Scientific Computing Tools for Advanced Mathematical Modelling
Supervisor: Prof. Stefano Pagani
Institution: Politecnico di Milano
Academic Year: 2023–2024
This is a mathematical and computational framework to reconstruct, simulate, and infer electrical propagation in the human heart from sparse electro-anatomical data.
Developed within the course Scientific Computing Tools for Advanced Mathematical Modelling, this project uses Gaussian Processes, Neural Networks, Autoencoders, and U-Net architectures to tackle challenges in cardiac activation modeling.
The codebase is structured in three main checkpoints:
Reconstruct the activation time field and conduction velocity from 20 sparse intracardiac recordings.
- Gaussian Process Regression (GPR) on 20-point input data
- Kernel tuning using cross-validated hyperparameter search
- Velocity estimated via gradient inversion + second-level GPR smoothing
Activation Time Field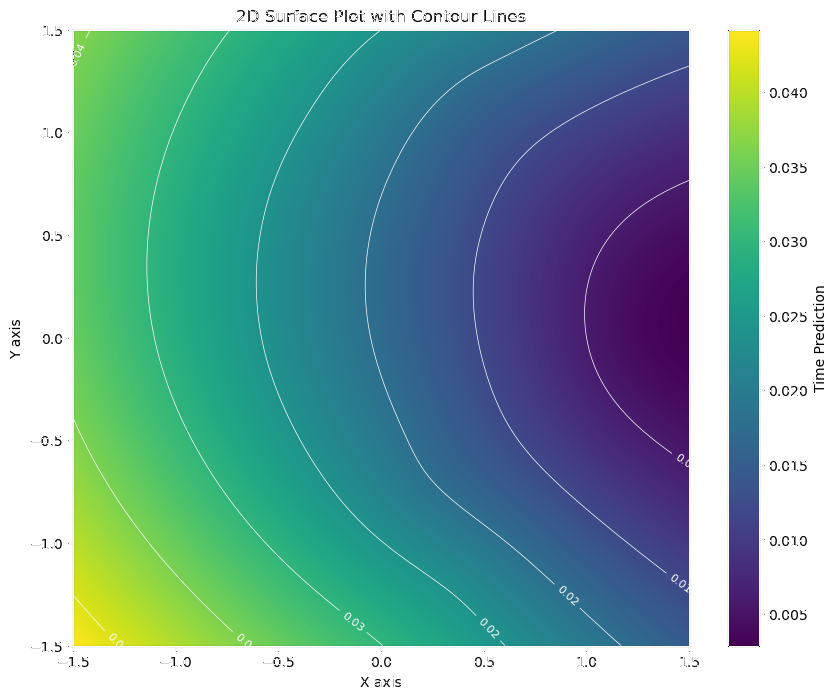
|
Velocity Field Estimation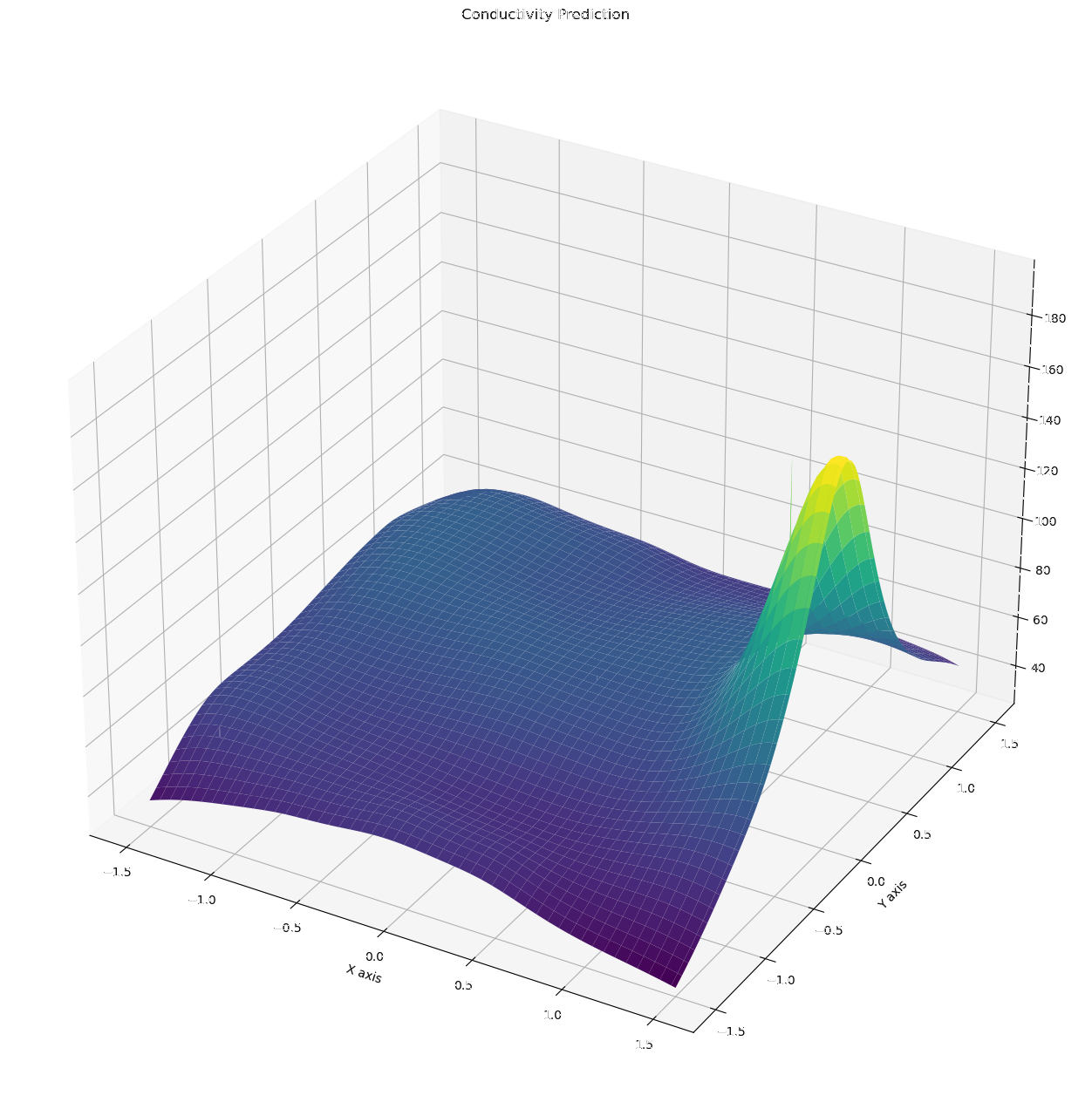
|
Infer three physiological parameters:
- Fiber angle (μ₁ ∈ [−π/10, π/10])
- Anisotropy ratio (μ₂ ∈ [1, 9])
- Activation origin y₀ (μ₃ ∈ [−1.5, 1.5])
- Train a Neural Network surrogate model for the Eikonal solver
anisotropic_FMM - Inputs: (μ₁, μ₂, μ₃, x, y); Output: activation time t
- Use grid search with progressive refinement to optimize parameters by minimizing the squared error functional:
Loss Evolution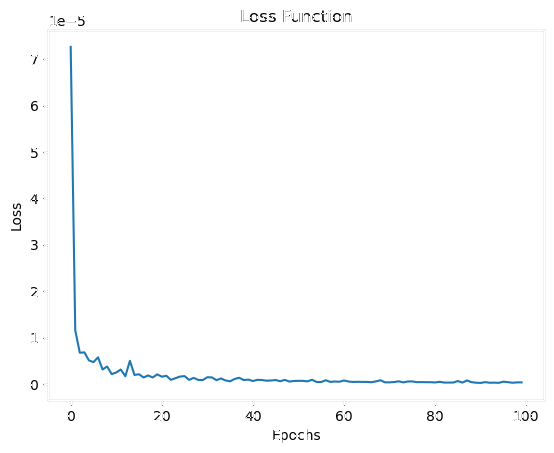
|
NN Prediction vs. Eikonal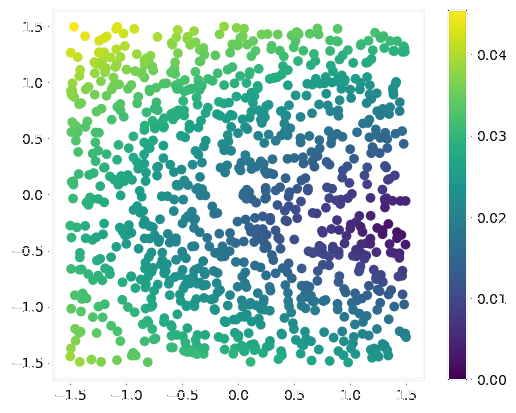
|
Estimate a 2D speed field ( c(x,y) ) from only 20 activation time recordings.
- Compress 151×151 speed fields into 8 latent parameters using an autoencoder
- Optimize latent parameters to minimize mismatch between real and simulated activation time fields
Autoencoder Architecture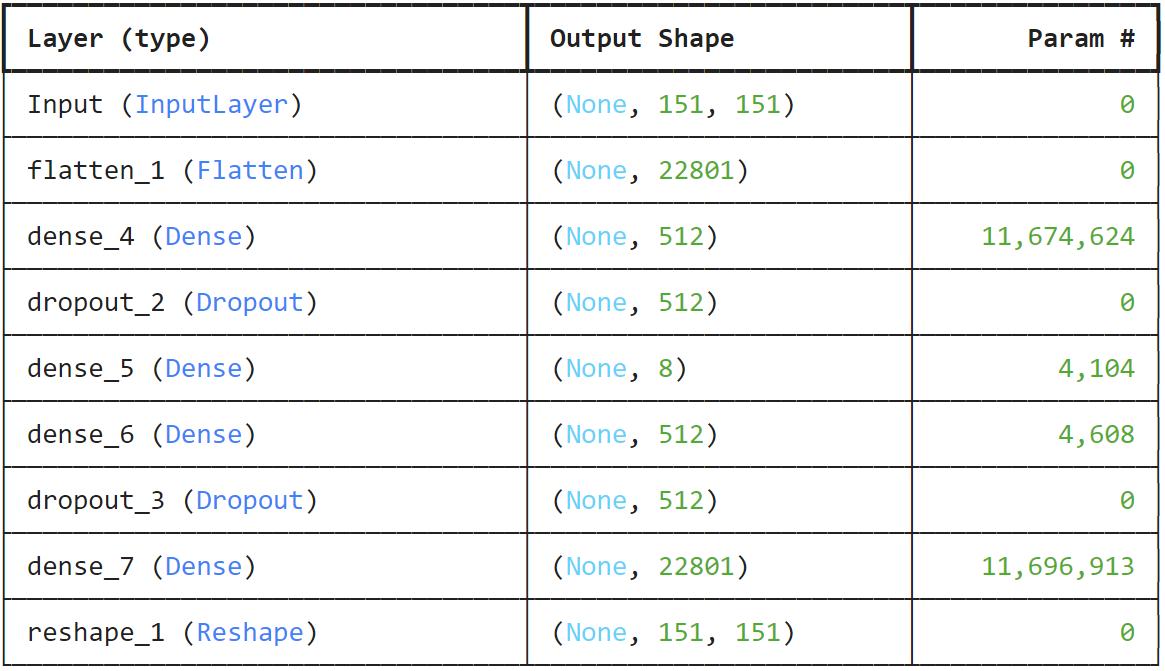
|
Original vs. Decoded Field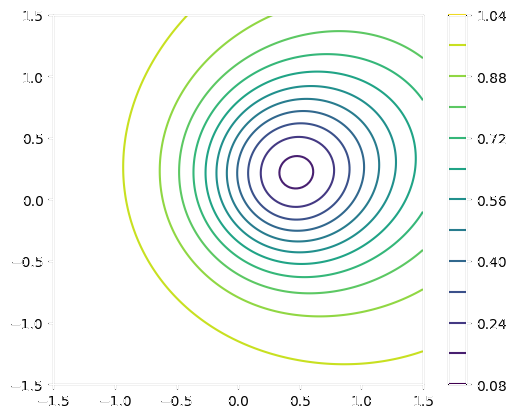
|
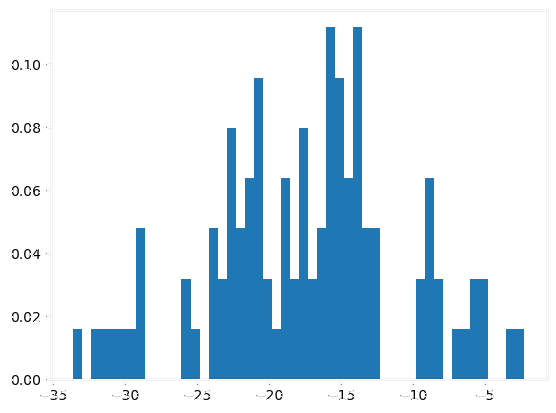 |
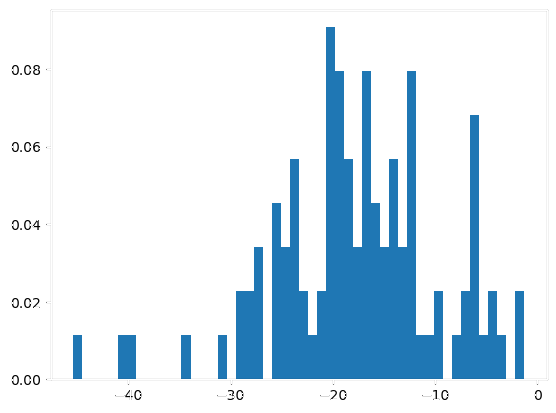 |
 |
 |
 |
 |
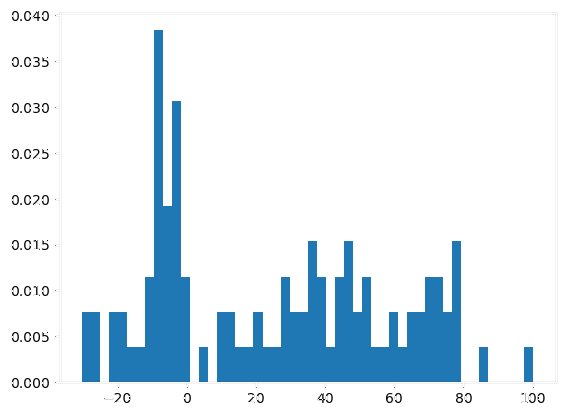 |
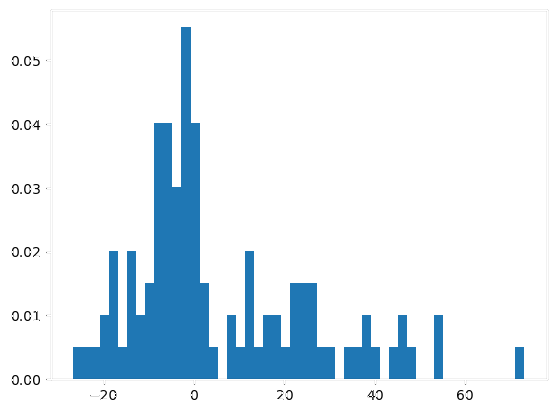 |
An alternative approach was tested using a U-net architecture to directly learn a mapping from activation time images to speed fields.
U-net Architecture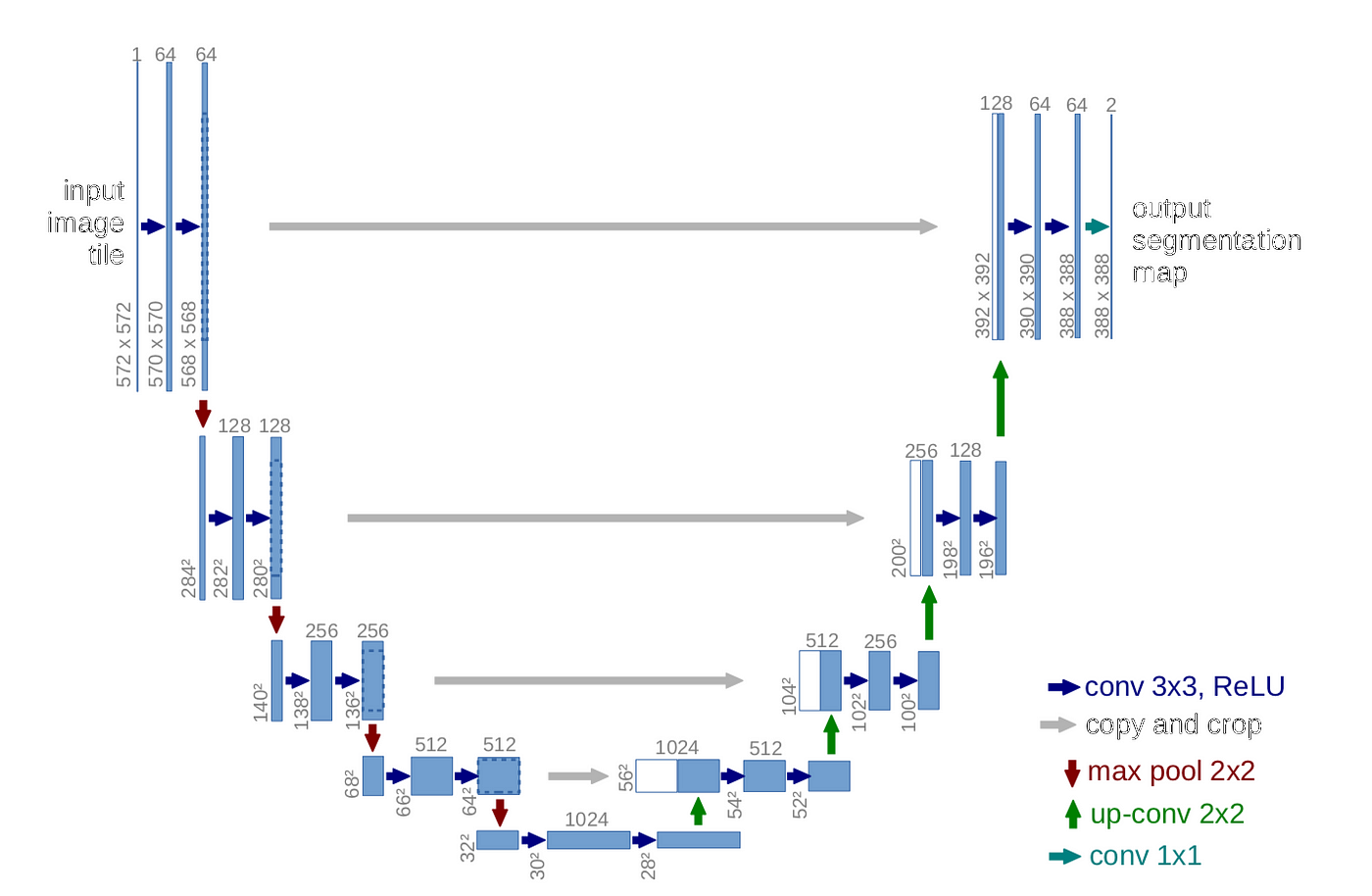
|
Reconstructed Speed Field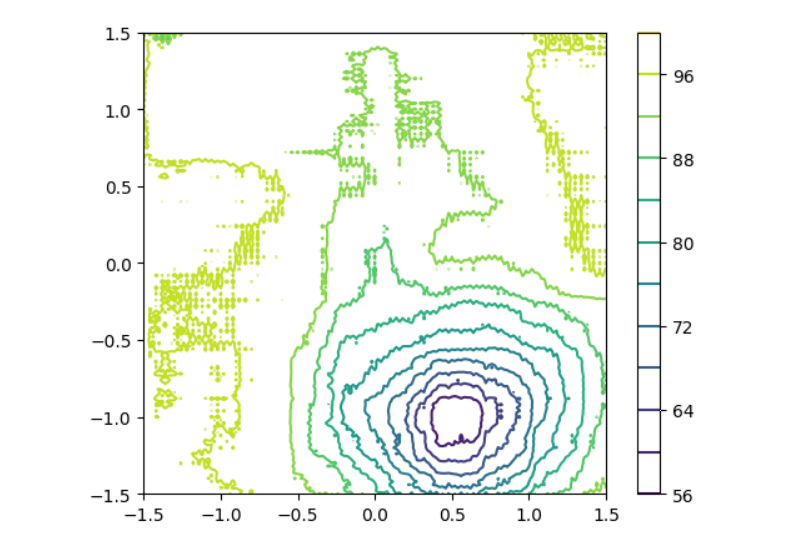
|
Checkpoint_1.ipynb– GPR for activation time & velocity fieldCheckpoint_2.ipynb+functions_CP2.py– NN surrogate + parameter optimizationCheckpoint_3_autoencoder.ipynb– speed field compression and estimationCheckpoint3_U-net.ipynb– U-net speed field regressionmodel_NN_new_training_2.keras– trained model used in CP2
- Activation time prediction accuracy: 0.96 / 1
- U-net reconstruction score: 1.51 / 2
- Autoencoder compression: effective despite minor information loss
- Eikonal solvers and Gaussian Processes: Rasmussen & Williams (2006)
- U-Net: Ronneberger et al. (2015)
- FMM Solver: Sethian (1996)
This project was completed as part of the course Scientific Computing Tools for Advanced Mathematical Modelling held by Prof. Stefano Pagani at Politecnico di Milano.