-
Notifications
You must be signed in to change notification settings - Fork 1
Design Document
Current methods of querying data from PrediXcan gene expression prediction models are tedious by command line and hard-coded in Python. Our project seeks to ease the process of .db file querying by specifying user inputs, such as R2 to filter genes by or a list of genes to query and output a user-readable .csv file for parsing. Our method is written in Python3, requires the libraries argparse, sqlite3 and pandas and is operated through the command line.
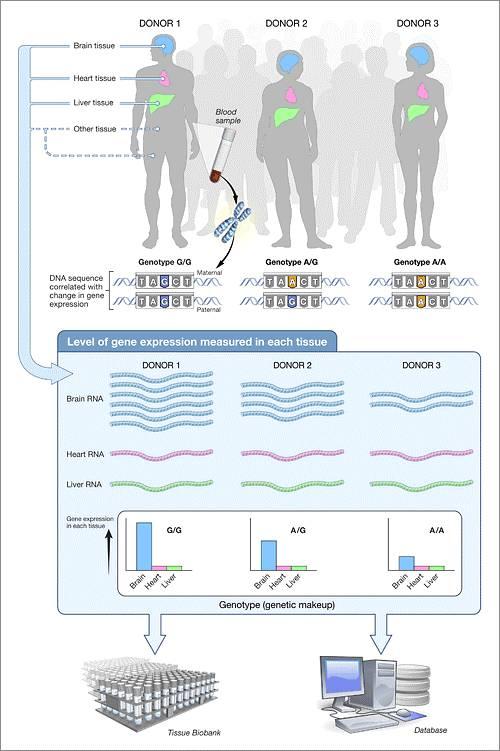
Transcriptome comparisons are an important aspect of genomics. These comparisons are commonly performed to examine differential gene expression across phenotypes, such as within a cell line before and after treatment with a drug or between tumor and normal tissue. Currently, the most popular method to collect and quantify transcriptomic data is with RNA-seq using next-generation sequencing technology. However, RNA-seq data is prohibitively expensive to collect for many labs and RNA expression is largely based on the tissue of origin, and many tissues except for blood are difficult to obtain from study participants. Fortunately, large genomic consortia such as the Genotype-Tissue Expression Project (GTEx) have been generating and publishing genomic and transcriptomic data on various tissues such as adipose, coronary artery, and skeletal muscle in recently deceased donors.
Both genetic and environmental factors contribute to gene expression. While environmental factors are often non-quantifiable and/or non-collectible, genetic data is easily measured and we can use this data to predict gene expression using previously measured genetic-transcriptomic associations. One such method is PrediXcan, a gene-based association method for mapping traits using reference transcriptome data. This method has an input of genotypes and phenotypes, and has outputs of predicted gene expression and predicted gene expression-phenotype associations. PrediXcan predicts gene expression using an additive model of gene expression trait trained in reference transcriptome data such as GTEx that weights SNPs by their association with gene expression. The user simply specifies the model to be used and the genotypes to be input.
The PrediXcan models are stored as .db files, with most models available at predictdb.org. Within these files include specific information on the overall model as well as each gene's predictive model. In the weights table, information of individual SNP's weights for a particular gene are described, along with allelic informatic, while the model summaries tables include information for each gene model, such as the gene's name, number of SNPs in the model, cross-validation R2 average, and the predictive performance R2. Though most users do not delve into the mechanics of the models, users may want to investigate the model behind a single gene or the weights for a single SNP. With current methods querying SQLite3 on the command line individually, this task becomes tedious. The goal of our project is to automate querying of the PrediXcan model .db file for the user, including varying parameters of genes to include and R2 to filter by, and output a .csv file for parsing by the user with the desired data.
- We will streamline user accessibility to data within .db files
- We will have example user input for various queries for demonstrating the abilities of our program
- We will validate our method by finding well-known genotype-gene expression associations
- As a far-reaching goal, we will include optional scripts to plot various data about the model
- As a far-reaching goal, we will make our method available using a web portal
- We will not be rewriting .db files
- We will not be recalculating SNP weights or other statistics
- 3/12
- Querying in command line examples
- Download and become familiar with necessary software
- Assign responsibilities
- Documentation - Angela
- SQL querying - Carlee
- .csv parsing - Shreya
- 3/19
- Querying with Python examples
- "Homework"
- Write Python examples to query multiple genes (APOE, PSRC1) for their SNPs and weights and output a .csv
- Write PowerPoint
- "Homework"
- Querying with Python examples
- 3/26
-
Group presentation
- Background and overview
-
Repo check #1
- Querying with Python examples (finish if not already completed)
- Pseudocode
- Create main methods
-
Group presentation
- 4/2
- Continue main methods
- 4/9
-
Repo check #2
- Main methods
- Finish up main methods
- Make example data
- Begin the app note draft
-
Repo check #2
- 4/16
-
Repo check #3
- Main methods
- Example data
- Debug methods
- Continue and finish app note draft
-
Repo check #3
- 4/23
- App note draft
- Write final group presentation
- Based on app note draft
- 4/30
- Final group presentation
Our method will comprise two main parts and a wrapper around them. The first main part will be querying the .db file and will be primarily authored by Carlee Bettler. We will offer various user parameters customized to the type of analysis required. For example, if a user found a gene of interest they would like to further examine, the user could query all possible information on the gene, such as the predictive performance of the model and all SNPs and their weights included. Alternatively, the user could desire information about the overall model and could query the sample size and the gene names of all genes available. These queries will be parsed from the command line by argparse and passed through commands from the Python sqlite3 package, where the results will be stored as a pandas dataframe.
The second main part of our method will further parse the SQLite3 output with user-specified parameters using methods within pandas. For example, a user may only want to retain information of genes with a cross-validation R2 of > 0.1 and specify this within the arguments so all mismatching data can be discarded. Another user may only want to keep genes on a certain chromosome or find gene names for certain ENSG ids. This method will mainly focus on filtering the output from the first part and will be primarily authored by Shreya Wadhwa. Main documentation, mentorship, and overall maintenance for the project will be performed by Angela Andaleon.