This guide covers everything a data site operator needs to participate in the GRASP glioblastoma survival prediction benchmark on MedPerf.
Create an account at https://www.medperf.org/ and log in.
Organise your data as follows before registering a dataset:
Raw data: one directory per subject, each containing a t1c/ and t2/ subdirectory with DICOM files:
data/
subject_001/
t1c/ (DICOM files)
t2/ (DICOM files)
subject_002/
t1c/
t2/
...
Labels: one plain-text file per subject named <subject_id>.txt (case-sensitive), containing Short-term or Long-term (case-insensitive):
labels/
subject_001.txt
subject_002.txt
...
Clone the MedPerf repository and install the CLI into a dedicated conda environment with Python 3.11:
git clone https://github.com/mlcommons/medperf.git
conda create -n medperf python=3.11
conda activate medperf
pip install ./medperf/cliActivate the environment before running any of the commands below.
Start the MedPerf Web UI:
medperf_webuiYou must obtain and register a certificate with MedPerf before you can be granted access to it.
Navigate to Settings under your profile icon in the top-right corner and scroll to the certificate section at the bottom.
1. Click the button to request a certificate. A URL and a short code are displayed.
2. Open the URL, paste the code into the form, and confirm.
3. Enter your email address and click Continue.
4. Enter the verification code sent to your inbox.
5. A success screen confirms the certificate has been issued.
6. Return to the certificate section in Settings. A new Submit button will appear. Click it and confirm to register your certificate with MedPerf.
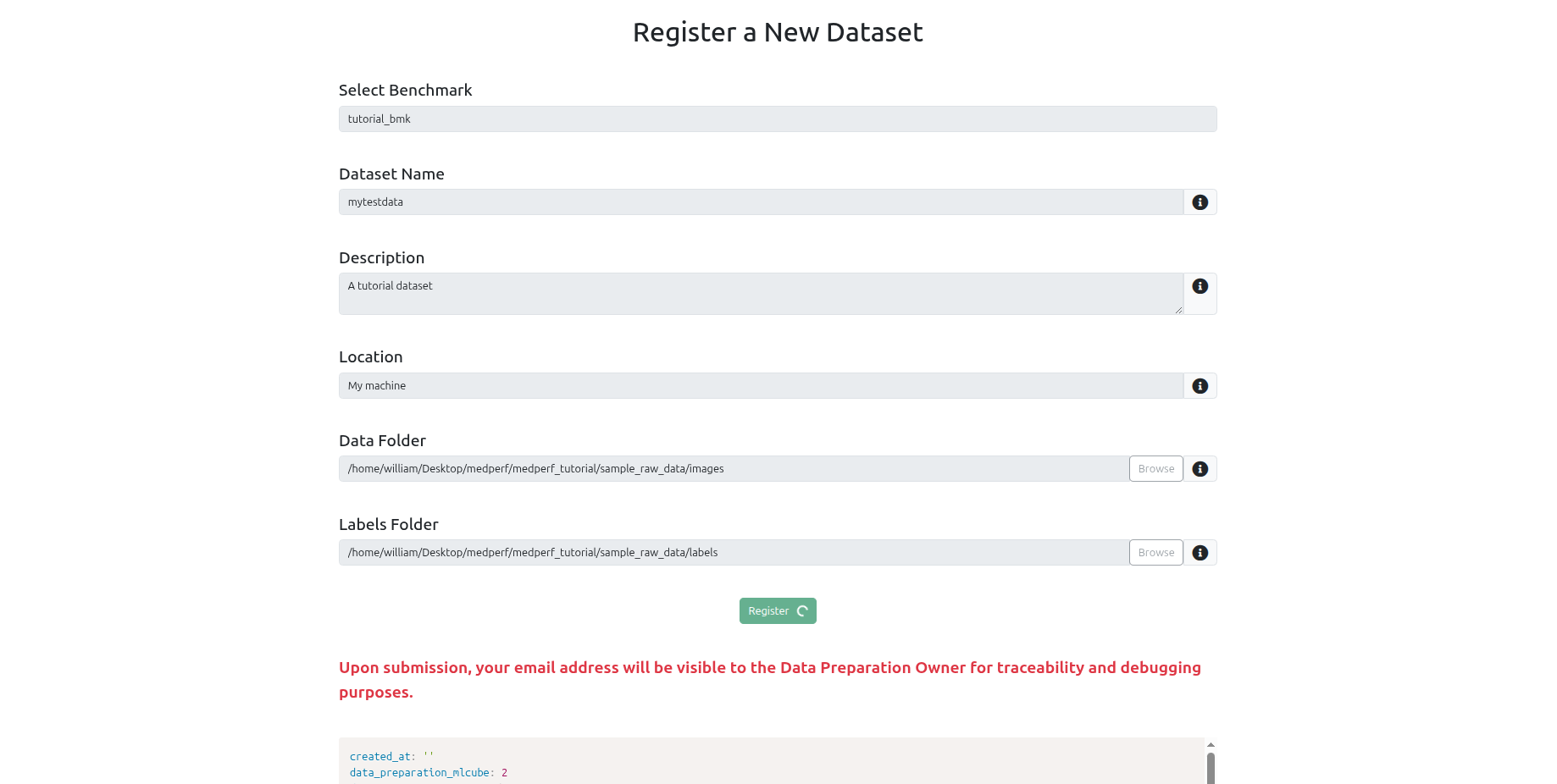
Navigate to the Datasets tab and click Register a new dataset.
Fill in:
- Name and description
- Location (e.g. your institution name)
- Data path: the directory containing your subject folders
- Labels path: the directory containing your
.txtlabel files - Benchmark: select the GRASP benchmark (named
grasp-<version>, e.g.grasp-0.1.0)
Your data never leaves your machine. Only summary statistics are uploaded to the MedPerf server.
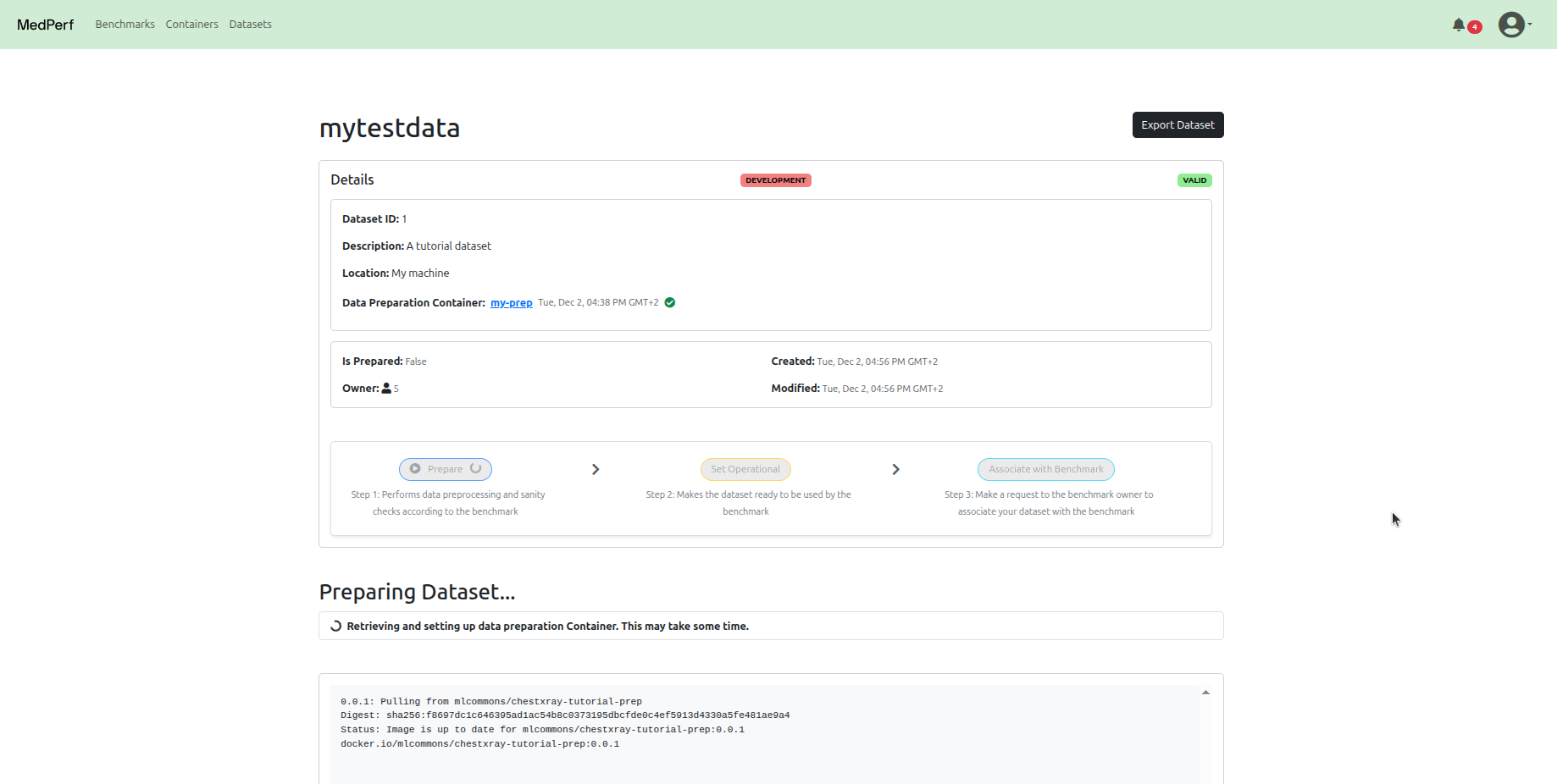
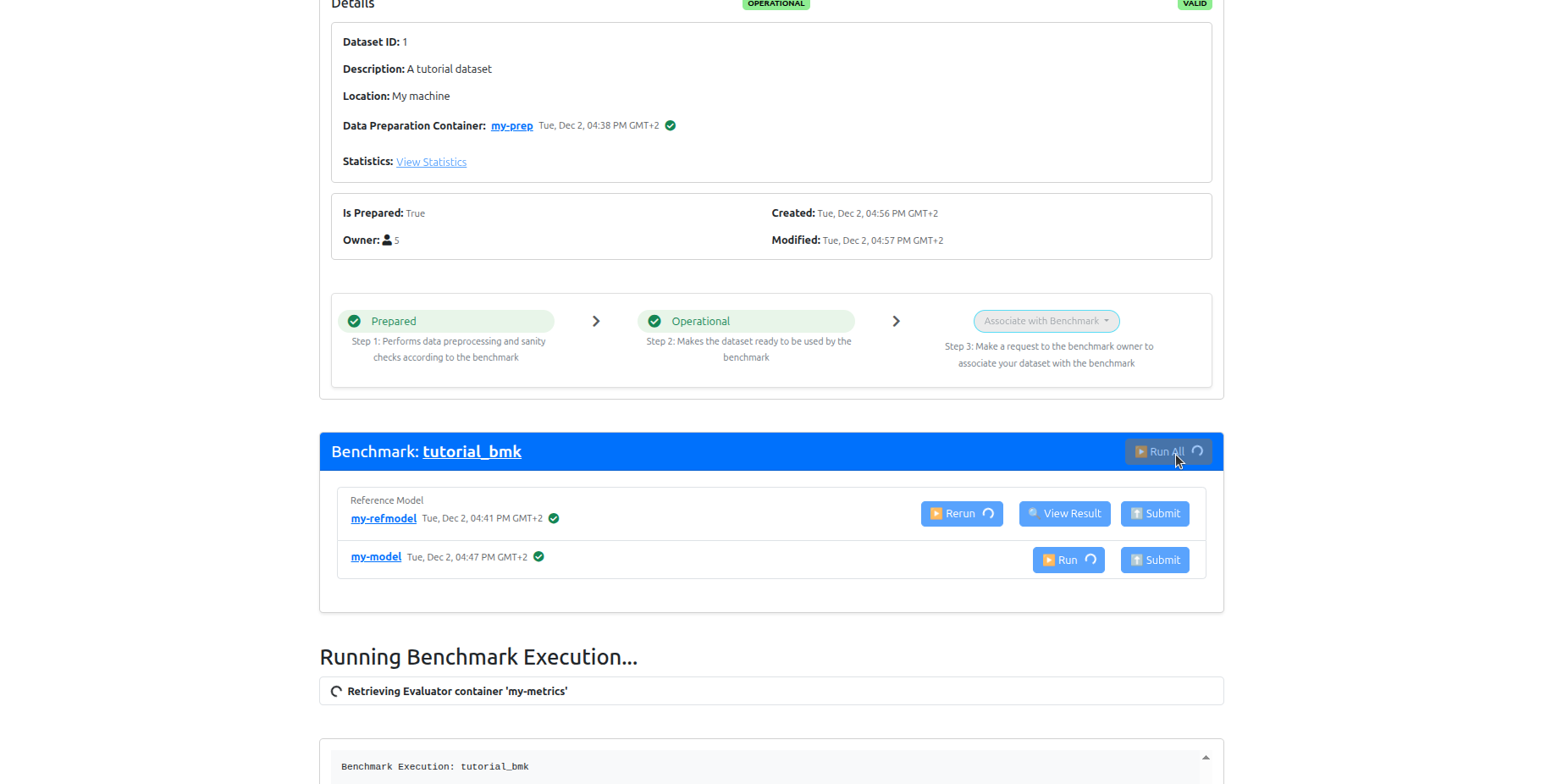
Navigate to your dataset page and click Prepare. The GRASP data preparator will preprocess your DICOM files into the format required for inference.
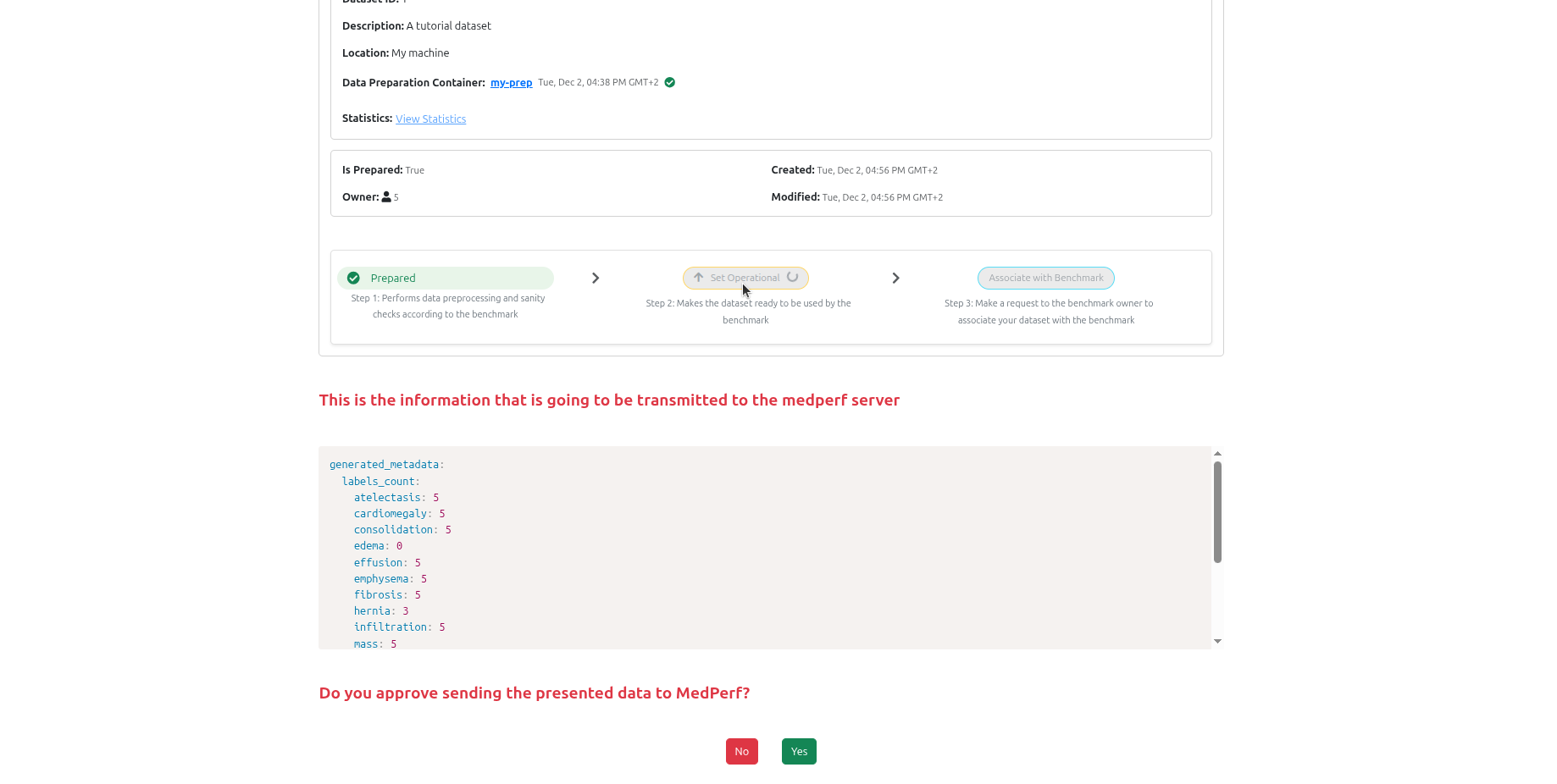
Once preparation completes, click Set Operational to mark the dataset as ready for benchmarking. This uploads anonymised statistics (subject count, label distribution) to the server.
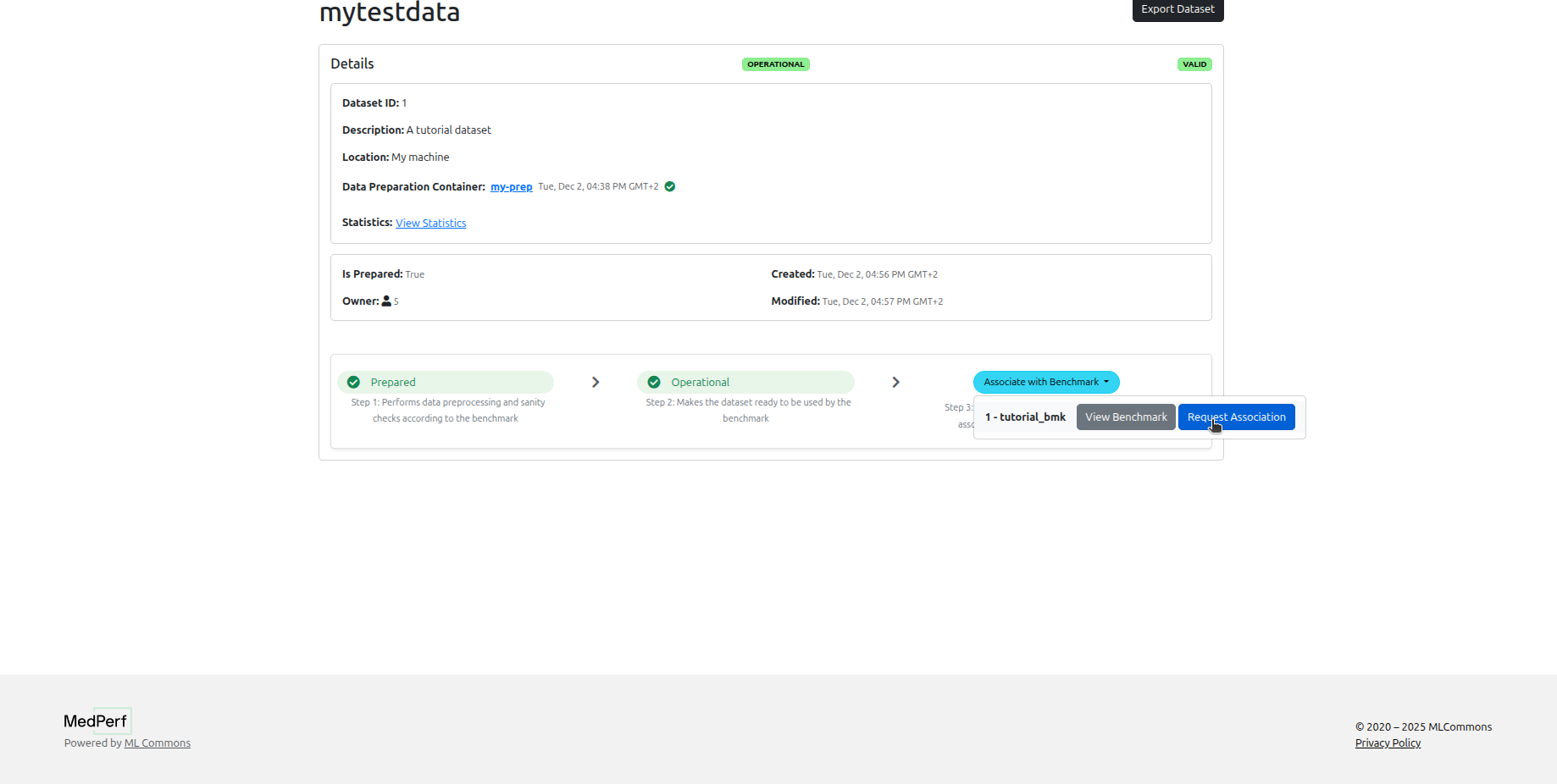
On your dataset page, click Associate with benchmark to submit a participation request to the benchmark owner.
Benchmarking cannot proceed until the benchmark owner approves your request and grants your account access to the encrypted model. You will be notified once this is done.
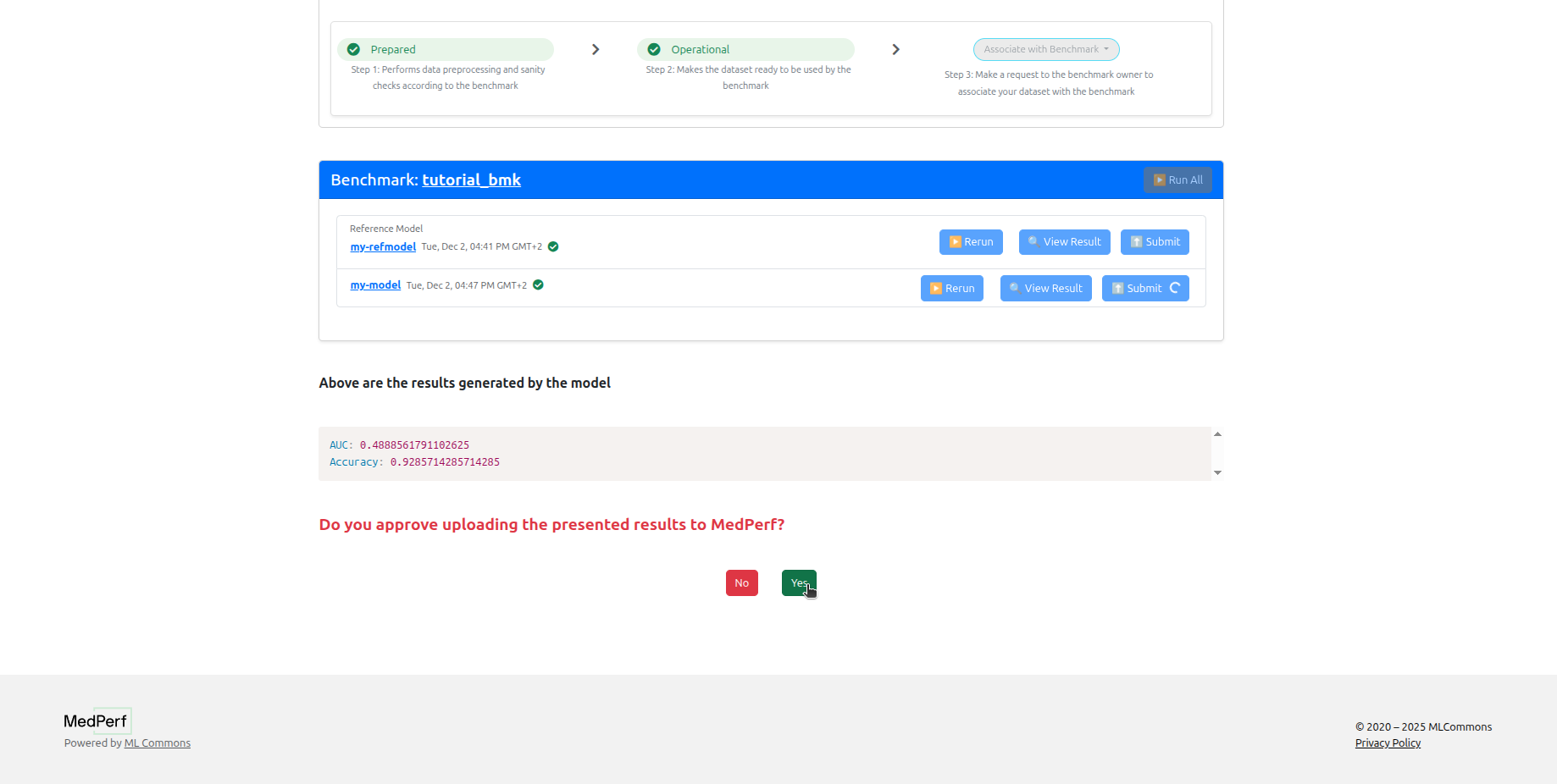
Once your participation request has been approved, return to your dataset page and click Run to execute the model.
Click Submit for each model to send results to the MedPerf server. You can click View Results beforehand to review them locally.
Results will be visible to the benchmark owner once submitted.