A proof-of-concept B-Fabric WebApp for invoking bulk transcriptomics processing with NF-Core pipelines, tightly integrated with B-Fabric.
Report Bug
·
Request Feature
Note: This repository was forked from the bfabric-web-app-template, and was built using the bfabric-web-apps Python library.
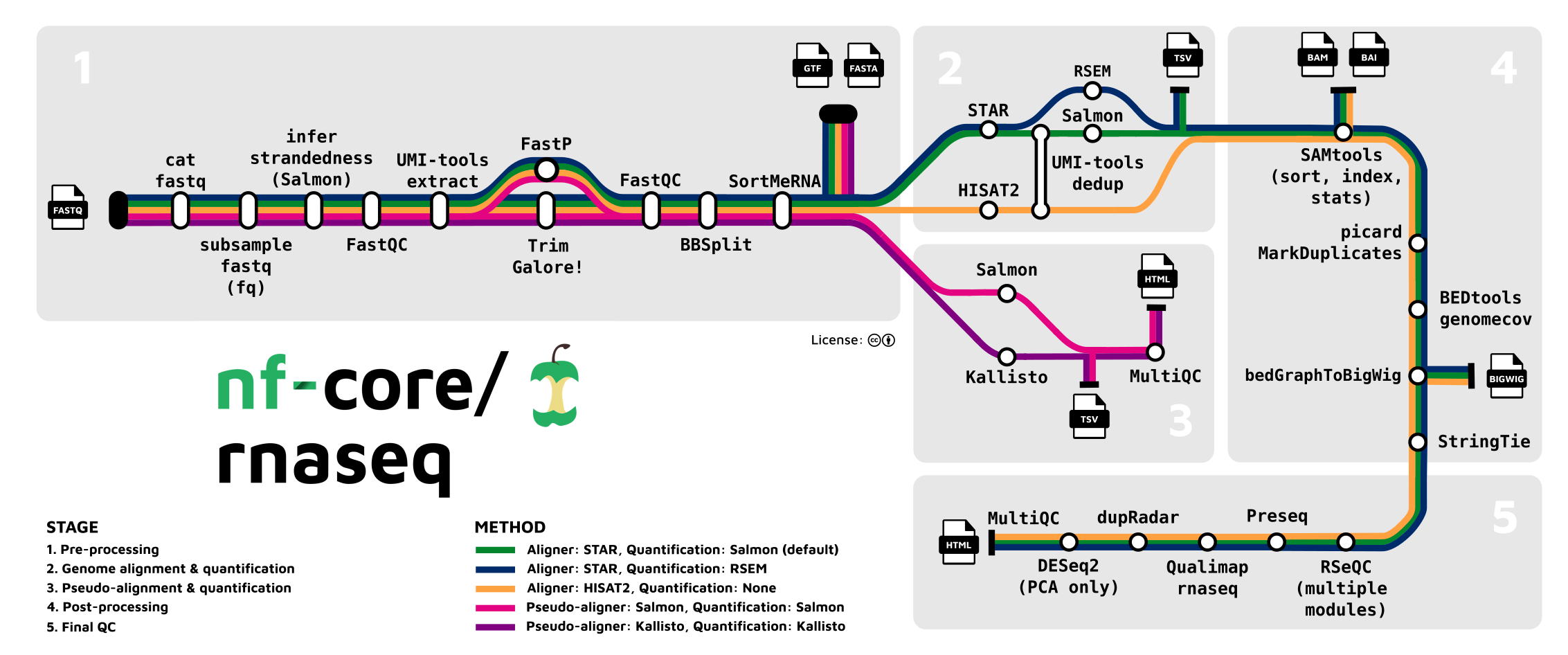
The NF-Core RNA-seq App demonstrates integration between B-Fabric, the Nextflow/NF-Core RNA-seq pipeline, and a Redis-based compute backend. Built using Dash and the bfabric-web-apps module, it enables a structured, interactive interface for RNA-seq data analysis.
- Retrieves and displays sample metadata from B-Fabric.
- Enqueues jobs for NF-Core RNA-seq execution using Redis.
- Links results back to B-Fabric automatically.
- Dash web UI with form-based job submission.
- Automated retrieval of metadata via B-Fabric API.
- Redis-powered job dispatch to remote compute server.
- Integrated output registration in B-Fabric.
Follow these steps to install and run the app locally:
git clone https://github.com/GWCustom/rnaseq.git
cd rnaseqpython3 -m venv venv
source venv/bin/activatepython -m venv venv
venv\Scripts\activateconda create -n rnaseq-app pip
conda activate rnaseq-apppip install -r requirements.txtCreate a file in your home directory at ~/.bfabricpy.yml with your credentials:
GENERAL:
default_config: PRODUCTION
PRODUCTION:
login: your_username
password: your_password
base_url: https://your-bfabric-api-endpointThis file is required to authenticate with the B-Fabric API.
python3 index.pyThen open http://localhost:8050 in your browser.
Distributed under the MIT License. See LICENSE for details.
GWC GmbH - GitHub
Griffin White - LinkedIn
Marc Zuber - LinkedIn