This repository is the official implementation of the Similarity-Hierarchical-Partitioning (SHiP) clustering framework proposed in Ultrametric Cluster Hierarchies: I Want `em All! This framework provides a comprehensive approach to clustering by leveraging similarity trees,
The whole project is implemented in C++ and Python bindings enable the usage within Python.
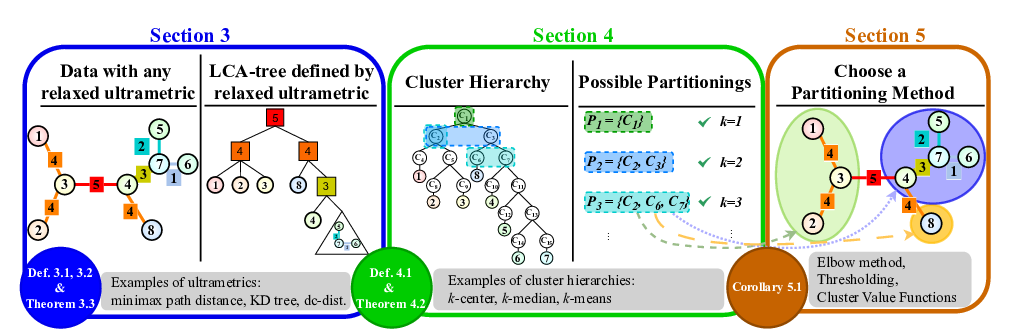
The SHiP framework operates in three main stages:
-
Similarity Tree Construction: A similarity tree is built for the given dataset. This tree represents the relationships and proximities between data points. Note that the default constructed tree corresponds to the
$k$ -center hierarchy (Section 3 in the paper). -
$(k,z)$ -Hierarchy Construction: Using the similarity tree, a$(k,z)$ -hierarchy can be constructed. These hierarchies correlate to common center based clustering methods, as e.g.,$k$ -median or$k$ -means (Section 4). - Partitioning: Finally, the data is partitioned based on the constructed hierarchy and a user-selected partitioning objective function (Section 5).
-
Similarity Trees: The package provides a set of similarity/ultrametric tree implementations:
-
$(k,z)$ -Hierarchies: It supports all possible$(k,z)$ -hierarchies, allowing flexibility in choosing the most suitable hierarchy for a given dataset.-
$z = 0$ →$k$ -center (actually in theory:$z = ∞$ , but in this implementation we use 0 for$∞$ ) -
$z = 1$ →$k$ -median -
$z = 2$ →$k$ -means - ...
-
-
Partitioning Functions: A wide range of partitioning functions are available, enabling users to select the most appropriate function based on their specific needs:
KElbowThresholdThresholdElbowQCoverageQCoverageElbowQStemQStemElbowLcaNoiseElbowLcaNoiseElbowNoTriangleMedianOfElbowsMeanOfElbowsStabilityNormalizedStability
-
Customization: Users can customize the framework by selecting from the available similarity trees,
$(k,z)$ - hierarchies, and partitioning functions.- E.g.,
DCTreewith$k$ -means ($z=2$ )-hierarchy and theElbowpartitioning method.from SHiP import SHiP # Build the `DCTree` ship = SHiP(data=data_points, treeType="DCTree") # Extract the clustering from the $k$-median hierarchy and the `Elbow` partitioning method labels = ship.fit_predict(hierarchy=2, partitioningMethod="Elbow")
- E.g.,
The current stable version can be installed by the following command:
pip install SHiP-framework (coming soon)
Note that a gcc compiler is required for installation. Therefore, in case of an installation error, make sure that:
- Windows: Microsoft C++ Build Tools is installed
- Linux/Mac: Python dev is installed (e.g., by running
apt-get install python-dev- the exact command may differ depending on the linux distribution)
The error messages may look like this:
error: command 'gcc' failed: No such file or directory
Could not build wheels for SHiP-framework, which is required to install pyproject.toml-based projects
Microsoft Visual C++ 14.0 or greater is required. Get it with "Microsoft C++ Build Tools
The current development version can be installed directly from git by executing:
sudo pip install git+https://github.com/pasiweber/SHiP-framework.git
Alternatively, clone the repository, go to the root directory and execute:
pip install .
from SHiP import SHiP
ship = SHiP(data=data, treeType="DCTree")
# or to load a saved tree
ship = SHiP(data=data, treeType="LoadTree", config={"json_tree_filepath": "<file_path>"})
# or additionally specify the tree_type of the loaded tree by adding {"tree_type": "DCTree"}
ship.hierarchy = 0
ship.partitioningMethod = "K"
labels = ship.fit_predict()
# or in one line
labels = ship.fit_predict(hierarchy = 1, partitioningMethod = "Elbow")
# optional: save the current computed tree
json = ship.get_tree().to_json()Our framework achieves the following performance:
| Dataset | DC-0-Stab. | DC-1-MoE | DC-2-Elb. | CT-0-Stab. | CT-1-MoE | CT-2-Elb. |
|
SCAR | Ward | AMD-DBSCAN | DPC |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Boxes | 90.1 | 99.3 | 97.9 | 2.6 | 42.1 ± 4.7 | 24.2 ± 1.6 | 93.5 ± 4.3 | 0.1 ± 0.1 | 95.8 | 63.9 | 25.9 |
| D31 | 79.7 | 42.7 | 82.9 | 46.5 ± 1.8 | 62.0 ± 5.4 | 67.7 ± 3.2 | 92.0 ± 2.7 | 41.7 ± 5.4 | 92.0 | 86.4 | 18.5 |
| airway | 38.0 | 65.9 | 58.8 | 0.8 | 18.2 ± 2.4 | 12.0 ± 1.4 | 39.9 ± 2.0 | -0.9 ± 0.5 | 43.7 | 31.7 | 65.1 |
| lactate | 41.0 | 41.0 | 67.5 | 0.1 | 4.1 ± 0.6 | 1.7 ± 0.2 | 28.6 ± 1.1 | 1.5 ± 1.0 | 27.7 | 71.5 | 0.0 |
| HAR | 30.0 | 46.9 | 52.8 | 14.7 ± 8.8 | 14.2 ± 4.7 | 9.6 ± 2.2 | 46.0 ± 4.5 | 5.5 ± 3.2 | 49.1 | 0.0 | 33.2 |
| letterrec. | 12.1 | 16.6 | 17.9 | 5.8 ± 0.2 | 7.2 ± 0.6 | 6.2 ± 0.3 | 12.9 ± 0.6 | 0.4 ± 0.1 | 14.7 ± 0.9 | 7.9 | 0.0 |
| PenDigits | 66.4 | 73.1 | 75.4 | 8.0 ± 0.8 | 12.0 ± 0.6 | 8.9 ± 0.5 | 55.3 ± 3.2 | 0.9 ± 0.3 | 55.2 | 55.6 | 28.8 ± 1.1 |
| COIL20 | 81.2 | 72.8 | 72.6 | 46.4 ± 4.4 | 46.6 ± 2.1 | 47.7 ± 2.0 | 58.2 ± 2.8 | 33.5 ± 2.0 | 68.6 | 39.2 | 35.9 ± 0.1 |
| COIL100 | 80.1 | 66.8 | 70.0 | 44.6 ± 4.2 | 46.6 ± 1.5 | 50.1 ± 1.2 | 56.1 ± 1.4 | 16.7 ± 0.8 | 61.4 | 14.2 | 0.2 |
| cmu_faces | 60.2 | 56.6 | 66.5 | 8.6 ± 3.1 | 37.1 ± 4.1 | 34.2 ± 2.1 | 53.2 ± 4.7 | 38.5 ± 2.9 | 61.6 | 0.7 | 0.6 |
| OptDigits | 55.3 | 77.0 | 77.0 | 40.9 ± 3.5 | 20.9 ± 2.3 | 18.1 ± 2.4 | 61.3 ± 6.6 | 14.4 ± 4.1 | 74.6 ± 2.4 | 63.2 | 0.0 |
| USPS | 33.7 | 29.3 | 29.3 | 12.0 ± 1.7 | 8.7 ± 1.0 | 11.2 ± 1.5 | 52.3 ± 1.7 | 2.9 ± 0.9 | 63.9 | 0.0 | 21.0 |
| MNIST | 19.7 | 41.7 | 46.0 | 11.1 ± 1.7 | 5.4 ± 0.6 | 5.4 ± 0.6 | 36.9 ± 1.0 | 1.3 ± 0.4 | 52.7 | 0.0 | - |
DC = DCTree,CT = CoverTreeStab. = Stability,MoE = MedianOfElbows,Elb. = Elbow- Competitors: k-means, SCAR, Ward, AMD-DBSCAN, DPC
The project is licensed under the BSD 3-Clause License (see LICENSE.txt).
[1] Connecting the Dots -- Density-Connectivity Distance unifies DBSCAN, k-Center and Spectral Clustering
[2] HST+: An Efficient Index for Embedding Arbitrary Metric Spaces
(Github)
[3] mlpack 4: a fast, header-only C++ machine learning library
(Github)