-
Notifications
You must be signed in to change notification settings - Fork 11
A Tutorial and Usage
liubingdong edited this page Nov 9, 2021
·
25 revisions
if(!require("devtools")) install.packages("devtools")
library(devtools)
install_github("xielab2017/EasyMicroPlot",subdir='Version_0.5.0')
# When something is wrong, please try this below
install_github("xielab2017/EasyMicroPlot",subdir='Version_0.5.0',upgrade = 'never')
## Analysis and results are generated in the output folder
EMP_MICRO(EMP$micro,EMP$mapping)
## The mapping and microbial relative abundance files have already been deposited in the working directory.
## Users only have to activate easy mode to perform a complete workflow with default parameters and generate results in the working-space.
## Demo data at this repository: Tutorial data/Easy_mode_data.7z
EMP_MICRO()
.Result/
├── RFCV_result
│ ├── Imprortance
│ ├── model
│ └── taxonomy
├── alpha_result
│ ├── Post-Hoc
│ ├── data
│ ├── html
│ └── pic
├── beta_result
│ ├── bray_0.001_0.7
│ └── jaccard_0.001_0.7
├── cooc_result
│ └── pic
├── core_data
│ ├── species_0.001_70%.txt
│ └── species_0.001_70%_info.txt
└── structure_result
├── pic
├── taxonomy
└── top_abundance
## Data preparation
meta_data <- EMP$meta
meta_data <- na.omit(meta_data)
meta_data <- meta_data[meta_data$Diarrhea=='n'&meta_data$Astriction=='n'&meta_data$Antibiotics=='n'&meta_data$Synbiotics=='n',]
## Identify the categorical and continuous variables
col_str <- 'MetS'
col_num <- colnames(meta_data)[!colnames(meta_data)%in%c('SampleID','MetS','Diarrhea','Astriction','Antibiotics','Synbiotics')]
## For regroup of meta data
re <- EMP_META(data = meta_data,estimate_group = 'MetS',missing_plot = F,new_col_id = 'Group',col_str = col_str,
col_num = col_num,keep_col = 'SampleID',regroup = T,clust_min = 2,clust_max = 9)
## For table generation
re$summary$summary_table
## For detailed summary of meta data
re$summary$summary_info
## Show cluster details
re$regroup$regroup_info
## Show regroup results
new_group <- re$regroup$regroup_data
## Prepare the bacteria taxa data
core_data <- data_filter(data = EMP$micro,design = EMP$mapping,
min_relative = 0.001,min_ratio = 0.7)
sp <- core_data$filter_data$species
## Analysis and plot
## When ** cor_output = T **, cooc picture is generated in working directory.
cor_re <- EMP_COR(data = sp,meta=meta,cor_output = T,method = 'spearman')
## Load data
library(EasyMicroPlot)
data(EMP)
## For summary of meta data
meta <- meta_summary(EMP$meta,estimate_group = 'MetS',keep_col = 'SampleID')

## For table
meta$summary_table

## For a detailed summary of meta data
meta$summary_info
Aginomoto Animal_oil Plant_oil Salt Sauce
Min. : 0.00 Min. : 0.00 Min. : 0 Min. : 0.0 Min. : 0.00
1st Qu.: 0.00 1st Qu.: 0.00 1st Qu.: 680 1st Qu.: 67.0 1st Qu.: 0.00
Median : 0.00 Median : 0.00 Median :1000 Median :100.0 Median : 0.00
Mean : 18.89 Mean : 24.92 Mean :1209 Mean :115.2 Mean : 25.42
3rd Qu.: 25.00 3rd Qu.: 0.00 3rd Qu.:1500 3rd Qu.:133.0 3rd Qu.: 25.00
Max. :200.00 Max. :1250.00 Max. :5333 Max. :400.0 Max. :500.00
NA's :17 NA's :14 NA's :23 NA's :28 NA's :11
Soy_sauce Sugar Fruit_juice Fruits Grains
Min. : 0.0 Min. : 0.0 Min. : 0 Min. : 0 Min. : 0
1st Qu.: 83.0 1st Qu.: 0.0 1st Qu.: 0 1st Qu.: 27124 1st Qu.: 51025
Median : 150.0 Median : 10.5 Median : 0 Median : 54750 Median : 73000
Mean : 217.5 Mean : 40.8 Mean : 7030 Mean : 70810 Mean : 83152
3rd Qu.: 250.0 3rd Qu.: 50.0 3rd Qu.: 4350 3rd Qu.: 91250 3rd Qu.:109226
Max. :6667.0 Max. :750.0 Max. :182500 Max. :547500 Max. :376500
NA's :21 NA's :20
Red_wine Rice_wine H_alcohol L_alcohol Livestock
Min. : 0 Min. : 0.0 Min. : 0 Min. : 0.0 Min. : 0
1st Qu.: 0 1st Qu.: 0.0 1st Qu.: 0 1st Qu.: 0.0 1st Qu.: 10950
Median : 0 Median : 0.0 Median : 0 Median : 0.0 Median : 23860
Mean : 2319 Mean : 288.3 Mean : 1105 Mean : 221.9 Mean : 33627
3rd Qu.: 900 3rd Qu.: 0.0 3rd Qu.: 0 3rd Qu.: 0.0 3rd Qu.: 42675
Max. :234000 Max. :54750.0 Max. :54750 Max. :18250.0 Max. :182500
Vegetables MetS Diarrhea Astriction Antibiotics Synbiotics
Min. : 0 Cases :154 n :573 n :561 n :560 n :531
1st Qu.: 73000 Control:464 y : 41 y : 53 y : 52 y : 80
Median :109500 NA's: 4 NA's: 4 NA's: 6 NA's: 7
Mean :127451
3rd Qu.:182500
Max. :821250
## Load data
library(EasyMicroPlot)
data(EMP)
## Data preparation
meta_data <- EMP$meta
meta_data <- na.omit(meta_data)
meta_data <- meta_data[meta_data$Diarrhea=='n'&meta_data$Astriction=='n'&meta_data$Antibiotics=='n'&meta_data$Synbiotics=='n',]
## Identify the categorical and continuous variables
col_str <- 'MetS'
col_num <- colnames(meta_data)[!colnames(meta_data)%in%c('SampleID','MetS','Diarrhea','Astriction','Antibiotics','Synbiotics')]
## For regroup of meta data
re <- meta_regroup(data = meta_data,new_col_id = 'Group',col_str = col_str,
col_num = col_num,keep_col = 'SampleID',regroup = T,clust_min = 2,clust_max = 9)
***********************************************************************
*** : The best number of clusters for Continuous variable data is 2
*** : The best number of clusters for Categorical variable data is 2
***** Conclusion *****
***: According to the majority rule, the best number of clusters is 4
***********************************************************************
## Show cluster details
re$regroup_info
$data_num_clust_best
[1] "2"
$data_str_clust_best
[1] 2
$data_clust_best
[1] 4
$data_num_clust_plot
## Show regroup result
new_group <- re$regroup_data
## Load data
library(EasyMicroPlot)
data(EMP)
## Result
core_data <- data_filter(data = EMP$micro,design = EMP$mapping,min_relative = 0.001,min_ratio = 0.7)
## Load data
library(EasyMicroPlot)
data(EMP)
## For Student-Test in comparation
alpha_re <- alpha_plot(data = EMP$micro,design = EMP$mapping,min_relative = 0.001,min_ratio = 0.7,
method = 'ttest')
## For one-way ANOVA and Post Hoc test
alpha_re <- alpha_plot(data = EMP$micro,design = EMP$mapping,min_relative = 0.001,min_ratio = 0.7,
method = 'LSD')
## Support ggplot theme parameters
library(ggplot2)
newtheme=newtheme_slope=theme(axis.text.x =element_text(angle = 90, hjust = -1,size = 15),
axis.title=element_text(size=15,face="bold",colour = 'black'))
alpha_re=alpha_plot(data = EMP$micro,design = EMP$mapping,min_relative = 0.001,min_ratio = 0.7,
method = 'ttest',mytheme = newtheme)
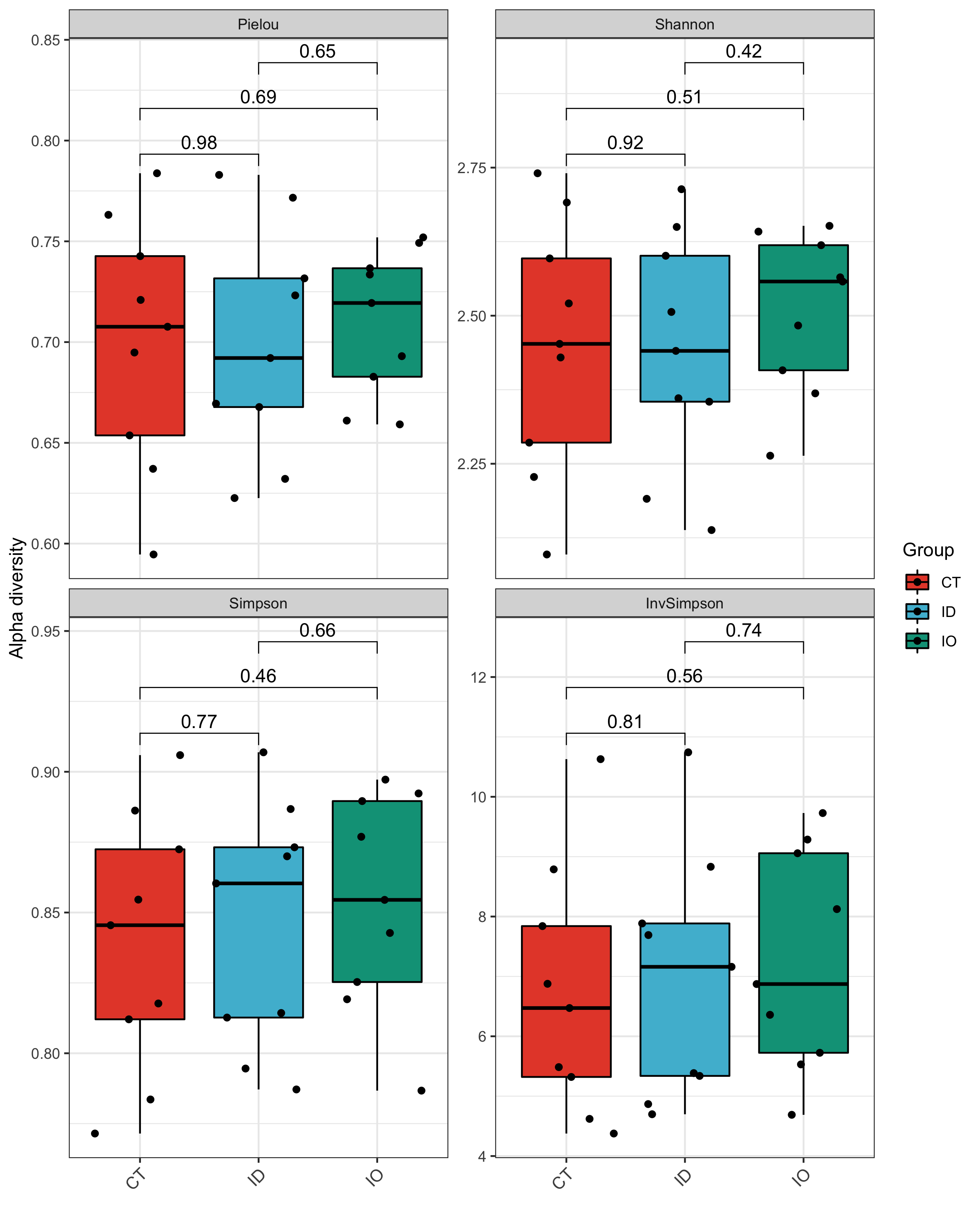
## Plot
alpha_re$plot$species$pic$Total
## Interactive visualization
alpha_re$plot$species$html$Total

## Load data
library(EasyMicroPlot)
data(EMP)
## Result
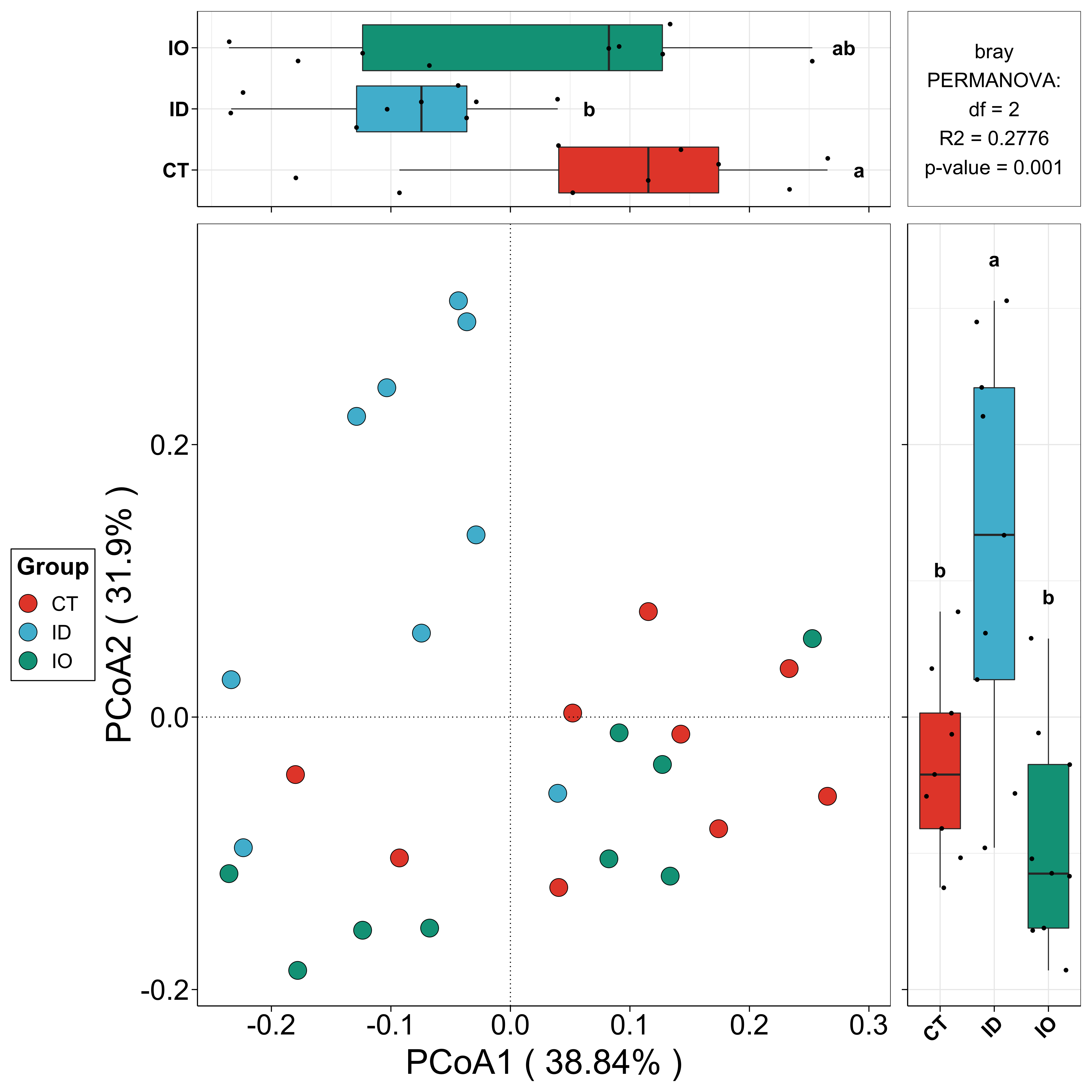
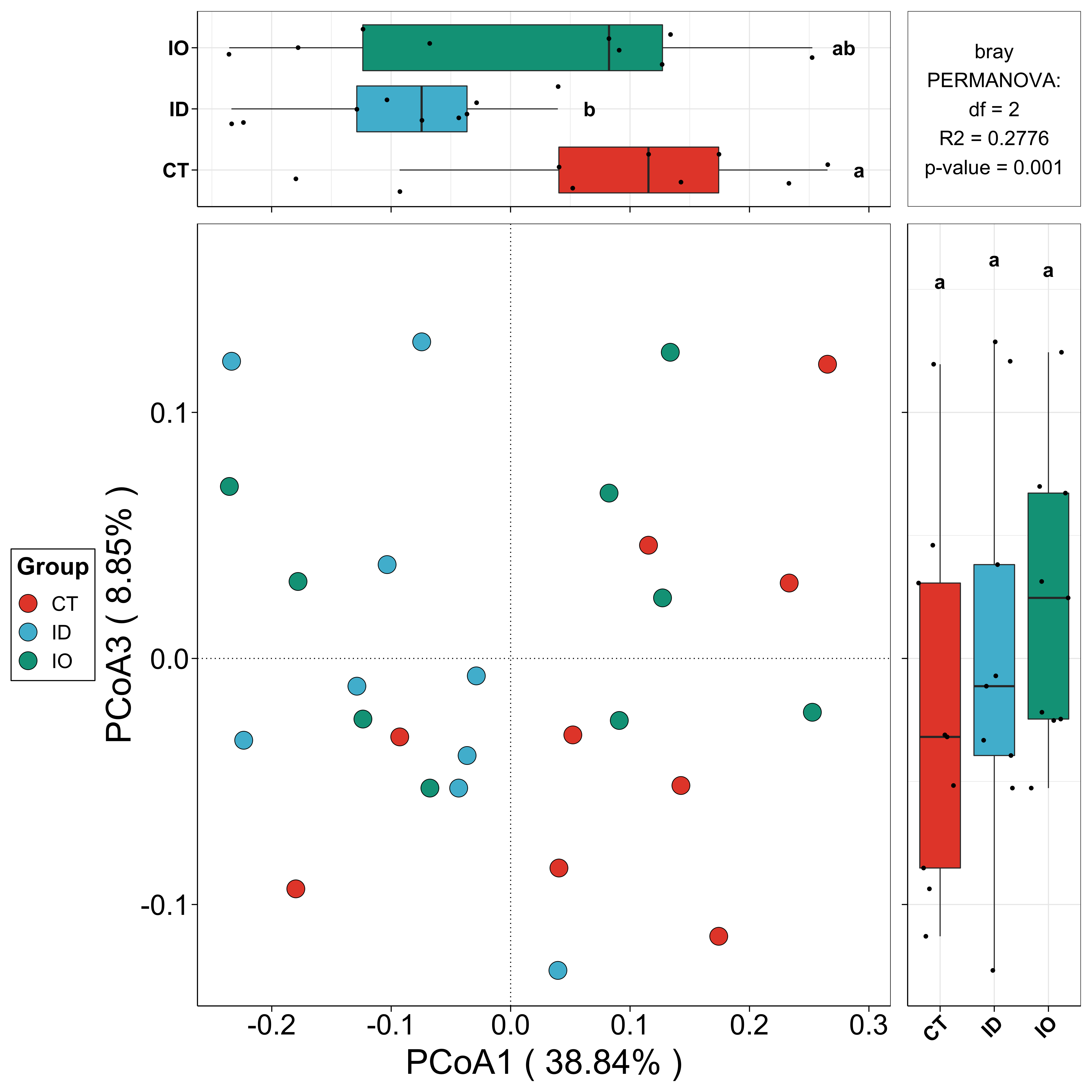
beta_re <- beta_plot(data = EMP$micro,design = EMP$mapping,min_relative = 0.001,min_ratio = 0.7,distance = 'bray')
## Plot
beta_re$plot$species$pic$p12
## Interactive visualization
beta_re$plot$species$html$p12

## Load data
data(EMP)
## Result
## When ** cooc_output = T **, cooc picture is generated in working directory.
cooc_re <- cooc_plot(data = EMP$micro,design = EMP$mapping,min_relative = 0.001,min_ratio = 0.7,cooc_method = 'spearman',cooc_output = T)
##show attributes of net analysis profile at different levels
cooc_re$cooc_profile$species
## If some globe parameters do not works well for specific plot,
## ** cooc_plot_each ** is prepared for this.
cooc_plot_each(result$plot$Control$species,cooc_output=T,vertex.size = 8,
vertex.label.cex =2 ,edge.width =2 )

## Load data
library(EasyMicroPlot)
data(EMP)
## Generate Top abundance barplot (easy running in default parameters)
structure_re <- structure_plot(data = EMP$micro,design = EMP$mapping,
min_relative = 0.001,min_ratio = 0.7)
## Plot
structure_re$pic$genus$barplot$Total
structure_re$pic$genus$barplot$CT
## Show top abundance taxonomy
structure_re$result$filter_data$species
structure_re$result$filter_data$species_ID
## Support ggplot theme parameters
library(ggplot2)
newtheme_slope=theme(axis.text.x =element_text(angle = 45, hjust = 1,size = 10))
structure_re <- structure_plot(data = EMP$micro,design = EMP$mapping,
min_relative = 0.001,min_ratio = 0.7,mytheme =newtheme_slope)
structure_re$pic$family$barplot$Total

## Load data
library(EasyMicroPlot)
data(EMP)
## Data preparation
core_data <- data_filter(data = EMP$micro,design = EMP$mapping,
min_relative = 0.001,min_ratio = 0.7)
sp <- core_data$filter_data$species
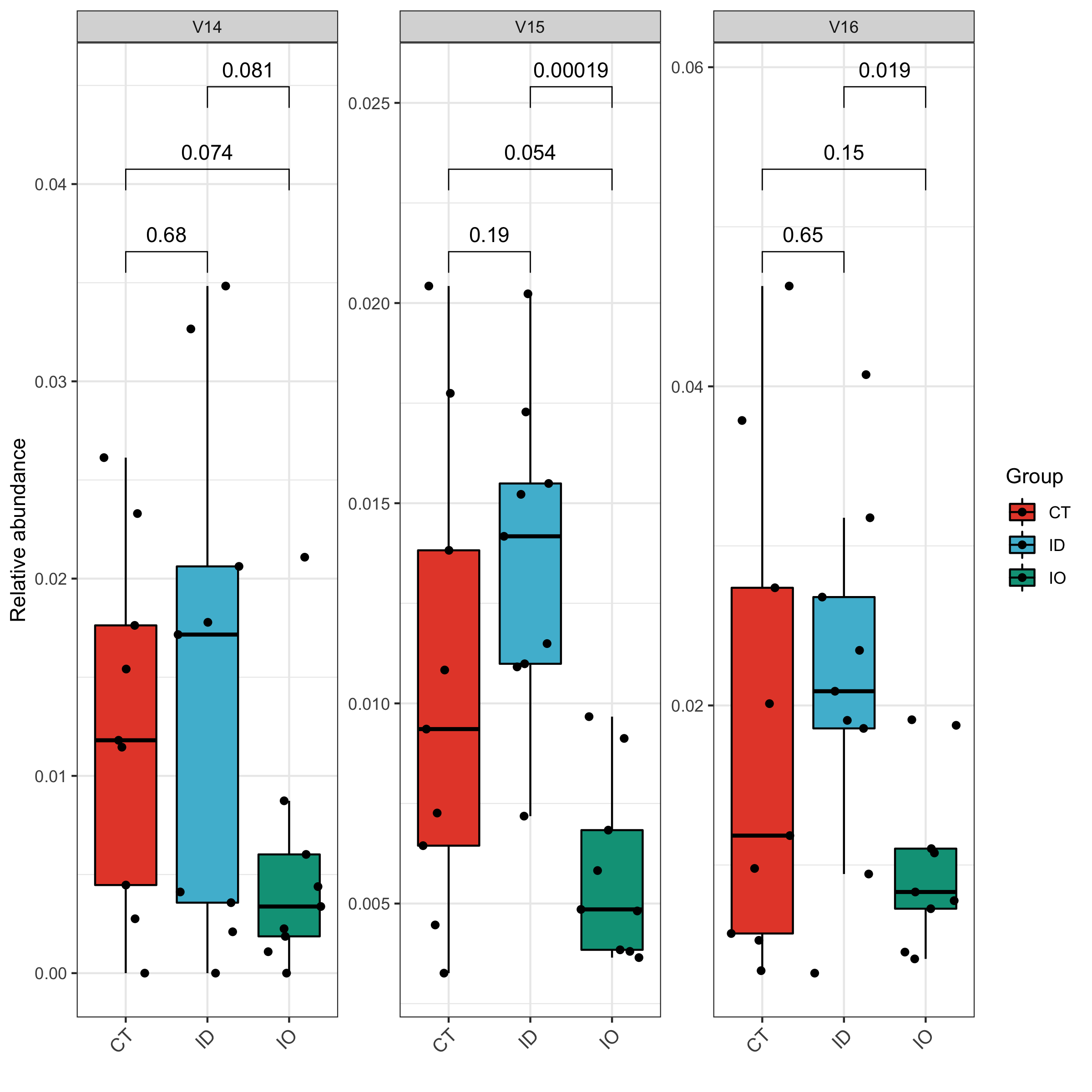
## Taxonomy boxplot
## For Student-Test in comparation
tax_re <- tax_plot(data = sp,tax_select = c('V14','V15','V16'),method = 'ttest')
## For one-way ANOVA and Post Hoc test
tax_re <- tax_plot(data = sp,tax_select = c('V14','V15','V16'),method = 'SNK')
## Plot
tax_re$pic$V14
## Interactive visualization
tax_re$html$V14
## Support ggplot theme parameters
library(ggplot2)
newtheme_slope=theme(axis.text.x =element_text(angle = 45, hjust = 1,size = 10))
tax_re <- tax_plot(data = sp,tax_select = c('V14','V15','V16'),method = 'ttest',mytheme = newtheme_slope)

## Load data
library(EasyMicroPlot)
data(EMP)
## Generate core data
core_data <- data_filter(data = EMP$micro,design = EMP$mapping,
min_relative = 0.001,min_ratio = 0.7)
## Prepare training data
sp <- core_data$filter_data$species
## Generate rfcv
RFCV_result <- RFCV(sp)
## Show RFCV curve
RFCV_result$RFCV_result_plot$curve_plot
## Show intersection and union tax among all the best models under different random number seed
RFCV_result$RFCV_result_plot$intersect_num
RFCV_result$RFCV_result_plot$union_num

## Load data
library(EasyMicroPlot)
data(EMP)
## Generate core data
core_data <- data_filter(data = EMP$micro,design = EMP$mapping,
min_relative = 0.001,min_ratio = 0.7)
## Prepare training data
sp <- core_data$filter_data$species
## Modify the group into two cases when multiple group exists.
sp <- RFCV_data_binary(sp,rf_estimate_group = 'ID',id_not = 'NO_ID')
## Load data
library(EasyMicroPlot)
data(EMP)
## Generate core data
core_data <- data_filter(data = EMP$micro,design = EMP$mapping,
min_relative = 0.001,min_ratio = 0.7)
## Prepare training data
sp <- core_data$filter_data$species
## Modify the group into two cases when multiple groups exist.
sp <- RFCV_data_binary(sp,rf_estimate_group = 'CT',id_not = 'Cases')
## Generate rfcv
RFCV_result <- RFCV(sp)
## Show intersection and union tax among all the best models under different random number seed
RFCV_result$RFCV_result_plot$intersect_num
RFCV_result$RFCV_result_plot$union_num
## ROC
## Parameter *rf_tax_select* allows users to select optional tax
## **union** and **intersect** means the intersection and union tax of models above
RFCV_roc(RFCV_result,rf_tax_select = c("V143", "V83" , "V101", "V121"),rf_estimate_group = 'Cases')
RFCV_roc(RFCV_result,rf_tax_select = 'union',rf_estimate_group = 'Cases')
RFCV_roc(RFCV_result,rf_tax_select = 'intersect',rf_estimate_group = 'Cases')

## Load data
library(EasyMicroPlot)
data(EMP)
## Prepare the bacteria taxa data
core_data <- data_filter(data = EMP$micro,design = EMP$mapping,
min_relative = 0.001,min_ratio = 0.7)
sp <- core_data$filter_data$species
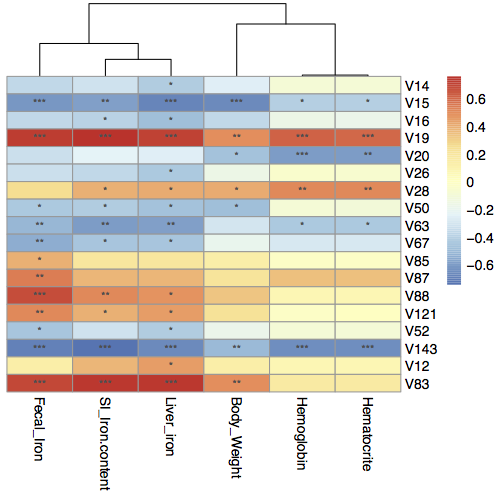
## Analysis and plot
## When ** cor_output = T **, cooc picture is generated in working directory.
cor_re <- EMP_COR(data = sp,meta=EMP$iron,cor_output = T,method = 'spearman',aes_value = 1)
cor_re <- EMP_COR(data = sp,meta=EMP$iron,cor_output = T,method = 'spearman',aes_value = 2)