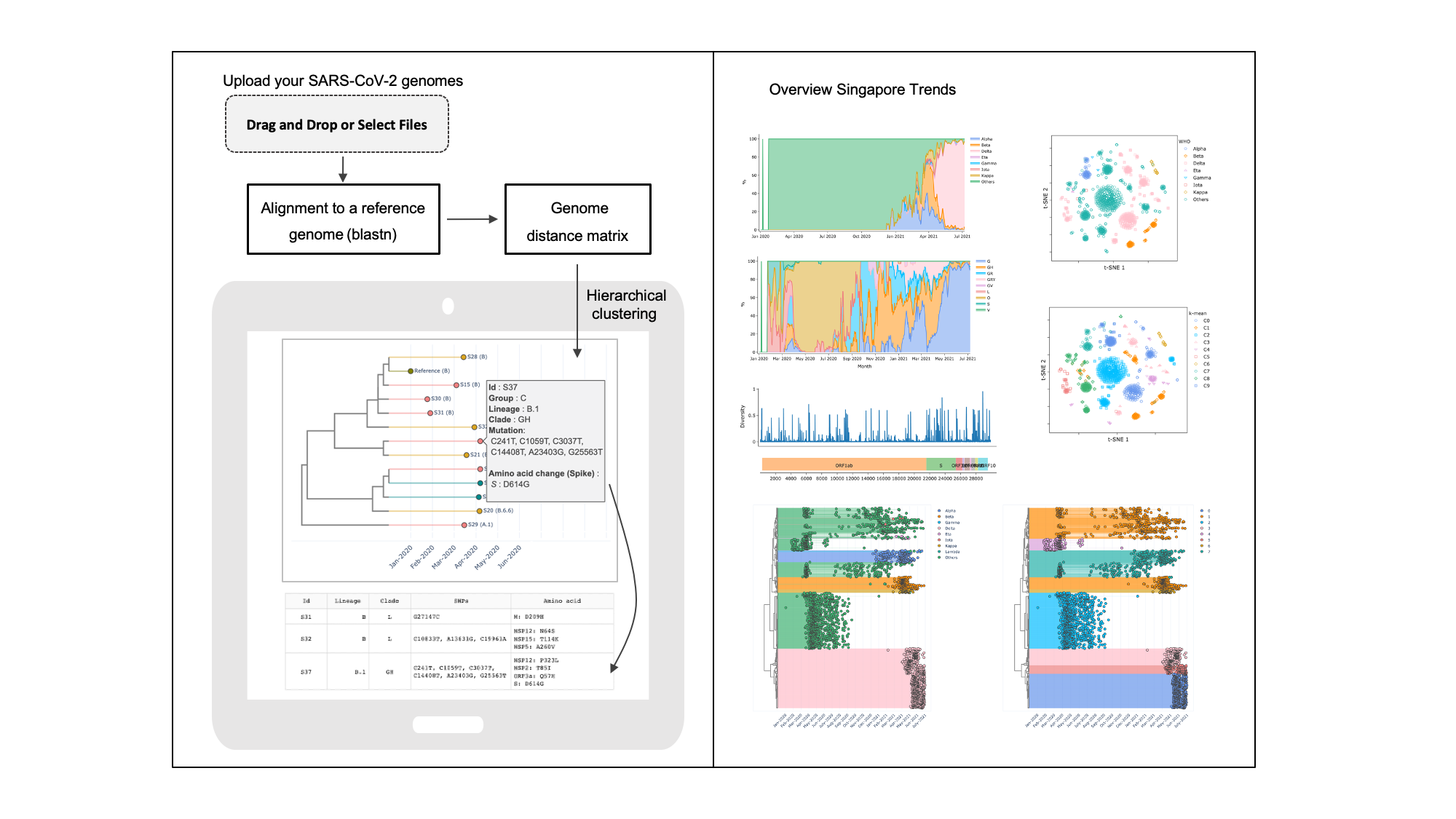
Rapid SARS‑CoV‑2 genome characterisation for outbreak tracking
CalmBelt's demo
Citation: Yingtaweesittikul, Hatairat, Karrie Ko, Nurdyana Abdul Rahman, Shireen Yan Ling Tan, Niranjan Nagarajan, and Chayaporn Suphavilai. "CalmBelt: rapid SARS-CoV-2 genome characterisation for outbreak tracking." Frontiers in Medicine: 2600.
Contact: suphavilaic@gis.a-star.edu.sg and hatairat.y@cmu.ac.th
In this example, we use the lightweight miniconda.
wget https://repo.anaconda.com/miniconda/Miniconda3-latest-Linux-x86_64.sh
bash Miniconda3-latest-Linux-x86_64.sh
conda create -n covid python=3.7A precompiled binary for iqtree2 can be downloaded from https://github.com/iqtree/iqtree2/releases/tag/v2.1.2. The remaining packages can be installed as follow:
conda activate covid
conda install -c bioconda blast
conda install mafft
git clone https://github.com/cov-lineages/pangolin.git
conda env update -n covid --file pangolin/environment.yml
cd pangolin
pip install .
cd ..
git clone https://github.com/neherlab/treetime.git
cd treetime
pip install .
cd ..
pip install -r requirement.txt
conda install -c plotly python-kaleidoToytree will require gs, so you might need to install ghostscript as well sudo apt-get install ghostscript.
This step would require users to prepare input files as follow:
genome_dircontain fasta filesgenome_metadata.tsvcontains region, clade, and lineage informationworld_metadata.tsvcontains continent, clade, and lineage information of samples around the world (can be obtained from GISAID).
python preprecess.py genome_dir reference_fname reference_gene_loci predefined_clade predefined_label preprocess_dir world_metadata_fname country_name subsampling_n_samplesFor example (an example dataset is available at calmbelt/preprocess.zip),
python preprocess.py genome_dir reference.fasta gene.tsv clade.tsv name_by_who.tsv preprocess world_metadata.tsv Singapore -1python covid_app.py preprocess CalmBelt is based on Dash/Flask framework (default port = 8050), and users can access the website at http://localthost:8050. Additional information for deploying CalmBelt on a remote server is avilable upon request.