @@
+"""Tests for mcpower.utils.mixed_models backward-compat re-exports."""
+
+import threading
+
+from mcpower.utils.mixed_models import (
+ _lme_analysis_wrapper,
+ _lme_thread_local,
+ reset_warm_start_cache,
+)
+
+
+class TestReExports:
+ """Verify that the backward-compatibility re-exports resolve correctly."""
+
+ def test_lme_analysis_wrapper_is_callable(self):
+ assert callable(_lme_analysis_wrapper)
+
+ def test_lme_thread_local_is_threading_local(self):
+ assert isinstance(_lme_thread_local, threading.local)
+
+ def test_reset_warm_start_cache_is_callable(self):
+ assert callable(reset_warm_start_cache)
+
+ def test_reset_warm_start_cache_clears_params(self):
+ _lme_thread_local.warm_start_params = "dummy"
+ reset_warm_start_cache()
+ assert _lme_thread_local.warm_start_params is None
 |
|  |
+
## Why MCPower?
Traditional power formulas break down with interactions, correlated predictors, categorical variables, or non-normal data. MCPower simulates instead — generates thousands of datasets like yours, fits your model, and counts how often the effects are detected.
@@ -297,19 +301,20 @@ model.set_effects("group[2]=0.4, group[3]=0.6, covariate=0.3")
# Use "vs" syntax for pairwise comparisons + correction="tukey"
model.find_power(
sample_size=150,
- target_test="group[0] vs group[1], group[0] vs group[2]",
+ target_test="group[1] vs group[2], group[1] vs group[3]",
correction="tukey"
)
```
### Test Individual Assumption Violations
```python
-# Manually add specific violations (without full scenario analysis)
-model.set_heterogeneity(0.2) # Effect sizes vary between people
-model.set_heteroskedasticity(0.15) # Violation of equal variance assumption
+# Add specific violations via custom scenario configs
+model.set_scenario_configs({
+ "my_test": {"heterogeneity": 0.2, "heteroskedasticity": 0.15}
+})
-# Run with your manual settings (no automatic scenario variations)
-model.find_sample_size(target_test="treatment")
+# Run with scenario variations
+model.find_sample_size(target_test="treatment", scenarios=True)
```
### Mixed-Effects Models
@@ -392,7 +397,7 @@ model.find_power(sample_size=200, progress_callback=False)
| **Factor effects** | **`model.set_effects("var[2]=0.5, var[3]=0.7")`** |
| Correlated predictors | `model.set_correlations("corr(var1, var2)=0.4")` |
| Multiple testing correction | Add `correction="FDR"`, `"Holm"`, `"Bonferroni"`, or `"Tukey"`|
-| Post-hoc pairwise comparison | `target_test="group[0] vs group[1]"` with `correction="tukey"` |
+| Post-hoc pairwise comparison | `target_test="group[1] vs group[2]"` with `correction="tukey"` |
| Mixed model (random intercept) | `MCPower("y ~ x + (1\|group)")` + `model.set_cluster(...)` |
| Random slopes | `MCPower("y ~ x + (1+x\|group)")` + `set_cluster(..., random_slopes=["x"], slope_variance=0.1)` |
| Nested random effects | `MCPower("y ~ x + (1\|A/B)")` + two `set_cluster()` calls |
@@ -424,7 +429,7 @@ model.find_power(sample_size=200, progress_callback=False)
- For simple models where all assumptions are clearly met.
- For large analyses with tens of thousands of observations, tiny effects, or very low alpha levels.
-## What Makes Scenarios Different? (Be careful, unvalidated, preliminary scenarios)
+## What Makes Scenarios Different? (Rule-of-thumb scenarios)
**Traditional power analysis assumes perfect conditions.** MCPower's scenarios add realistic "messiness":
@@ -478,8 +483,8 @@ model.set_variable_type("treatment=(factor,3), education=(factor,4)")
# Set effects for specific levels
model.set_effects("treatment[2]=0.5, treatment[3]=0.7, education[2]=0.3")
-# Or set same effect for all levels of a factor
-model.set_effects("treatment=0.5") # Applies to treatment[2] and treatment[3]
+# Each non-reference level needs its own effect
+model.set_effects("treatment[2]=0.5, treatment[3]=0.7")
# Important: Factors cannot be used in correlations
# This will error: model.set_correlations("corr(treatment, education)=0.3")
@@ -508,12 +513,31 @@ model.set_alpha(0.01) # Stricter significance (p < 0.01)
model.set_simulations(10000) # High precision (slower)
```
+### Model Misspecification Testing
+
+Use `test_formula` to generate data with one model but test with a simpler one -- useful for evaluating the power impact of omitting variables:
+
+```python
+# Generate with 3 predictors, test with 2 (omitting x3)
+model = MCPower("y = x1 + x2 + x3")
+model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+model.find_power(100, test_formula="y = x1 + x2")
+
+# Generate with clusters, test without (ignoring clustering)
+model = MCPower("y ~ treatment + (1|school)")
+model.set_cluster("school", ICC=0.2, n_clusters=20)
+model.set_effects("treatment=0.5")
+model.find_power(1000, test_formula="y ~ treatment")
+```
+
+See the [Test Formula Tutorial](https://github.com/pawlenartowicz/MCPower/wiki/Tutorial-Test-Formula) for details.
+
### Formula Syntax
```python
# These are equivalent:
-"y = x1 + x2 + x1*x2" # Assignment style
-"y ~ x1 + x2 + x1*x2" # R-style formula
-"x1 + x2 + x1*x2" # Predictors only
+"y = x1 + x2 + x1:x2" # Assignment style
+"y ~ x1 + x2 + x1:x2" # R-style formula
+"x1 + x2 + x1:x2" # Predictors only
# Interactions:
"x1*x2" # Main effects + interaction (x1 + x2 + x1:x2)
@@ -538,9 +562,8 @@ model.set_correlations("(x1, x2)=0.3, (x1, x3)=-0.2")
## Requirements
- Python ≥ 3.10
-- NumPy, matplotlib, joblib
+- NumPy (≥1.26.0), matplotlib, joblib, tqdm
- pandas (optional, for DataFrame input — install with `pip install mcpower[pandas]`)
-- statsmodels (optional, for mixed-effects models — install with `pip install mcpower[all]`)
## Documentation
@@ -549,11 +572,11 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
- [Quick Start](https://github.com/pawlenartowicz/MCPower/wiki/Quick-Start)
- [Model Specification](https://github.com/pawlenartowicz/MCPower/wiki/Model-Specification)
-- [Variable Types](https://github.com/pawlenartowicz/MCPower/wiki/Variable-Types)
-- [Effect Sizes](https://github.com/pawlenartowicz/MCPower/wiki/Effect-Sizes)
-- [Mixed-Effects Models](https://github.com/pawlenartowicz/MCPower/wiki/Mixed-Effects-Models) (random intercepts, slopes, nested effects)
-- [ANOVA & Post-Hoc Tests](https://github.com/pawlenartowicz/MCPower/wiki/ANOVA-and-Post-Hoc-Tests)
-- [Scenario Analysis](https://github.com/pawlenartowicz/MCPower/wiki/Scenario-Analysis)
+- [Variable Types](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Variable-Types)
+- [Effect Sizes](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Effect-Sizes)
+- [Mixed-Effects Models](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Mixed-Effects) (random intercepts, slopes, nested effects)
+- [ANOVA & Post-Hoc Tests](https://github.com/pawlenartowicz/MCPower/wiki/Tutorial-ANOVA-PostHoc)
+- [Scenario Analysis](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Scenario-Analysis)
- [API Reference](https://github.com/pawlenartowicz/MCPower/wiki/API-Reference)
## Need Help?
@@ -568,8 +591,8 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
- ✅ C++ native backend (pybind11 + Eigen, 3x speedup)
- ✅ Mixed Effects Models (random intercepts, random slopes, nested effects) — [validated against lme4](https://github.com/pawlenartowicz/MCPower/wiki/Concept-LME-Validation)
- 🚧 Logistic Regression (coming soon)
-- 🚧 ANOVA (coming soon)
-- 🚧 Guide about methods, corrections (coming soon)
+- ✅ ANOVA (factor variables as ANOVA, post-hoc pairwise comparisons)
+- ✅ Guide about methods, corrections
- 📋 2 groups comparison with alternative tests
- 📋 Robust regression methods
@@ -578,16 +601,18 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
GPL v3. If you use MCPower in research, please cite:
-Lenartowicz, P. (2025). MCPower: Monte Carlo Power Analysis for Statistical Models. Zenodo. DOI: 10.5281/zenodo.16502734
+Lenartowicz, P. (2025). MCPower: Monte Carlo Power Analysis for Complex Statistical Models (Version
|
+
## Why MCPower?
Traditional power formulas break down with interactions, correlated predictors, categorical variables, or non-normal data. MCPower simulates instead — generates thousands of datasets like yours, fits your model, and counts how often the effects are detected.
@@ -297,19 +301,20 @@ model.set_effects("group[2]=0.4, group[3]=0.6, covariate=0.3")
# Use "vs" syntax for pairwise comparisons + correction="tukey"
model.find_power(
sample_size=150,
- target_test="group[0] vs group[1], group[0] vs group[2]",
+ target_test="group[1] vs group[2], group[1] vs group[3]",
correction="tukey"
)
```
### Test Individual Assumption Violations
```python
-# Manually add specific violations (without full scenario analysis)
-model.set_heterogeneity(0.2) # Effect sizes vary between people
-model.set_heteroskedasticity(0.15) # Violation of equal variance assumption
+# Add specific violations via custom scenario configs
+model.set_scenario_configs({
+ "my_test": {"heterogeneity": 0.2, "heteroskedasticity": 0.15}
+})
-# Run with your manual settings (no automatic scenario variations)
-model.find_sample_size(target_test="treatment")
+# Run with scenario variations
+model.find_sample_size(target_test="treatment", scenarios=True)
```
### Mixed-Effects Models
@@ -392,7 +397,7 @@ model.find_power(sample_size=200, progress_callback=False)
| **Factor effects** | **`model.set_effects("var[2]=0.5, var[3]=0.7")`** |
| Correlated predictors | `model.set_correlations("corr(var1, var2)=0.4")` |
| Multiple testing correction | Add `correction="FDR"`, `"Holm"`, `"Bonferroni"`, or `"Tukey"`|
-| Post-hoc pairwise comparison | `target_test="group[0] vs group[1]"` with `correction="tukey"` |
+| Post-hoc pairwise comparison | `target_test="group[1] vs group[2]"` with `correction="tukey"` |
| Mixed model (random intercept) | `MCPower("y ~ x + (1\|group)")` + `model.set_cluster(...)` |
| Random slopes | `MCPower("y ~ x + (1+x\|group)")` + `set_cluster(..., random_slopes=["x"], slope_variance=0.1)` |
| Nested random effects | `MCPower("y ~ x + (1\|A/B)")` + two `set_cluster()` calls |
@@ -424,7 +429,7 @@ model.find_power(sample_size=200, progress_callback=False)
- For simple models where all assumptions are clearly met.
- For large analyses with tens of thousands of observations, tiny effects, or very low alpha levels.
-## What Makes Scenarios Different? (Be careful, unvalidated, preliminary scenarios)
+## What Makes Scenarios Different? (Rule-of-thumb scenarios)
**Traditional power analysis assumes perfect conditions.** MCPower's scenarios add realistic "messiness":
@@ -478,8 +483,8 @@ model.set_variable_type("treatment=(factor,3), education=(factor,4)")
# Set effects for specific levels
model.set_effects("treatment[2]=0.5, treatment[3]=0.7, education[2]=0.3")
-# Or set same effect for all levels of a factor
-model.set_effects("treatment=0.5") # Applies to treatment[2] and treatment[3]
+# Each non-reference level needs its own effect
+model.set_effects("treatment[2]=0.5, treatment[3]=0.7")
# Important: Factors cannot be used in correlations
# This will error: model.set_correlations("corr(treatment, education)=0.3")
@@ -508,12 +513,31 @@ model.set_alpha(0.01) # Stricter significance (p < 0.01)
model.set_simulations(10000) # High precision (slower)
```
+### Model Misspecification Testing
+
+Use `test_formula` to generate data with one model but test with a simpler one -- useful for evaluating the power impact of omitting variables:
+
+```python
+# Generate with 3 predictors, test with 2 (omitting x3)
+model = MCPower("y = x1 + x2 + x3")
+model.set_effects("x1=0.5, x2=0.3, x3=0.2")
+model.find_power(100, test_formula="y = x1 + x2")
+
+# Generate with clusters, test without (ignoring clustering)
+model = MCPower("y ~ treatment + (1|school)")
+model.set_cluster("school", ICC=0.2, n_clusters=20)
+model.set_effects("treatment=0.5")
+model.find_power(1000, test_formula="y ~ treatment")
+```
+
+See the [Test Formula Tutorial](https://github.com/pawlenartowicz/MCPower/wiki/Tutorial-Test-Formula) for details.
+
### Formula Syntax
```python
# These are equivalent:
-"y = x1 + x2 + x1*x2" # Assignment style
-"y ~ x1 + x2 + x1*x2" # R-style formula
-"x1 + x2 + x1*x2" # Predictors only
+"y = x1 + x2 + x1:x2" # Assignment style
+"y ~ x1 + x2 + x1:x2" # R-style formula
+"x1 + x2 + x1:x2" # Predictors only
# Interactions:
"x1*x2" # Main effects + interaction (x1 + x2 + x1:x2)
@@ -538,9 +562,8 @@ model.set_correlations("(x1, x2)=0.3, (x1, x3)=-0.2")
## Requirements
- Python ≥ 3.10
-- NumPy, matplotlib, joblib
+- NumPy (≥1.26.0), matplotlib, joblib, tqdm
- pandas (optional, for DataFrame input — install with `pip install mcpower[pandas]`)
-- statsmodels (optional, for mixed-effects models — install with `pip install mcpower[all]`)
## Documentation
@@ -549,11 +572,11 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
- [Quick Start](https://github.com/pawlenartowicz/MCPower/wiki/Quick-Start)
- [Model Specification](https://github.com/pawlenartowicz/MCPower/wiki/Model-Specification)
-- [Variable Types](https://github.com/pawlenartowicz/MCPower/wiki/Variable-Types)
-- [Effect Sizes](https://github.com/pawlenartowicz/MCPower/wiki/Effect-Sizes)
-- [Mixed-Effects Models](https://github.com/pawlenartowicz/MCPower/wiki/Mixed-Effects-Models) (random intercepts, slopes, nested effects)
-- [ANOVA & Post-Hoc Tests](https://github.com/pawlenartowicz/MCPower/wiki/ANOVA-and-Post-Hoc-Tests)
-- [Scenario Analysis](https://github.com/pawlenartowicz/MCPower/wiki/Scenario-Analysis)
+- [Variable Types](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Variable-Types)
+- [Effect Sizes](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Effect-Sizes)
+- [Mixed-Effects Models](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Mixed-Effects) (random intercepts, slopes, nested effects)
+- [ANOVA & Post-Hoc Tests](https://github.com/pawlenartowicz/MCPower/wiki/Tutorial-ANOVA-PostHoc)
+- [Scenario Analysis](https://github.com/pawlenartowicz/MCPower/wiki/Concept-Scenario-Analysis)
- [API Reference](https://github.com/pawlenartowicz/MCPower/wiki/API-Reference)
## Need Help?
@@ -568,8 +591,8 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
- ✅ C++ native backend (pybind11 + Eigen, 3x speedup)
- ✅ Mixed Effects Models (random intercepts, random slopes, nested effects) — [validated against lme4](https://github.com/pawlenartowicz/MCPower/wiki/Concept-LME-Validation)
- 🚧 Logistic Regression (coming soon)
-- 🚧 ANOVA (coming soon)
-- 🚧 Guide about methods, corrections (coming soon)
+- ✅ ANOVA (factor variables as ANOVA, post-hoc pairwise comparisons)
+- ✅ Guide about methods, corrections
- 📋 2 groups comparison with alternative tests
- 📋 Robust regression methods
@@ -578,16 +601,18 @@ Full documentation is available on the **[MCPower Wiki](https://github.com/pawle
GPL v3. If you use MCPower in research, please cite:
-Lenartowicz, P. (2025). MCPower: Monte Carlo Power Analysis for Statistical Models. Zenodo. DOI: 10.5281/zenodo.16502734
+Lenartowicz, P. (2025). MCPower: Monte Carlo Power Analysis for Complex Statistical Models (Version