Merged
Conversation
Contributor
Author
|
Note: commit messages ccf38bb and earlier are from a merge conflict from #54 that I addressed but resulted in my ajlee master branch getting out of sync with greenelab master. So these commits are already merged in greenelab master but still showing in the history. These commits do not reflect the current changes made in this PR |
ben-heil
approved these changes
Jan 27, 2021
 ben-heil
left a comment
ben-heil
left a comment
There was a problem hiding this comment.
Looks good! Sorry for leaving so many comments, it's late (for me) so I'm more confused than usual
pseudomonas_analysis/2_identify_generic_genes_pathways_pao1.ipynb
Outdated
Show resolved
Hide resolved
pseudomonas_analysis/2_identify_generic_genes_pathways_pao1.ipynb
Outdated
Show resolved
Hide resolved
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
Sign up for free
to join this conversation on GitHub.
Already have an account?
Sign in to comment
Add this suggestion to a batch that can be applied as a single commit.This suggestion is invalid because no changes were made to the code.Suggestions cannot be applied while the pull request is closed.Suggestions cannot be applied while viewing a subset of changes.Only one suggestion per line can be applied in a batch.Add this suggestion to a batch that can be applied as a single commit.Applying suggestions on deleted lines is not supported.You must change the existing code in this line in order to create a valid suggestion.Outdated suggestions cannot be applied.This suggestion has been applied or marked resolved.Suggestions cannot be applied from pending reviews.Suggestions cannot be applied on multi-line comments.Suggestions cannot be applied while the pull request is queued to merge.Suggestion cannot be applied right now. Please check back later.
This PR performs a similar validation, previously performed on recount2 data, to pseudomonas data. Here we are comparing the ranking of genes generated by SOPHIE vs those from a manually curated dataset, GAPE.
The following changes were made:
0_prepare_reference_gene_file.ipynbnotebook that processes the curated ANOVA results to get gene rankings. These gene rankings will be what we compare our SOPHIE rankings against2_identify_generic_genes_pathways.ipynbnotebook to compare SOPHIE rankings vs the manually curated ones.0_subset_training_compendium.ipynbto create new training compendium and2_identify_generic_genes_pathways_pao1.ipynbto perform validation analysis on the new training compendium. The code in this notebook is nearly identical to2_identify_generic_genes_pathways.ipynband so doesn't need much review. There were some cutsom edits that needed to be made due to limitations in ponyo. Which I have created an issue for.The main result is here:
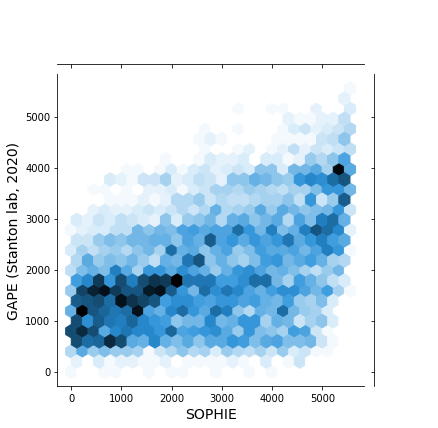
Using only PAO1 samples we get:
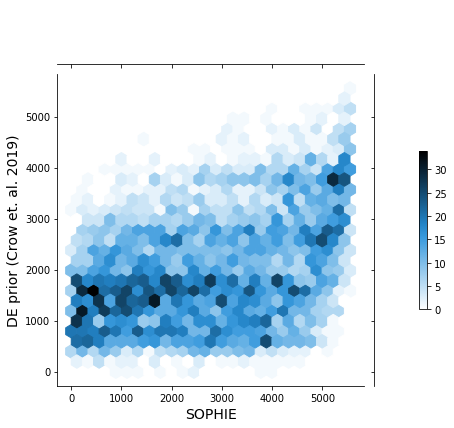
This inconsistency in genes found to be generic by SOPHIE and not by the manually curated set of experiments appears in this analysis using P. aeruginosa data and also human data. Some other hypotheses to test in the future include: